Week 2 HW: DNA Read, Write, & Edit
Part 1: Benchling & In-silico Gel Art
Simulate Restriction Enzyme Digestion
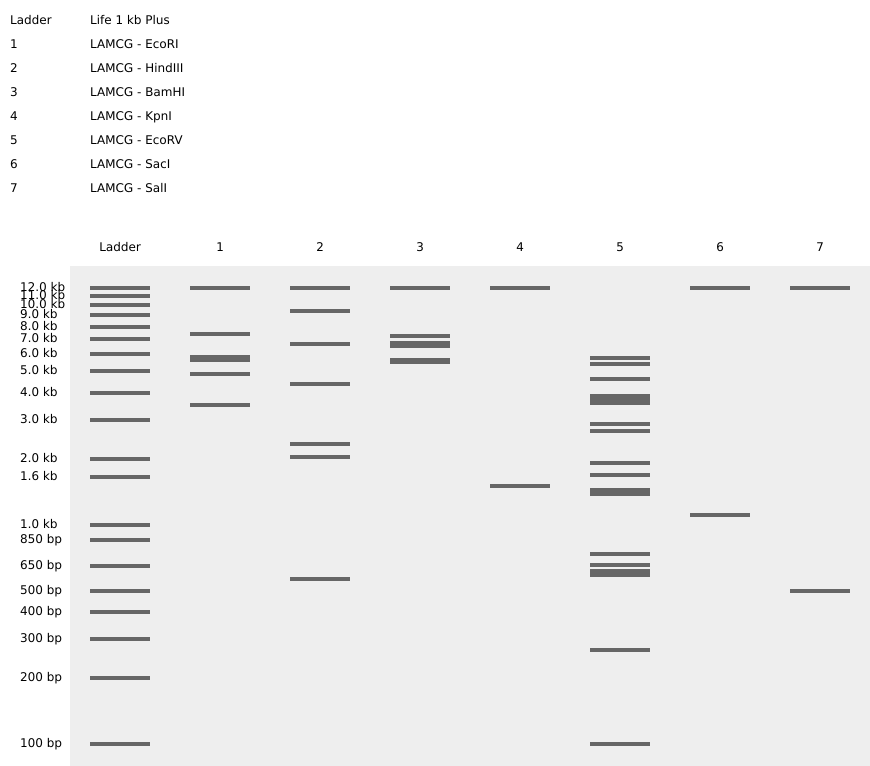


Image in the style of Paul Vanouse’s Latent Figure Protocol artworks
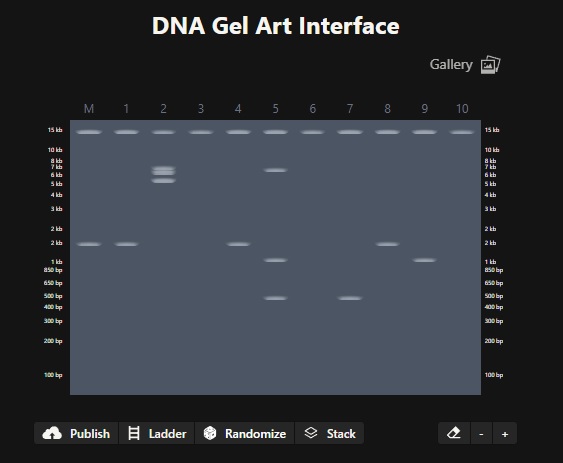

