Week 6: Genetic Circuits Pt 1
DNA Assembly
(Question 1) Within Phusion High-Fidelity Master Mix [1], there is a Phusion DNA Polymerase (which enzymatically synthesizes the DNA in the 5’ to 3’ direction), nucleotides (the building blocks of the synthesized DNA), and an optimized reaction buffer (maintains optimal conditions for the polymerase).
(Question 2) Some factors that determine primer annealing temperature are the primer length, the GC content, and the salt concentration.
(Question 3) While PCR and restriction enzyme digest both produce linear DNA, they differ in terms of when they can be used. For example, with PCR, it requires the design of fragment-specific primers and relies on the use of a polymerase to synthesize the specified DNA fragment. For restriction enzyme digest, it requires the presence of restriction enzyme sites in appropriate locations within the plasmid as well as incubation with the restriction enzyme itself.
(Question 4) To ensure that the DNA sequence will be appropriate for Gibson cloning, it is essential to determine that the sequence contains homologous overhangs (~20-40 bp) to the fragment that you want to combine it with.
(Question 5) When transforming E. coli, the plasmid enters the bacterial cell through pores in the membrane, which are made chemically or through heat shock [2].
(Question 6) Golden Gate is similar to Gibson assembly, in the sense that it combines two or more fragments. For Golden Gate, however, it utilizes type IIS endonucleases in order to make single-stranded fragment-specific overhangs. This process hinges on the creation of primers that will not only amplify the desired fragment, but will also add on the specific overhang, a type IIS restriction enzyme site, and a non-specific overhang on each side of the fragment. Once