Week 4: Protein Design Pt 1
Conceptual Questions
(Question 1) A 500g piece of meat would weight about 3.011x1026 Daltons, and since each amino acid is equal to about 100 Daltons, that would mean that by consuming this piece of meat, you are consuming 3.011x1024 amino acids.
(Questions 2) When we eat sources of meat, we physically and enzymatically break down the proteins into amino acids, fatty acids, and sugars, which in turn are used to provide energy to our bodies.
(Question 3) The 20 natural amino acids that are used most regularly today represent the 20 amino acids that contributed to early evolution and allowed for both efficiency and redundancy in order to prevent disasterous mutations[1].
(Question 5) The Miller-Urey experiment [2] famously showed that the chemistry within the early Earth’s atmosphere contributed to the abiogenesis of amino acids, proving their initial evolution through chemical synthesis.
(Question 6) Most natural proteins form the L configuration, resulting in right-handed alpha helices. Since D-amino acids are mirror images of L-amino acids, this would mean that they would form left-handed alpha helices.
(Question 7) There are other helice formations besides the alpha helix, such as the pi helix, which is a wider helix [3]. New helical structures can also be designed using synthetic biology by incorporating non-canonical amino acids in order to design new hydrogen-bonding patterns [4].
(Question 8) Most molecular helices are right-handed becuase life mostly uses L-amino acids, which favors right-handed helices for stability.
(Question 9) Beta sheets tend to aggregate becuase their backbone is extendedly exposed, which means that hydrogen bonds can form easier between subparts without steric hindrance. These hydrogen bonds are the driving force for beta sheet aggregation.
