Week 2: DNA Read, Write, and Edit
Benchling and In-Silico Gel Art
Simulate Restriction Enzyme Digest
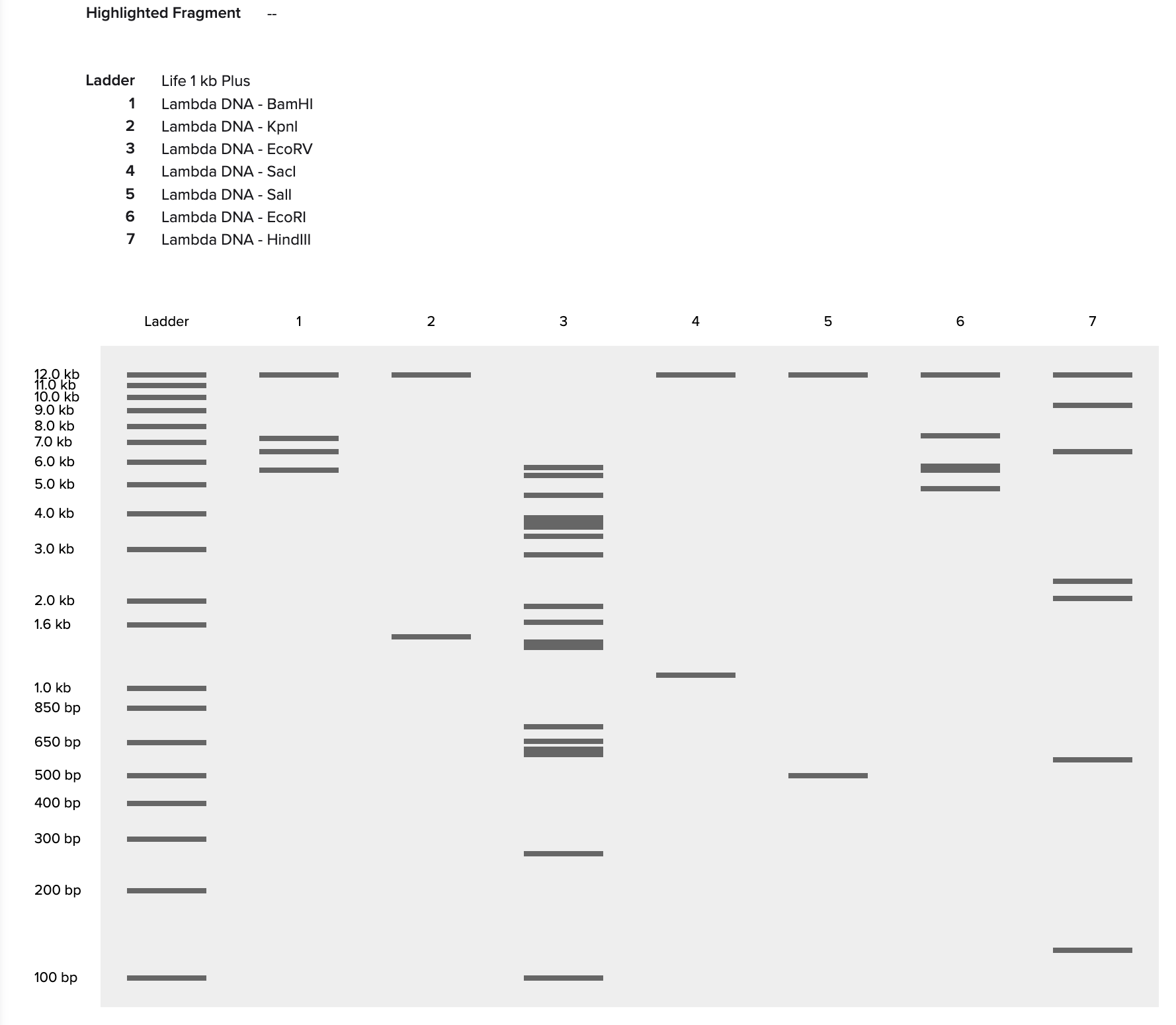


I found this process quite intuitive, as I’ve done similar simulations with the application SnapGene, but it was interesting to notice the small interface differences between the two!