all of the fragments are amplified using PCR, Golden Gate is performed in a one-pot reaction, combining the fragments, the selected restriction enzyme, ligase, and a buffer.
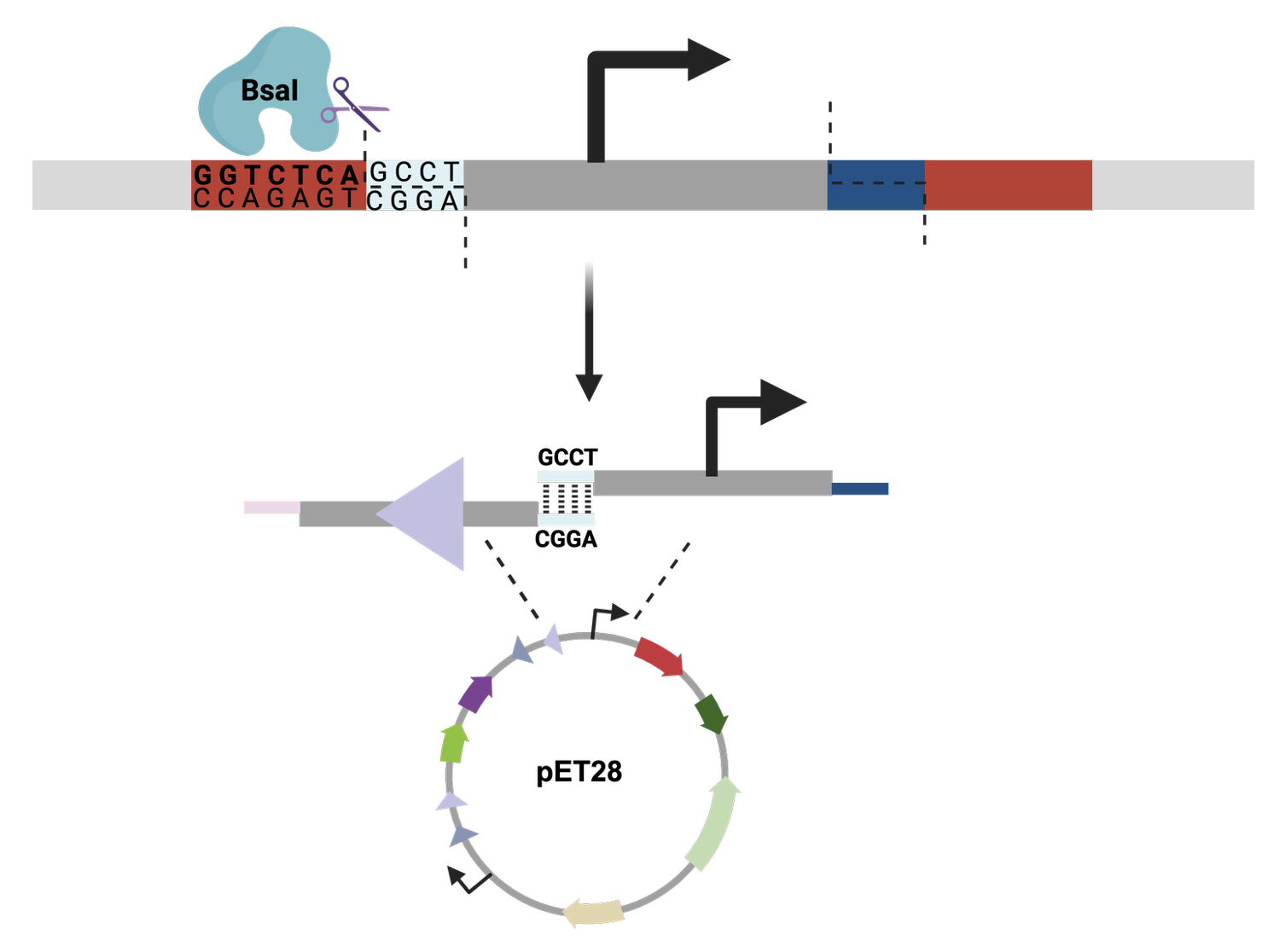

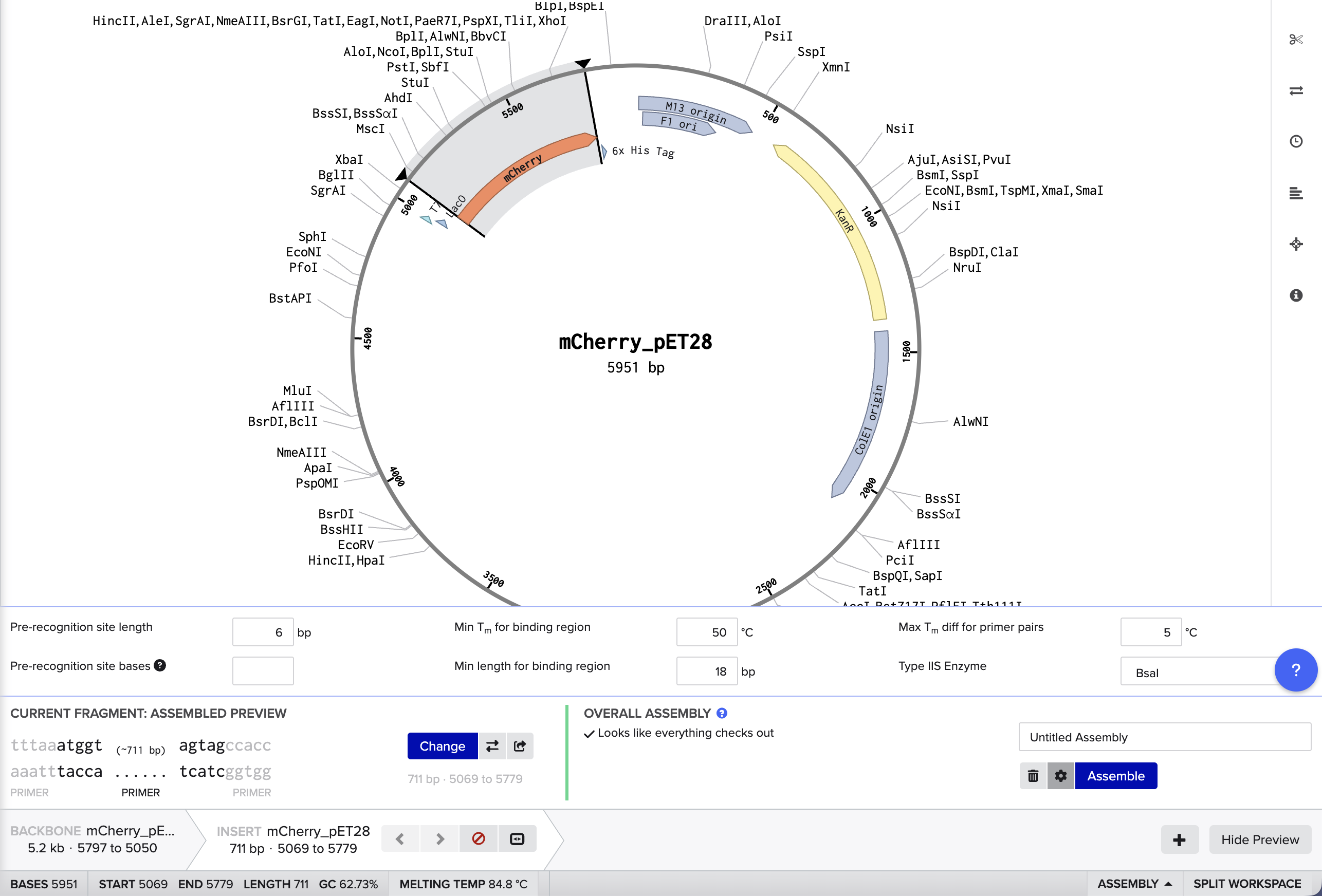
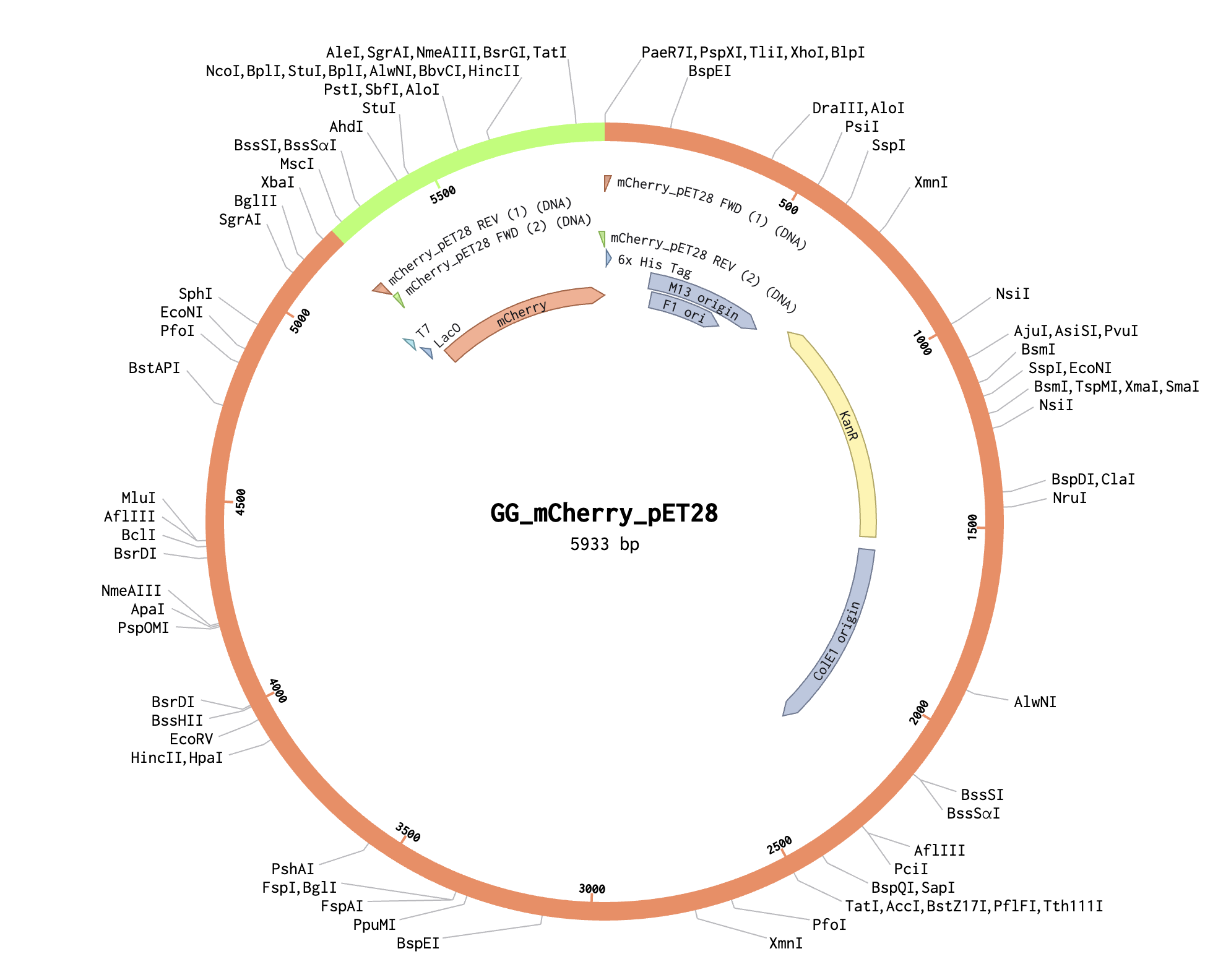
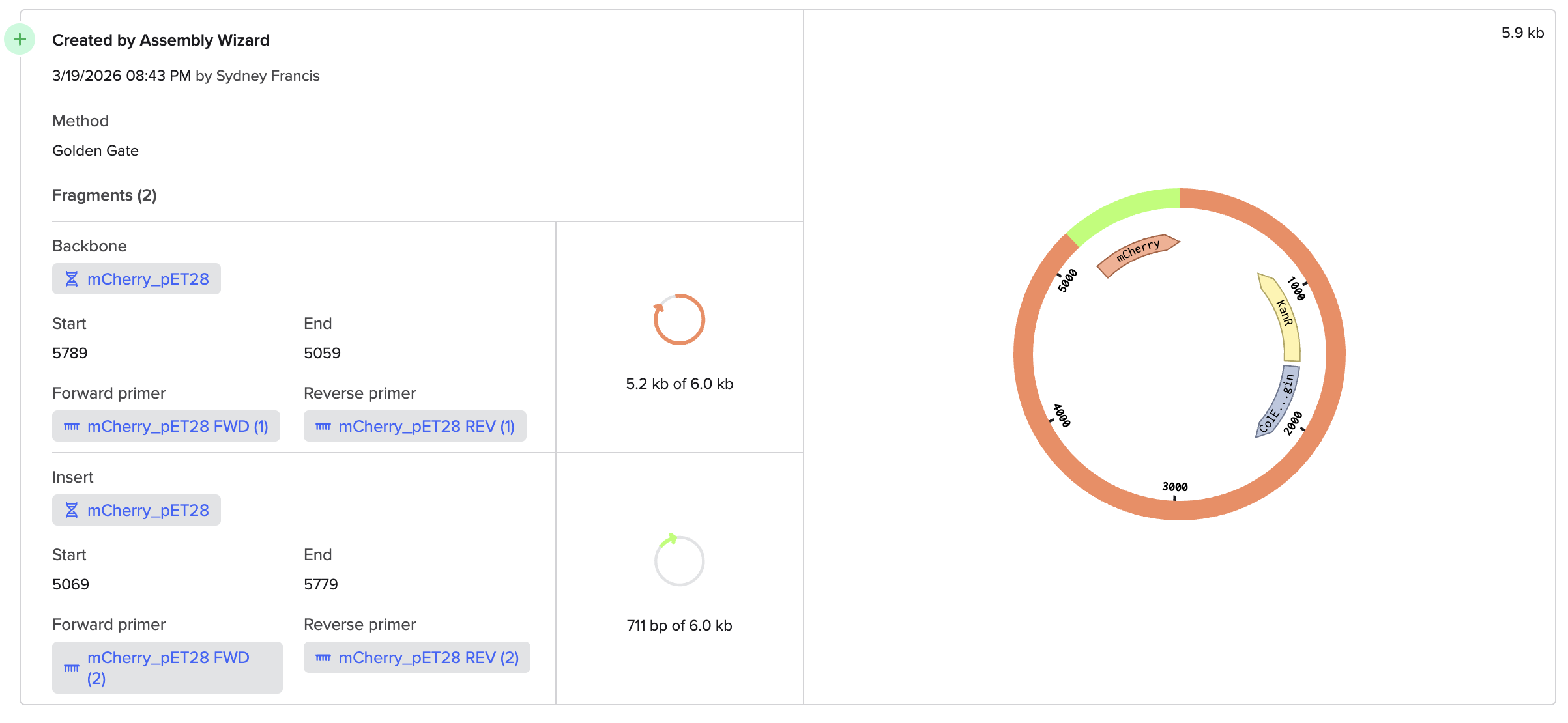
I used Benchling to try to simulate using golden gate to insert mCherry into a pET28 backbone. It was quite simple! I used BsaI as the restriction enzyme.
Asimov Kernel
I have a bit of experience in constructing genetic circuits in the past, but this was my first time using a designated program for it. It was quite interesting to play around with the different circuits and observe the production of certain aspects based on the arrangement.
Next, I moved on to replicating the repressilator on my own by referencing the demo repressilator, which is as follows.