(Question 10) Amyloid diseases form beta sheets often due to the ability to aggregate and their extreme stability [5]. This is what also makes these amyloid beta sheets attractice material candidates due to their strength and ability to self-assemble.
Protein Analysis and Visualization
I wanted to expand upon what I had done in week 2 with Calreticulin (CALR), since I also used it as the premise of one of my final project ideas. CALR is a pro-healing cue in wound healing [6] and supports the progression through the four wound healing phases (hemostasis, inflammation, proliferation, and remodeling) [7], which is classically disrupted during chronic wounds [8].
Protein sequence[9]:
This protein sequence is 416 amino acids long and the most frequent amino acid is aspartate (D), which appears 55 times. There are 250 identified homologs [10] Calreticulin belongs to the calreticulin family of proteins, which are highly conserved ER-resident caperones [11].
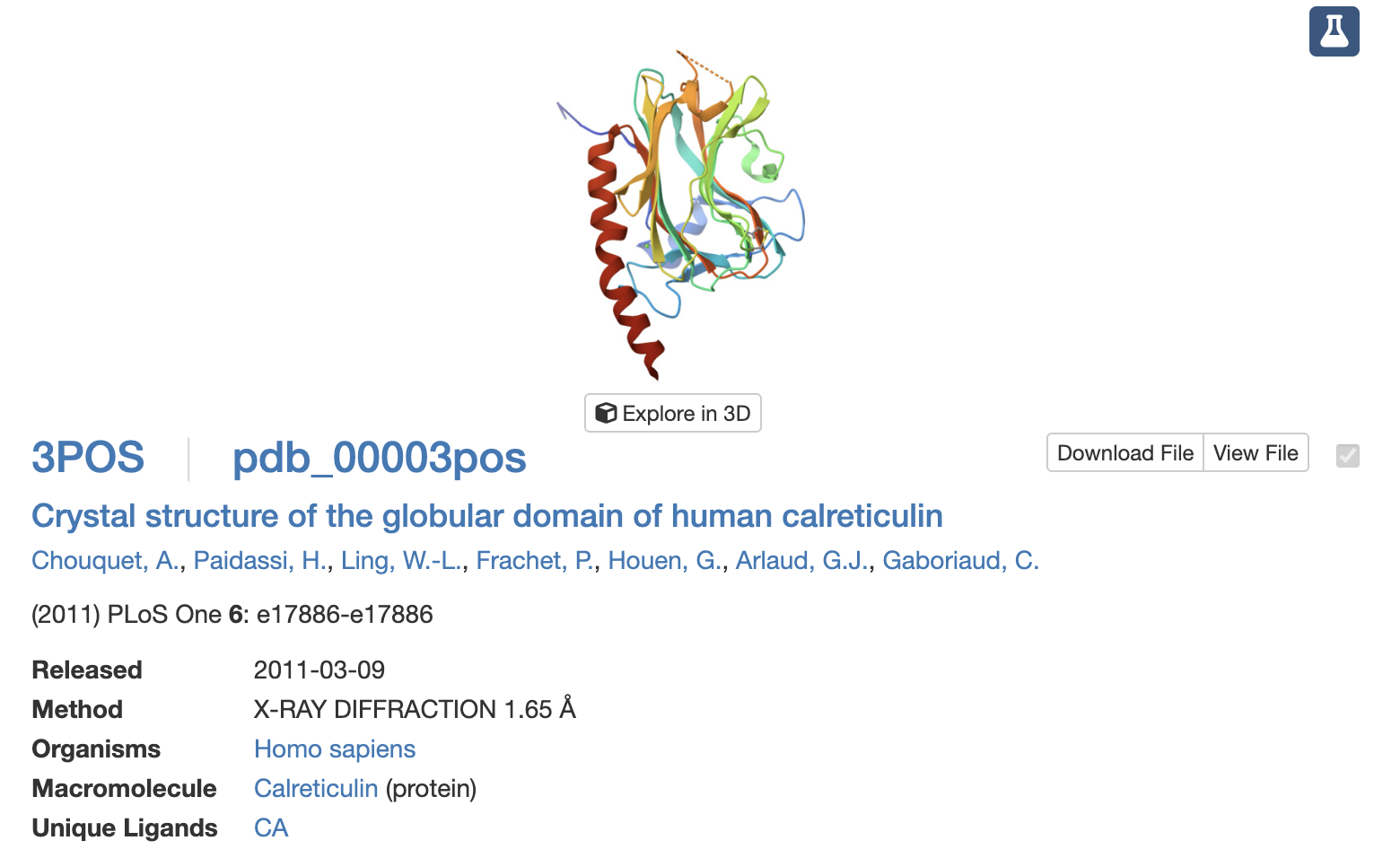

 This structure was discovered in 2011 and has a resolution of 1.65A. It belongs to the Concanavalin A-like lectins/glucanases structural classification family [12].