Pattern in the style of Paul Vanouse
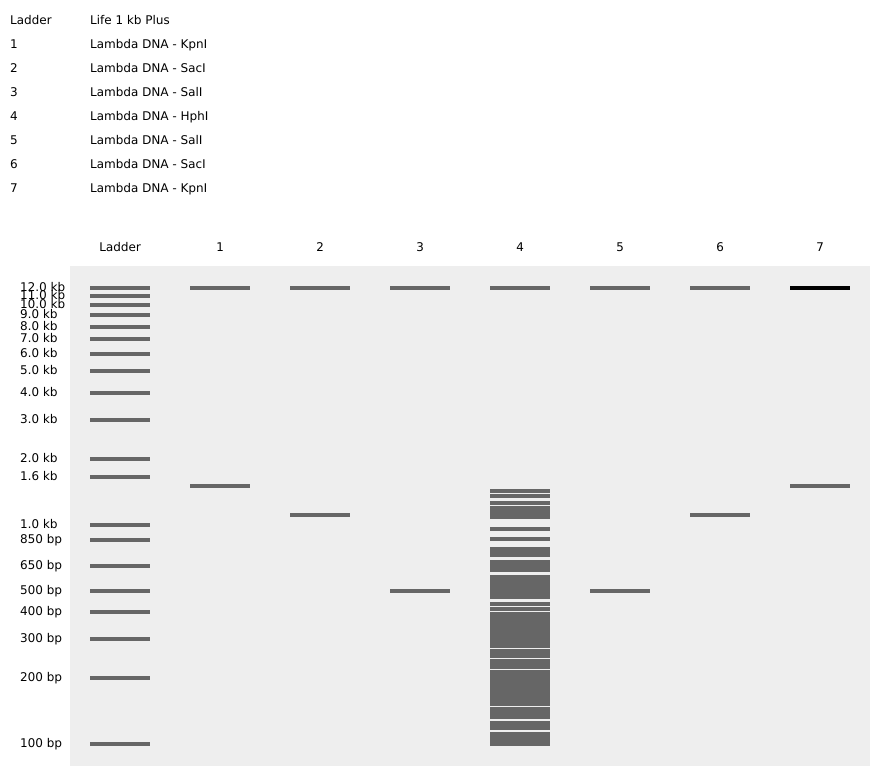


I attempted to make a “Y” for Yonsei, but it turned out to be more difficult than I expected and this was the closest I ended up getting… Huge respect to the people who were able to make a more comprehensive image like the ones that spell MIT!
DNA Design Challenge
My Chosen Protein
I chose to explore Calreticulin (CALR) as my protein of interest for this week due to its role as a pro-healing cue in wound healing[1]. CALR typically serves to support the progression through the four wound healing phases (hemostasis, inflammation, proliferation, and remodeling) [2], which is classically disrupted during chronic wounds [3].
Protein sequence[4]:
Nucleotide sequence[5]:
Codon optimization[6]:
Codon optimization is essential to ensuring the proper and efficient protein expression of a given protein within a specific organism. Typically, different organisms favor different codons that ultimately encode the same amino acid [7], which is why optimizing to the specific organism you intend to use to produce the protein verifies that frequently used codons are encoded instead of rarely used ones within your expression host of choice[8].
In this case, I think that the most applicable technology to produce CALR would be to use HEK293T cells. These cells are human derived and are quickly replicable, meaning that they would be able to prouduce this protein with great efficiency. In order to do this, first I would need to clone my protein insert into a mammalian expression vector with a strong promoter, which would then be tranfected into the HEK293T cells (for example, by lipofection). Within the cells, the RNAP recognizes the promoter and would transcribe the plasmid into RNA. This mRNA would then be translated into protein by the ribosome.