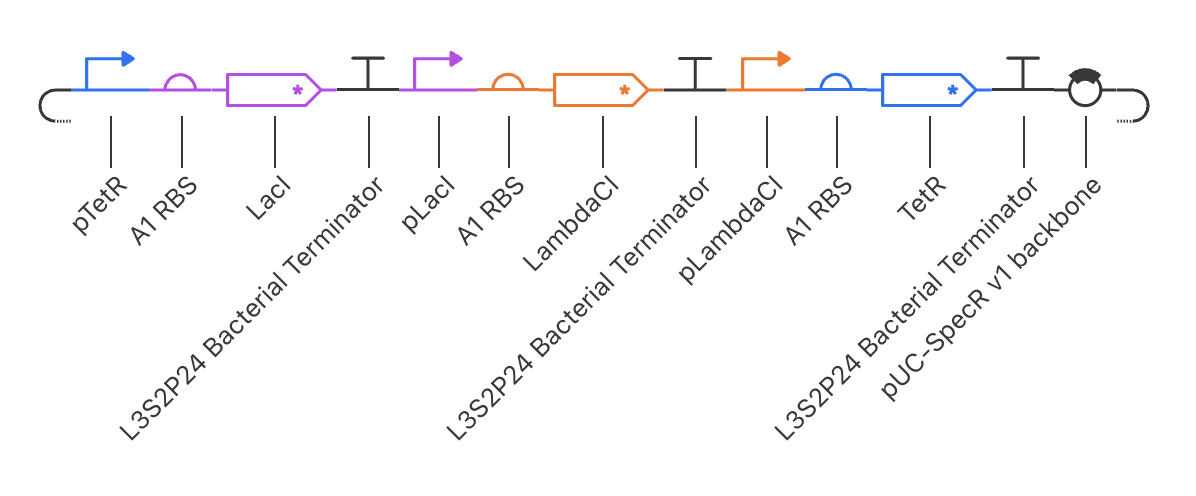
 It ended up turning out like this:
It ended up turning out like this:
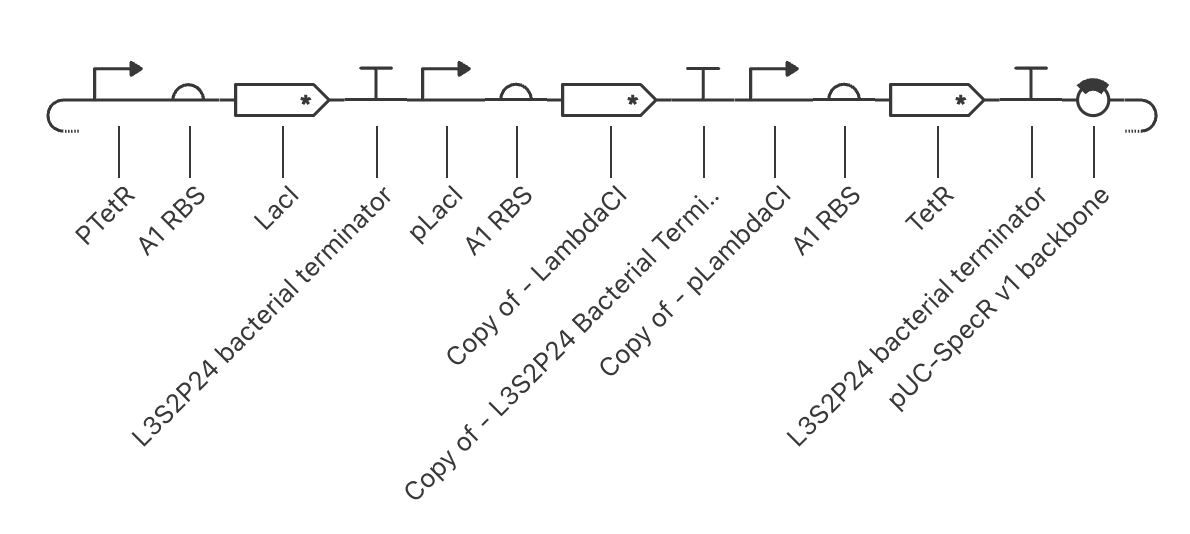