This structure was discovered in 2011 and has a resolution of 1.65A. It belongs to the Concanavalin A-like lectins/glucanases structural classification family [12].
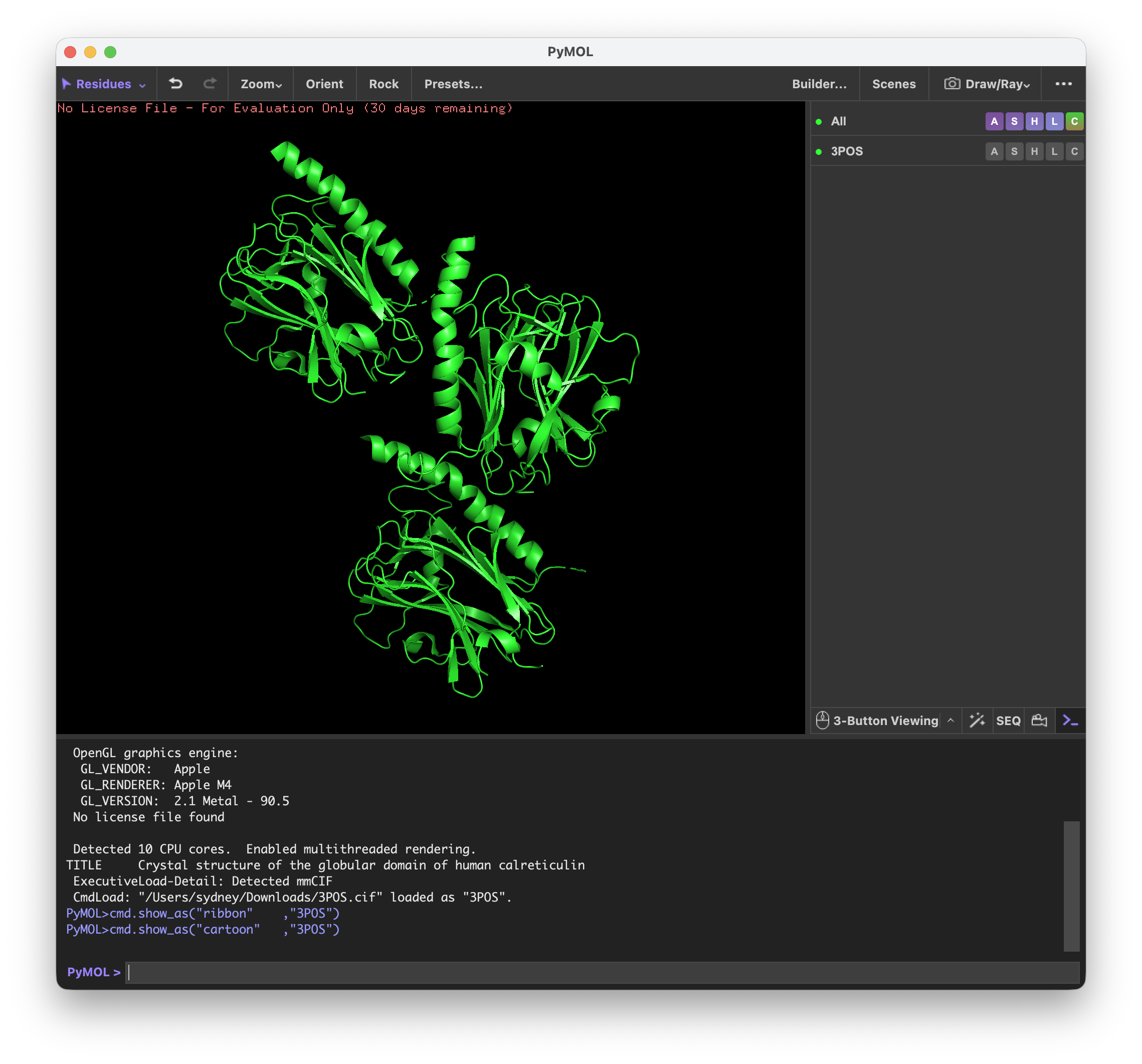
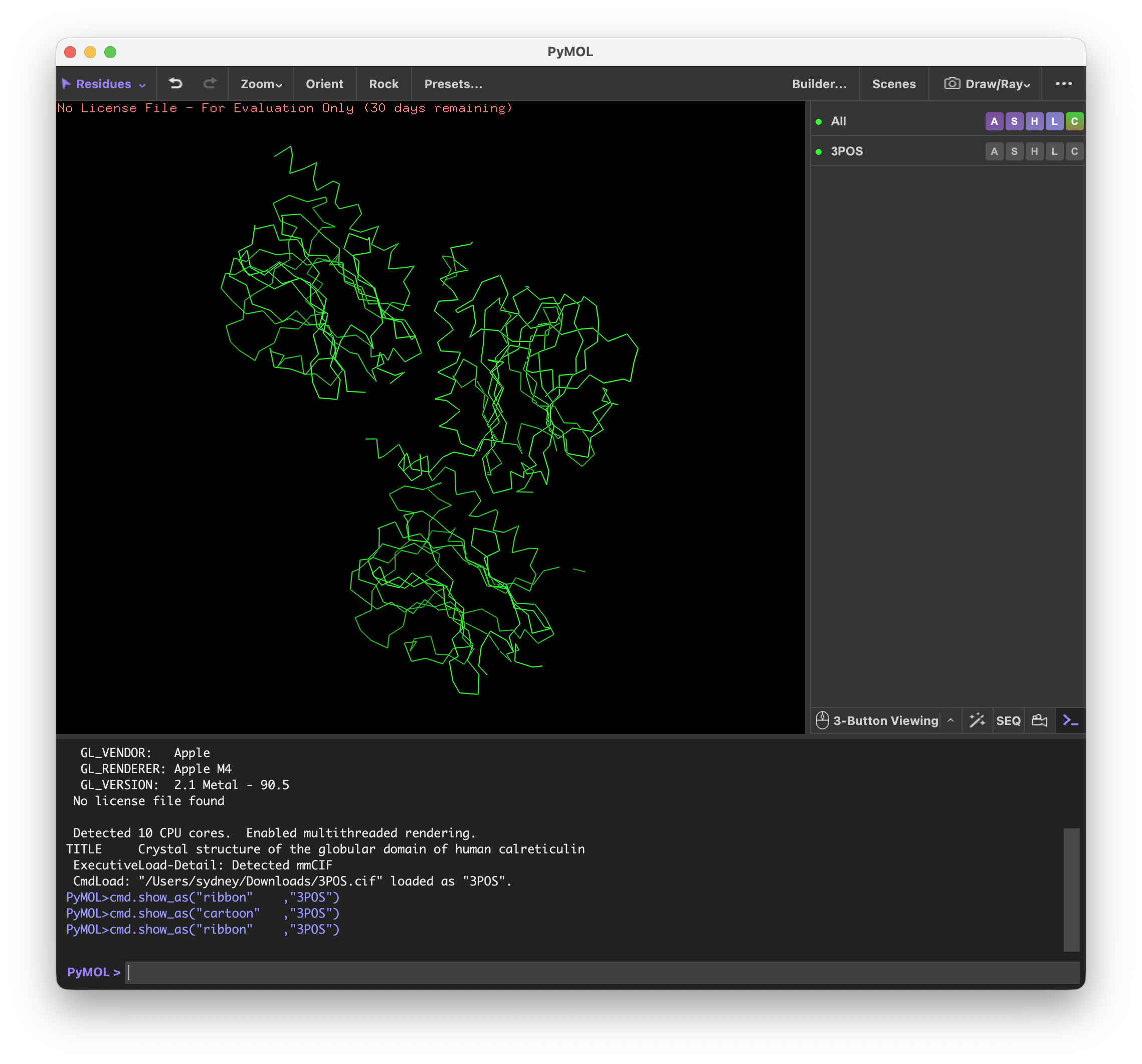
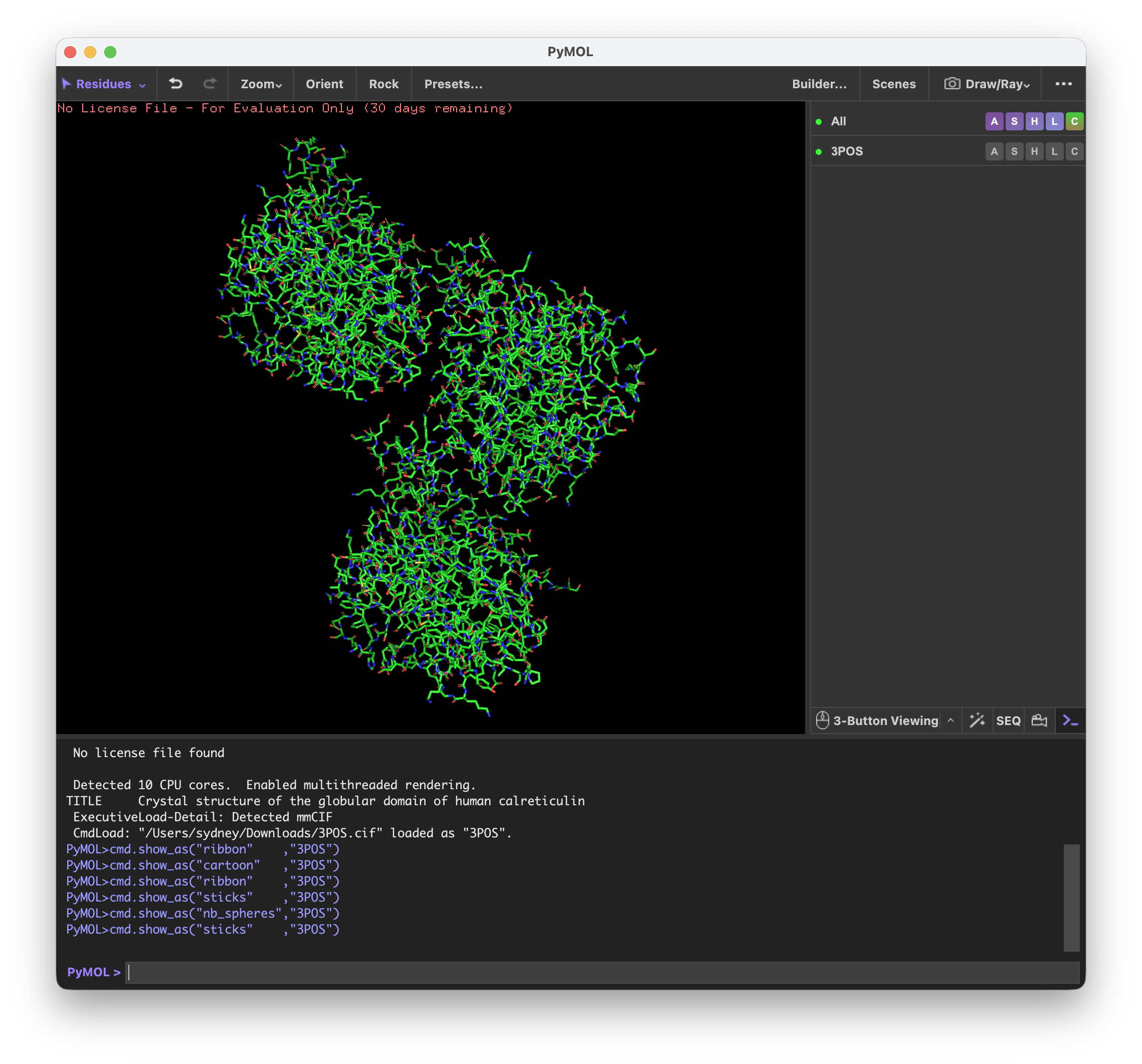
I’ve never used PyMol before, so it was quite interesting to explore the different functions.
First I visualized CALR in the cartoon, ribbon, and ball and stick visualization.
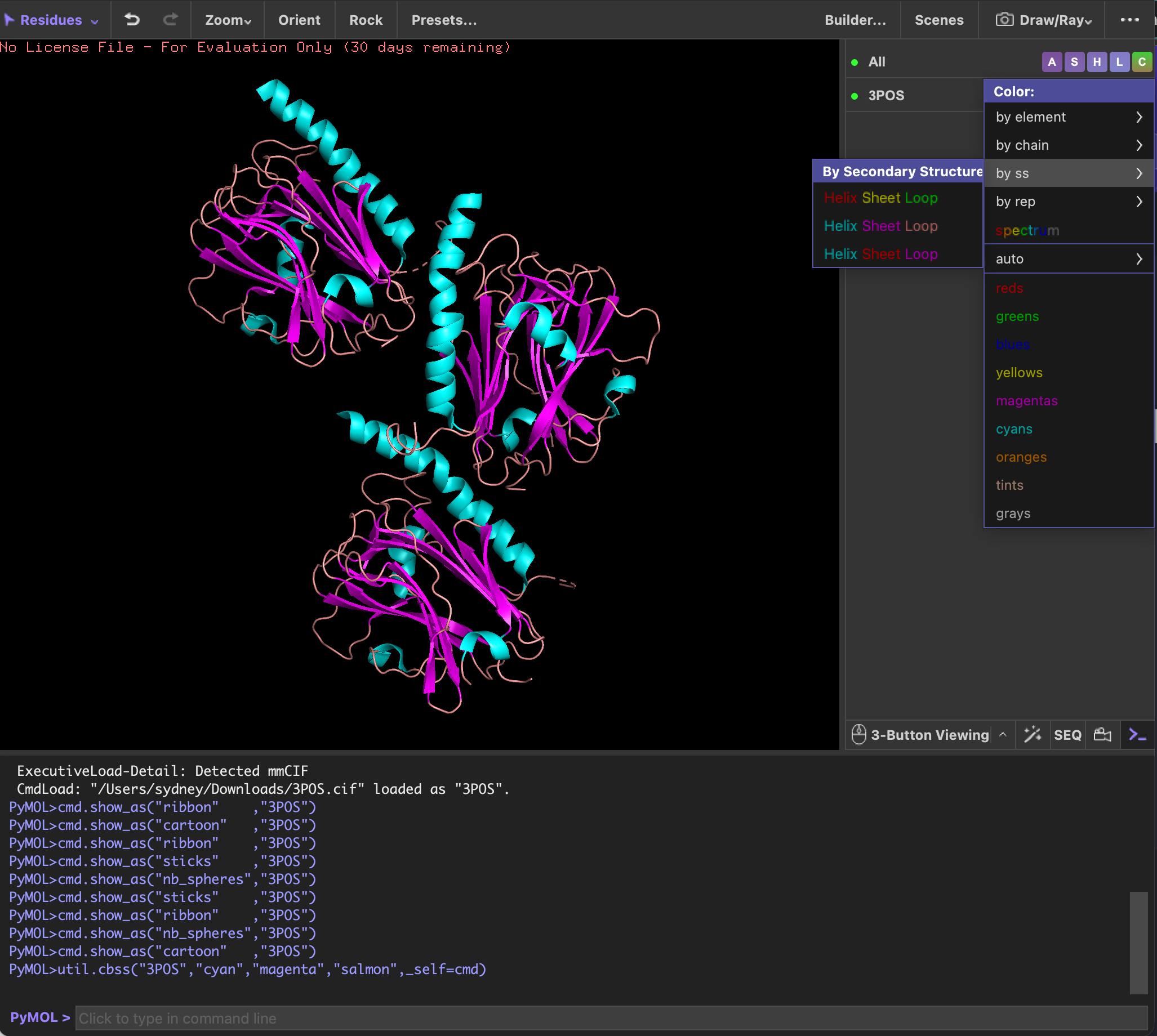

Next, I colored it by the secondary structures and noticed that the structure is mostly made up of beta sheets.
I wasn’t able to figure out how to color the structure by residues.