Prepare a Twist DNA Synthesis Order
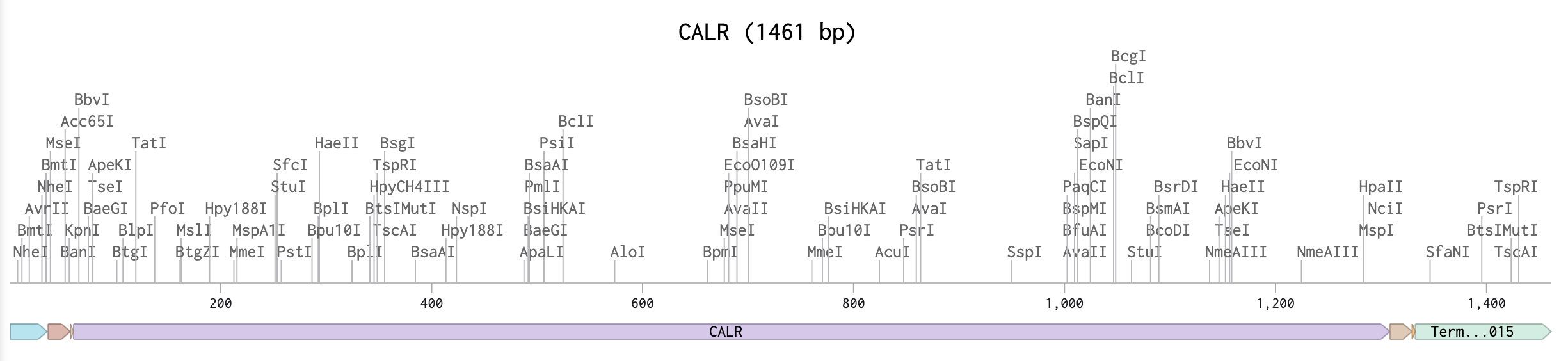
 I prepared my optimized gene within Benchling for Twist [9].
I prepared my optimized gene within Benchling for Twist [9].

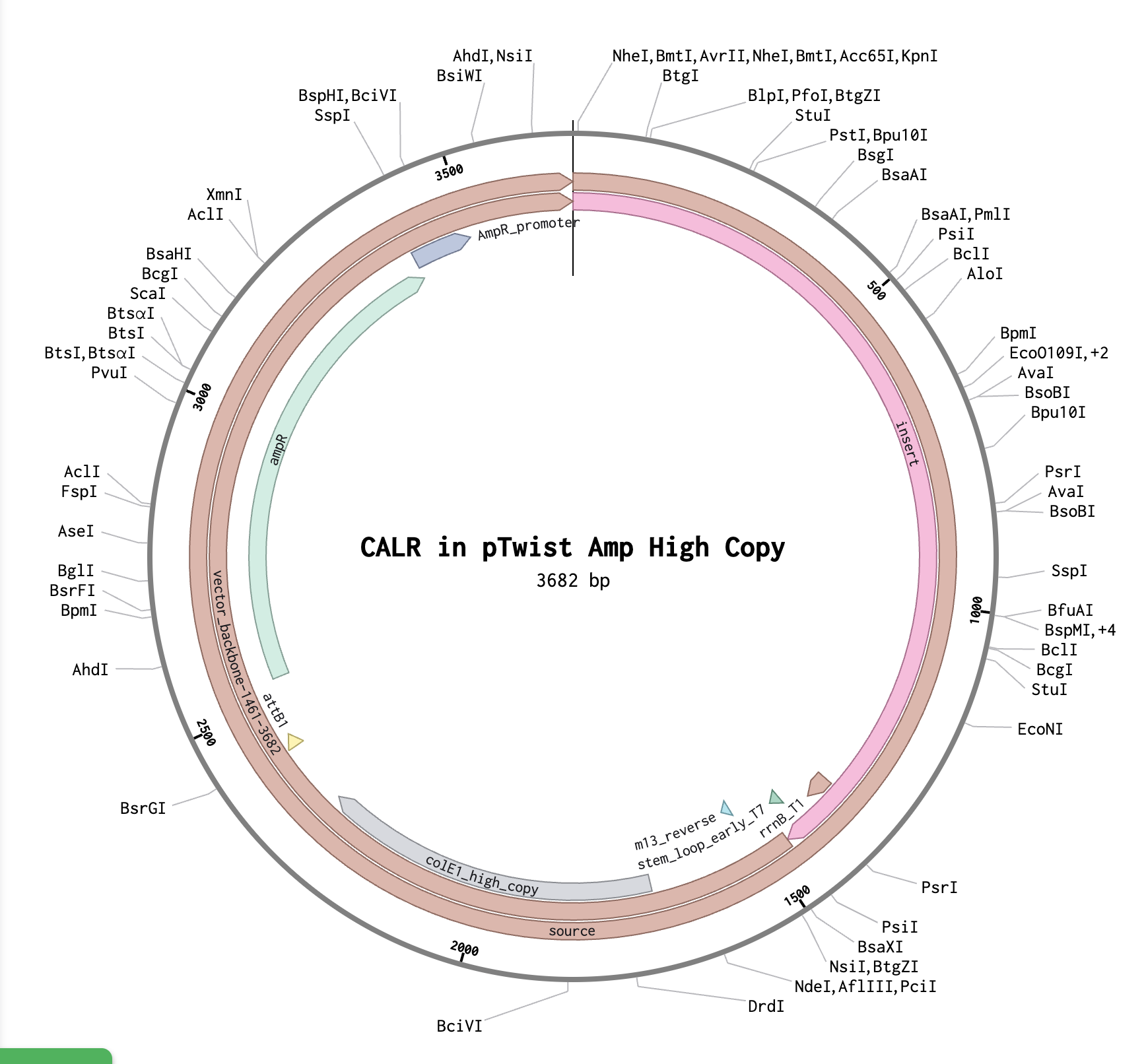
 This order page was so simple compared to other ordering sites I used, and I liked that you could export the entire plasmid as well.
This order page was so simple compared to other ordering sites I used, and I liked that you could export the entire plasmid as well.
DNA Read/Write/Edit
DNA Read
On a similar theme as my previous assignments, one of my main interests is in finding the key molecular mechanisms and differences that distinguish successful wound repair from chronic, non-healing counterparts. For this reason, I’d be interested in being able to compare the mitochondrial genome of healthy (efficiently healing) and chronic wound patients, as the mitochondria has been proven to play a central role in wound metabolism [10][11]. By sequencing these genomes and contrasting the two, it may reveal variants within the genome that could predispose individuals or make them more vulnerable to chronic wound development.
In order to achieve this goal, I would try to make use of Oxford Nanopore sequencing, which is a third-generation sequencing technology. Since the main goal of this reading would be to read specifically mitochondrial DNA, the first step would be to extract the DNA from a wound tissue/normal tissue sample and quantify it. Next, I should perform long-amplicon PCR in order to highlight the mitochondrial DNA, subsequently adding an A tail so that sequenecing adapters can ligate efficiently. Following this, the sequencing adapters will be attached and and then sequencing will be started. In order to decode the bases of the DNA sample, Oxford Nanopore sequencing relies on ionic currents to detect which bases are passing through the nanopore. Since each basepair emits a different current value, we are able to trace the sequence that subsequently passes through the pore by decoding each current value. The final output of this sequencing technology is a FASTQ file that includes the DNA sequence along with a per-base quality score.