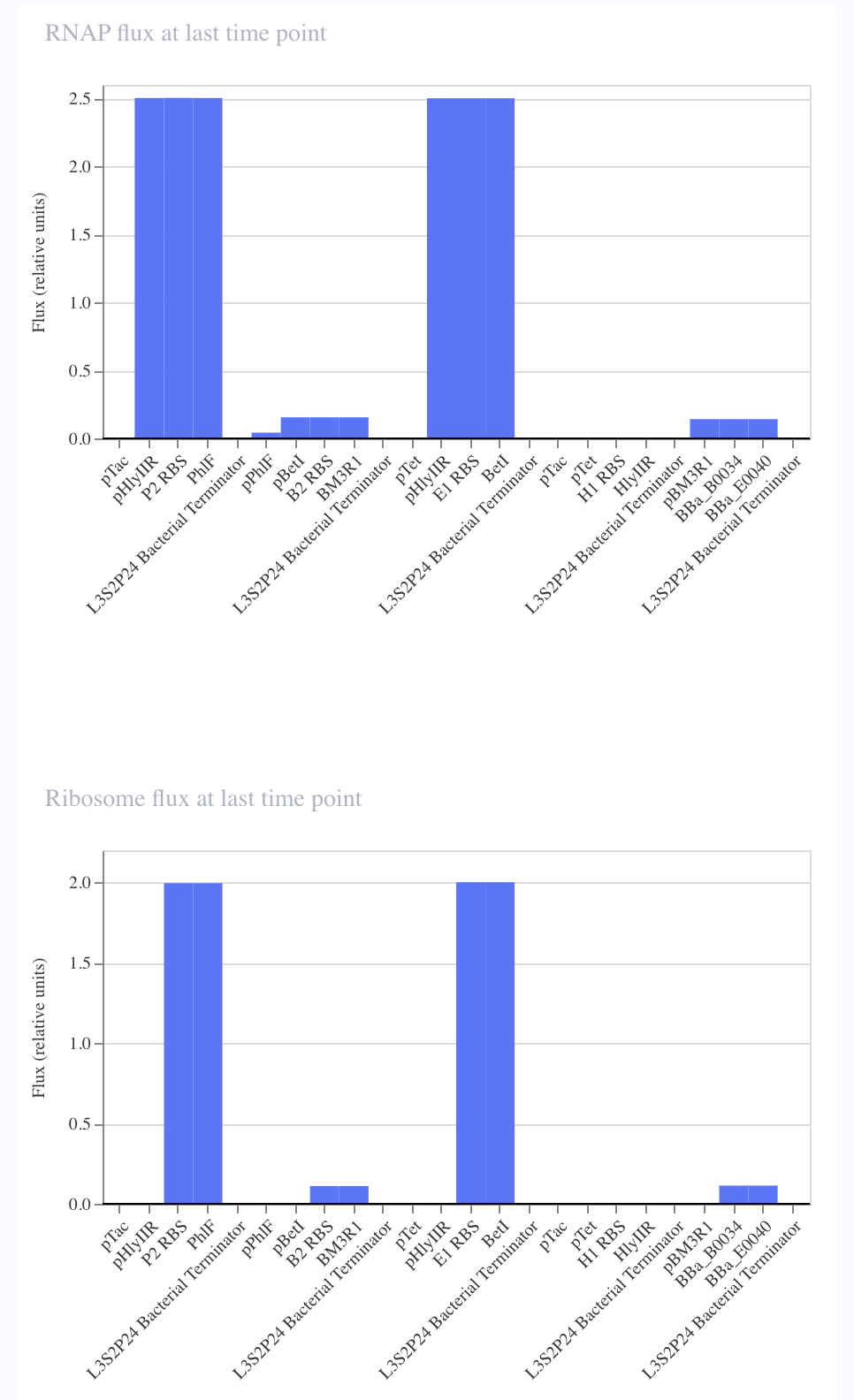
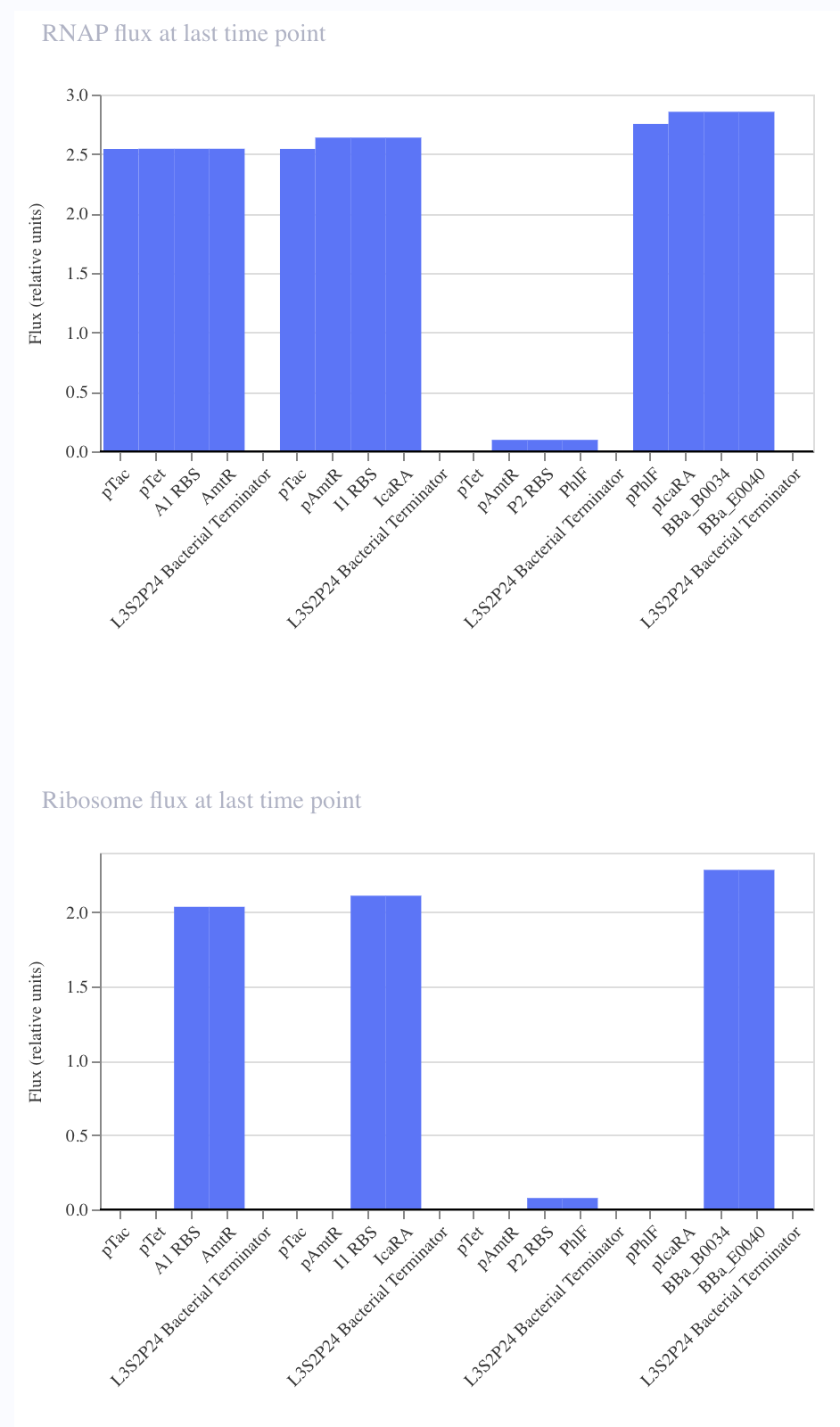
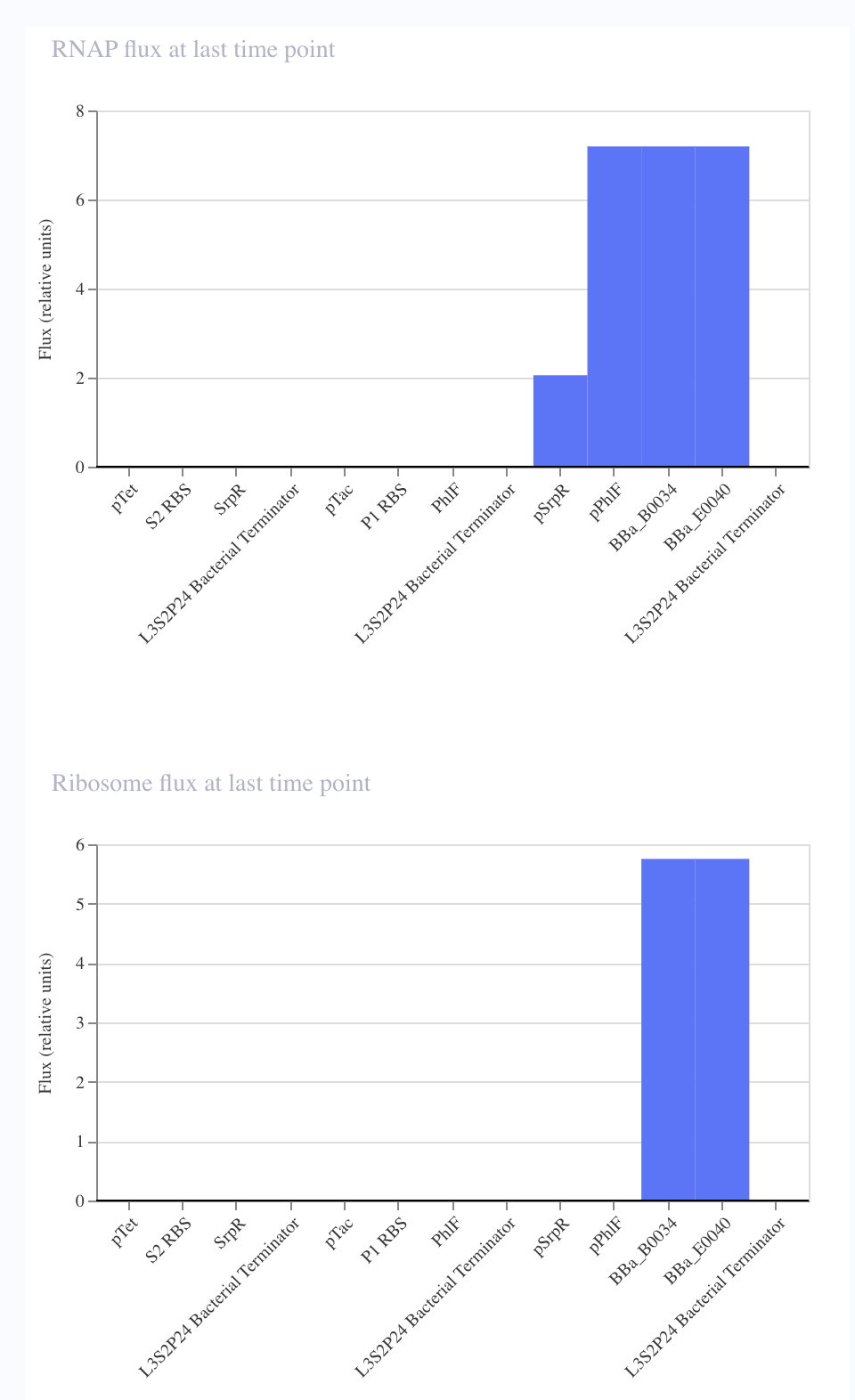
In order to further verify that I replicated the repressilator correctly, I