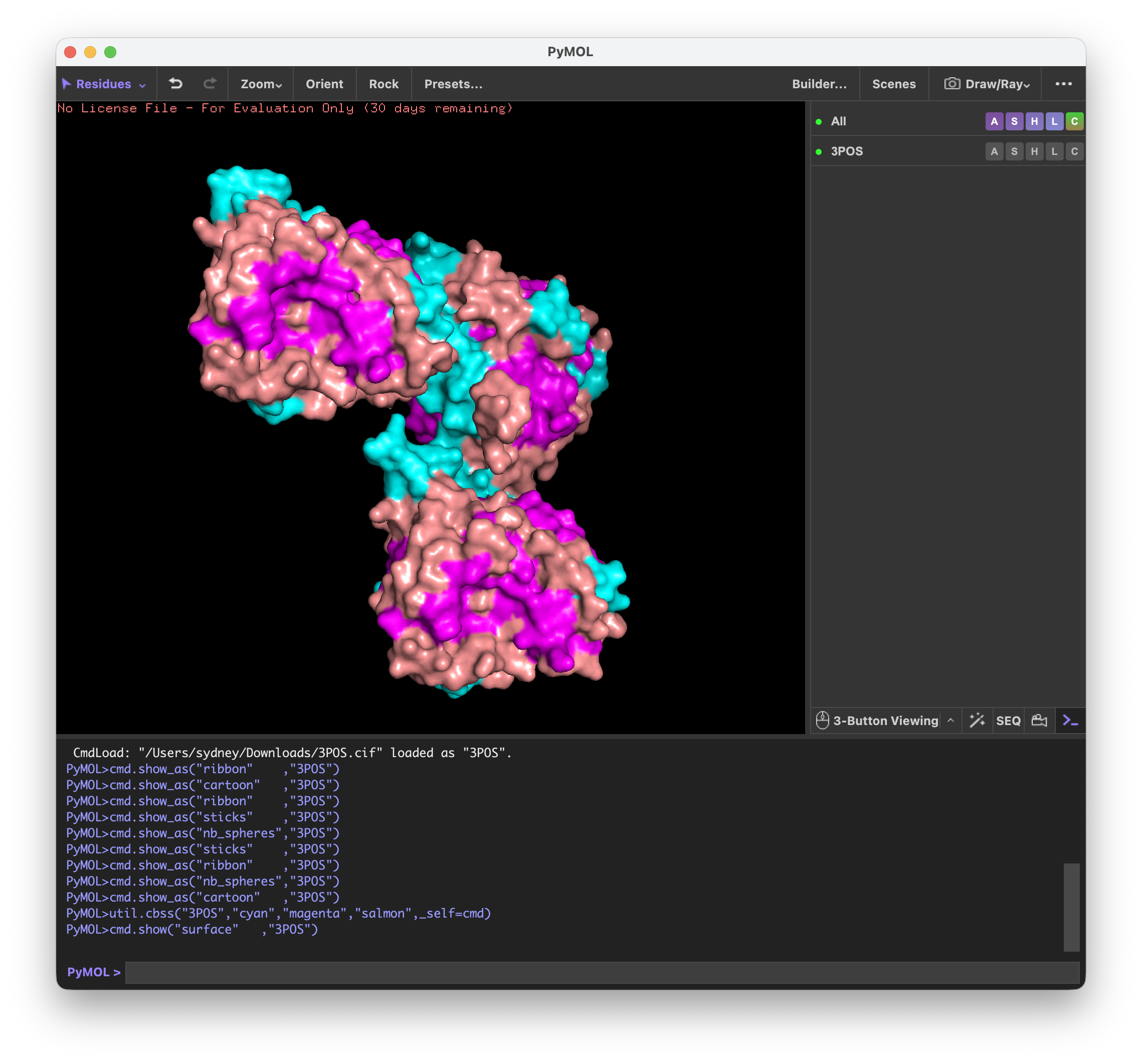

By looking at the surface of CALR, I determined that the binding pocket from this part of the protein was most likely the hook part of the ‘upside down L’.
Using ML-Based Protein Design Tools
Protein Language Modeling
I once again decided to use the crystal structure of the calreticulin lectin domain as my model protein for this part of the homework.
1. Mutation Scan Heatmap
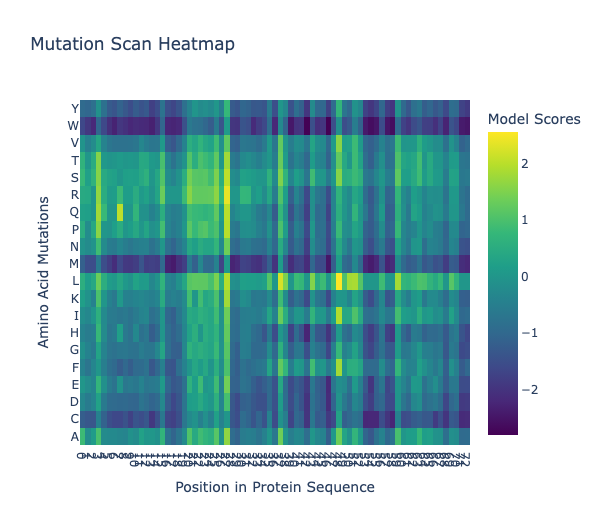
 In the leucine row, it appears that most leucine mutations would be tolerated, apparent by its bright yellow to green band.