AI citation
Peplexity - “Can you explain in simple terms how Oxford Nanopore sequencing is prepared and what is the outcome?”
DNA Write
One of the long-term projects I’ve been working on at the Designer Cells lab was to synthesize a genetic circuit for chronic wounds. Since the main obstacle of chronic wound healing is their persistent inflammation, I designed a FLEx (Flip Excision) switch [12] that first expresses an anti-inflammatory gene set and then switches irrersibly to a migratory gene set after sensing a biomarker indicative of the end of the inflammation phase of wound healing.
Since this genetic circuit insert ended up being close to 4.5k, I think that the most efficient method to synthesize this would be an enzymatic synthesis approach. This synthesis method is similar to how primers are synthesized. Some limitations of this synthesis method is that it has limited synthesizing ability for longer strands of DNA.
DNA Edit
One point of DNA I’ve been looking at is to edit Calreticulin to add domains that recognize damage-associated extracellular matrix patterns. This would allow these engineered proteins to more effectively seek out damaged tissues (such as in the case fo chronic wounds) in order to facilitate wound healing.
In order to achieve this, I could utilize CRISPR-Cas HDR. In order to achieve this, I would need to design a gRNA that targets the desired insertion site as well as the edits that would need to be made. Then, I would need to ensure that the gRNA was delivered to the appropriate cells of my choosing. The potential limitations could revolve around structure integrity as well as the efficiency of HDR.