wanted to compare the simulation of each of the circuits.
| Original Repressilator | Replicated Repressilator |
|---|---|
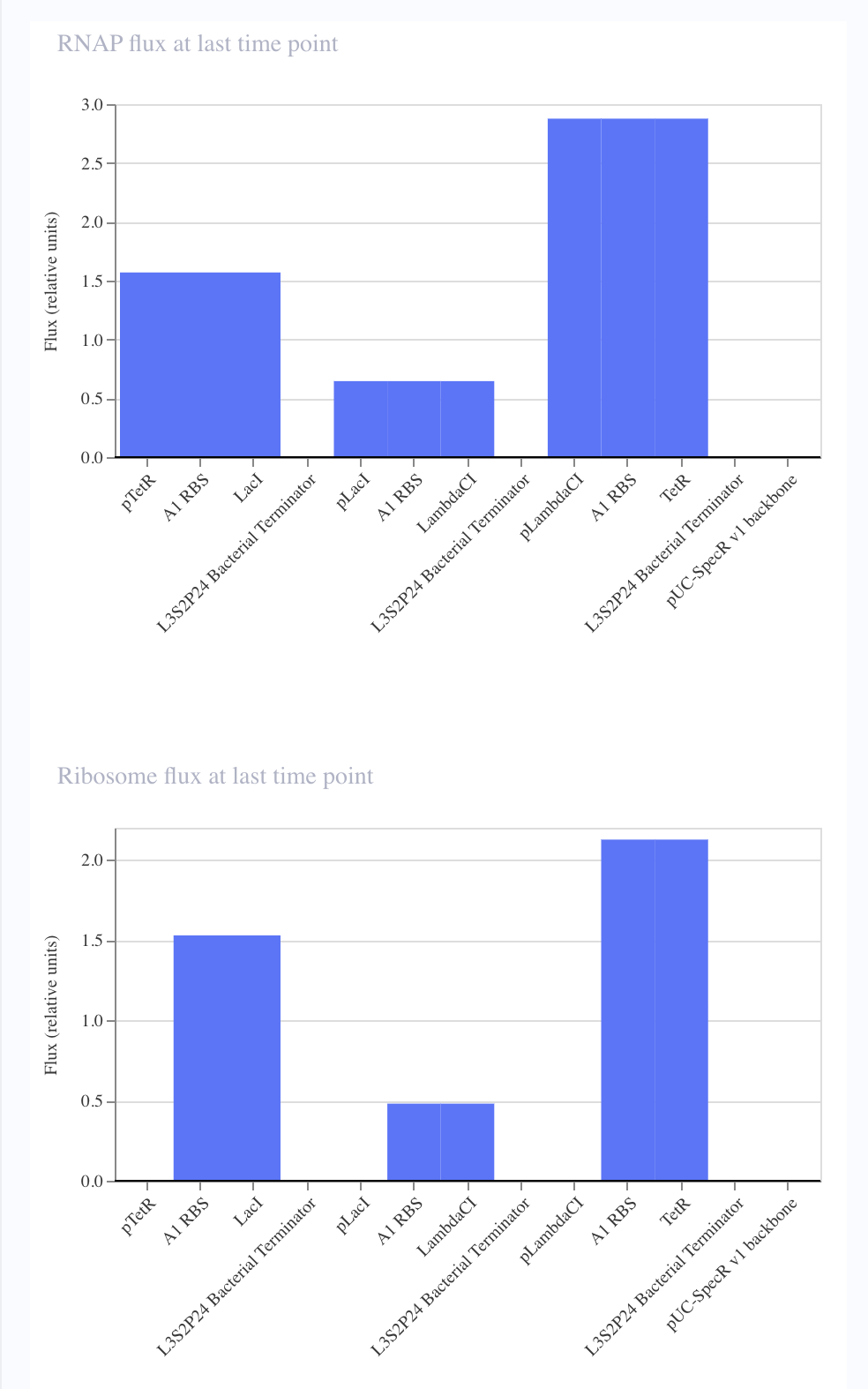