In the leucine row, it appears that most leucine mutations would be tolerated, apparent by its bright yellow to green band.
Protein Folding
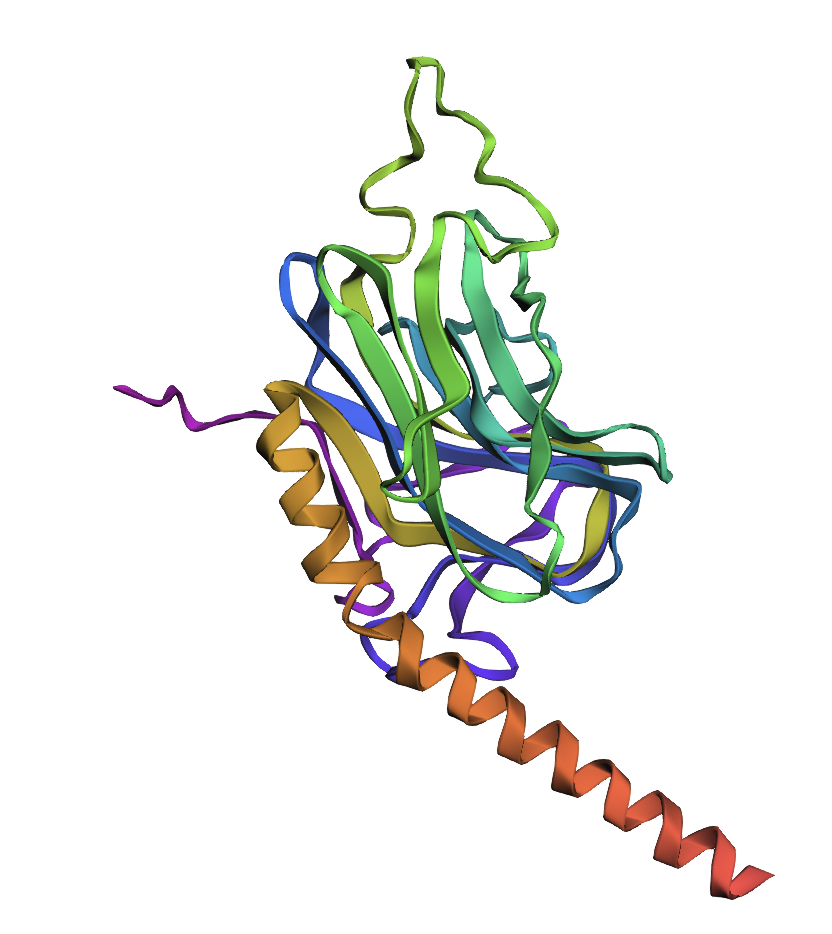

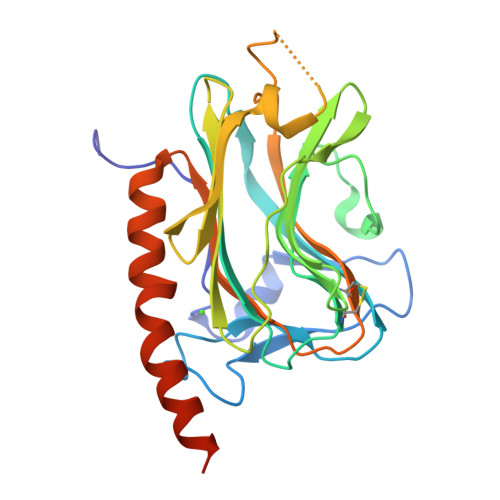
 The first image is of the generated structure by ESMFold and the second is the proven structure, which seems to match quite well.
The first image is of the generated structure by ESMFold and the second is the proven structure, which seems to match quite well.