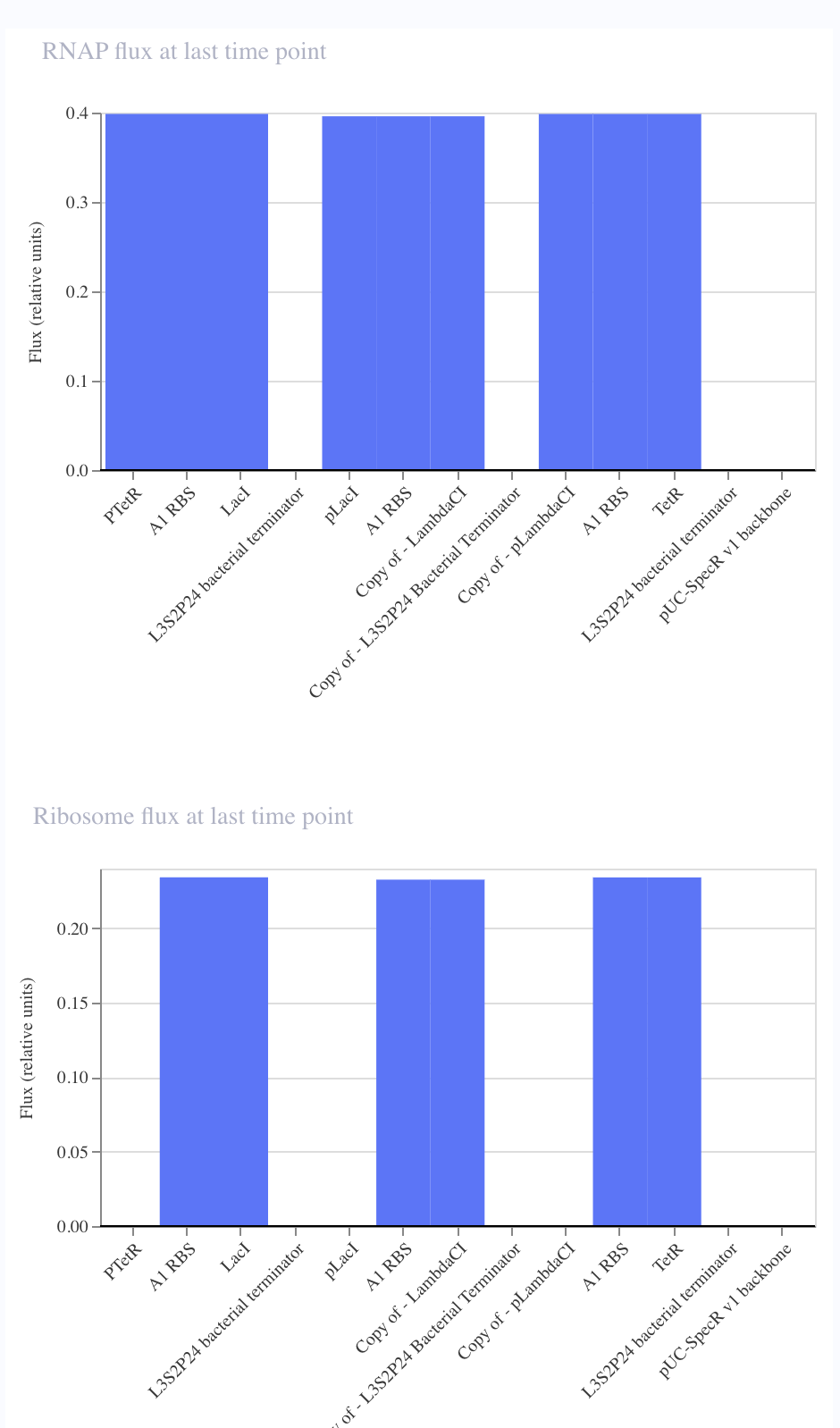
 | 
 |
I noticed that although I copied the repressilator exactly, the expression of the parts did not vary as they did in the original.
After this, I moved towards simulating three of my own constructs.
| Autonomous FLEx Switch | 2 | 3 |
|---|---|---|
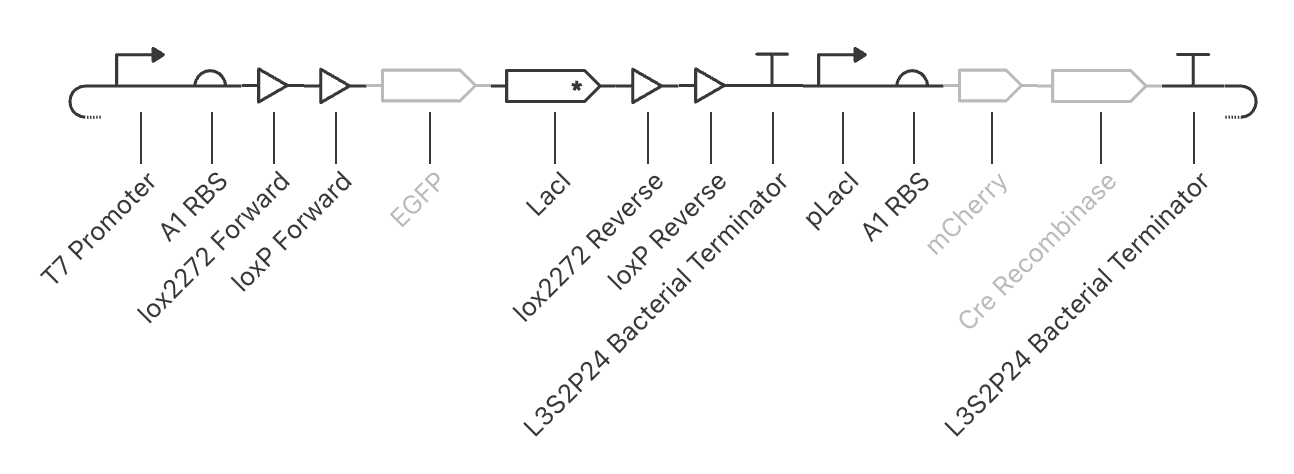
 | ||
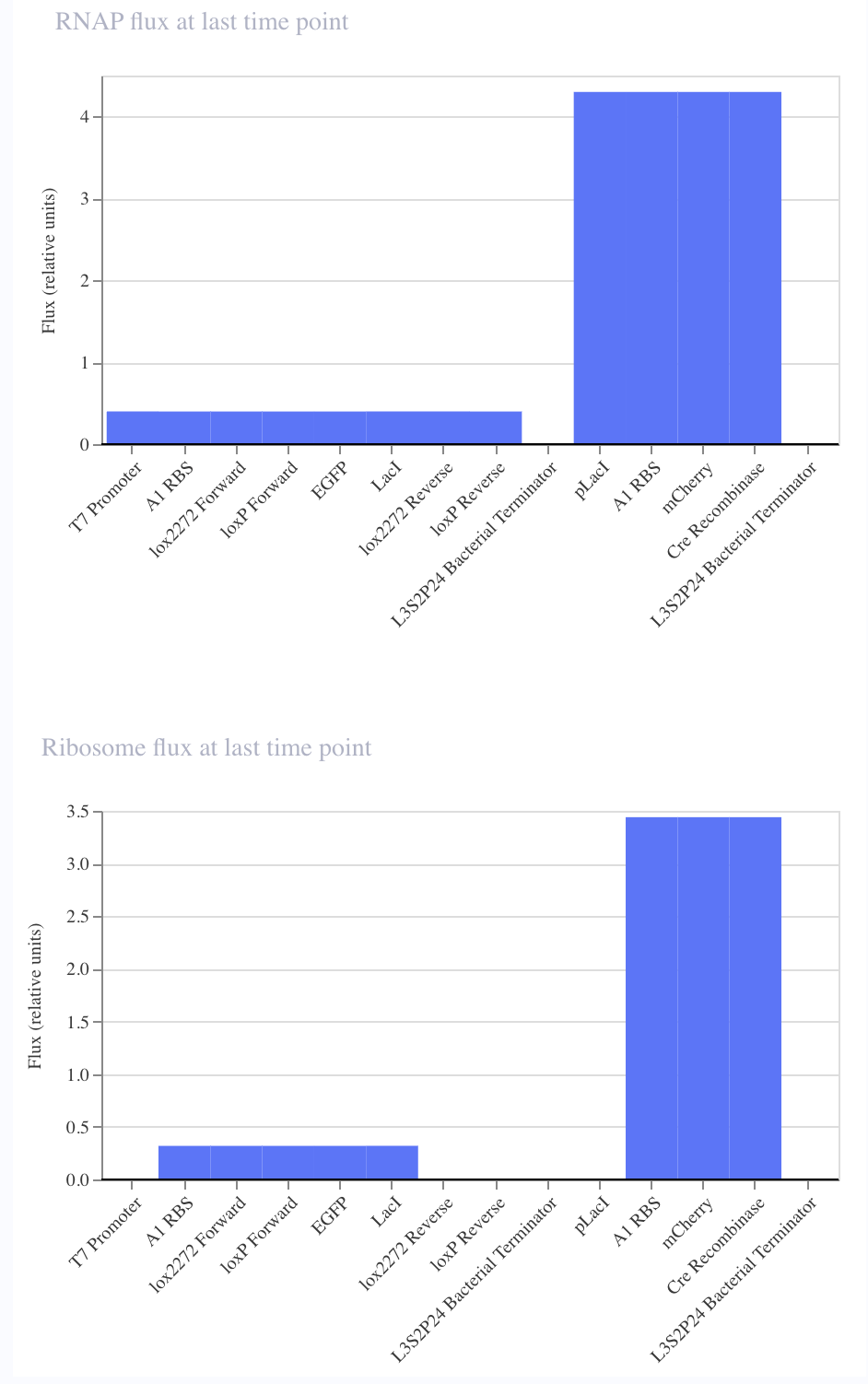
 |
The main concept I used to formulate these circuits was the underlying principle of my final project idea. At its core, my final project relies on the production of two gene sets subsequently. Upon the production of the first gene set, the circuit will then respond to the consequentially produced environment in order to then express the second gene set. However, in order to avoid permanently altering the biological environment, both of the gene sets need to be eventually irreversibly turned off.
The first strategy I used to achieve this was a modification of the FLEx switch to become an autonomously responding circuit. It then uses Cre recombinase and lox2272/loxP sites to then shutoff the first gene set.