Part 3: DNA Design Challenge
3.1 For my homework, I chose the p53 protein because it plays a key role in regulating the cell cycle and protecting the body from tumor growth. This protein is often called the “guardian of the genome” because it is activated when DNA is damaged and can halt cell division or trigger apoptosis.
I found it interesting to study p53 because its mutations are associated with the development of many types of cancer. Therefore, it is important not only for a fundamental understanding of cellular molecular biology but also for medicine and the development of anticancer drugs.
MEEPQSDLSIELPLSQETFSDLWKLLPPNNVLSTLPSSDSIEELFLSENVTGWLEDSGGALQGVAAAAAS TAEDPVTETPAPVASAPATPWPLSSSVPSYKTFQGDYGFRLGFLHSGTAKSVTCTYSPSLNKLFCQLAKT CPVQLWVNSTPPPGTRVRAMAIYKKLQYMTEVVRRCPHHERSSEGDSLAPPQHLIRVEGNLHAEYLDDKQ TFRHSVVVPYEPPEVGSDCTTIHYNYMCNSSCMGGMNRRPILTIITLEDPSGNLLGRNSFEVRICACPGR DRRTEEKNFQKKGEPCPELPPKSAKRALPTNTSSSPPPKKKTLDGEYFTLKIRGHERFKMFQELNEALEL KDAQASKGSEDNGAHSSYLKSKKGQSASRLKKLMIKREGPDSD
to
3.2 ATGGAGGAGCCGCAGTCAGATCCTAGCGTCGAGCCCCCTCTGAGTCAGGAAACATTTTCAGACCTATGGAAACTACTTCCTGAAAACAACGTTCTGGTAAGGACAAGGGTGGTTGGGAGTAGATGGAGCCTGGTTGATCCTGAGGCTGGGCTCCGGGGACACTTTGCGTTCAGTGGGAGAGGAGCTGCTGGTG…
3.3 MTAMESSQSDISLELPLSQETFSGLWKLLPPEDILPSPHCMDDLLLPQDVEEFFEGPSEALRVSGLLQHQDPVTETPGPVAPAPATPWPLSSSVPSQKTYQGNYGFHLGFLQSGTAKSVMC
3.4 Once I have the DNA sequence of the TP53 gene, I can use it to produce the p53 protein. This can be done using either cell-dependent (in vivo) or cell-free expression system.
Cell-dependent protein expression (in vivo)
This is the most common method and involves expressing the gene inside living cells (such as E. coli, yeast, or mammalian cells).
Steps:
- Cloning The TP53 DNA sequence is inserted into an expression vector (plasmid). The plasmid contains:
promoter (e.g., T7 promoter),
ribosome binding site,
transcription terminator.
Transformation The plasmid is introduced into host cells.
Transcription RNA polymerase reads the DNA template and synthesizes messenger RNA (mRNA). During transcription, all thymine (T) bases are replaced with uracil (U).
Translation The ribosome reads the mRNA in groups of three nucleotides (codons). Transfer RNA (tRNA) molecules bring the corresponding amino acids. A polypeptide chain is formed, resulting in the p53 protein.
3.5 A single gene can produce multiple proteins at the transcriptional level through several regulatory mechanisms that generate different RNA transcripts from the same DNA sequence.
The most important mechanism is alternative splicing. After transcription, eukaryotic genes produce a precursor mRNA (pre-mRNA) containing both exons (coding regions) and introns (non-coding regions). During RNA processing, introns are removed and exons are joined together. However, the cell can combine exons in different ways. Some exons may be included or skipped, producing multiple mature mRNA variants from the same gene. Each mRNA variant is translated into a slightly different protein isoform.
Another mechanism is the use of alternative promoters. A gene may contain multiple promoter regions, allowing transcription to start at different sites. This can produce mRNAs with different 5′ ends, which may alter the resulting protein’s structure or regulation.
Additionally, alternative polyadenylation can generate transcripts with different 3′ ends, affecting mRNA stability, localization, or translation efficiency.
Through these mechanisms, one gene can generate multiple distinct mRNA transcripts, each encoding different protein variants. This greatly increases protein diversity without increasing the number of genes in the genome, and it is especially common in eukaryotic organisms.
ATG GAG GAG CCG CAG TCA GAT CCT AGC GTC GAG CCC CCT CTG AGT CAG GAA ACA TTT TCA GAC CTA TGG AAA CTA CTT CCT GAA AAC AAC GTT CTG GTA AGG ACA AGG GTG GTT GGG AGT AGA TGG AGC CTG GTT GAT CCT GAG GCT GGG CTC CGG GGA CAC TTT GCG TTC AGT GGG AGA GGA GCT GCT GGT G
Part 4: Prepare a Twist DNA Synthesis Order

Part 5: DNA Read/Write/Edit
5.1 If I were to choose DNA to sequence, I would focus on circulating tumor DNA (ctDNA) found in human blood. Circulating tumor DNA consists of small fragments of DNA released into the bloodstream by cancer cells. Sequencing this DNA allows researchers and clinicians to detect cancer-associated mutations without performing invasive tissue biopsies.
To sequence circulating tumor DNA (ctDNA), I would use a combination of Illumina Next-Generation Sequencing (NGS) and, when necessary, Oxford Nanopore sequencing.
Illumina sequencing is a second-generation sequencing technology. It is considered second-generation because it performs massively parallel sequencing of millions of short DNA fragments simultaneously after clonal amplification.
Oxford Nanopore is a third-generation sequencing technology, because it sequences single DNA molecules directly without PCR amplification and can generate very long reads.
The input is cell-free DNA (cfDNA) isolated from blood plasma.
Sample preparation steps:
Blood collection
Plasma separation (centrifugation)
DNA extraction
Fragment size selection (ctDNA is usually ~150–200 bp)
End repair and A-tailing
Adapter ligation (platform-specific adapters added)
PCR amplification (library enrichment)
Library quality control (quantification and fragment analysis)
Essential steps of Illumina sequencing (How bases are decoded) Illumina uses sequencing by synthesis (SBS):
Library fragments bind to a flow cell coated with complementary adapters.
Bridge amplification creates clusters of identical DNA copies.
Fluorescently labeled reversible terminator nucleotides are added.
Only one nucleotide incorporates per cycle.
A camera captures fluorescence to determine which base (A, T, C, or G) was added.
The terminator is chemically removed, and the cycle repeats.
Base calling occurs by detecting the fluorescent signal emitted by each incorporated nucleotide.
Millions of short sequence reads (typically 100–150 bp)
5.2 In a bioart context, I would choose to synthesize fluorescent protein genes, such as GFP (green fluorescent protein) or its variants. These genes can be inserted into harmless bacterial or yeast cells to create living artworks that glow under specific light.
The reason for choosing fluorescent proteins is twofold:
Aesthetic impact: Fluorescence provides an immediate visual effect, allowing the audience to see biological processes in real time. Artists can manipulate colors, patterns, or intensity to create dynamic, living installations.
Conceptual significance: By choosing a specific DNA sequence and controlling its expression, the artist highlights the interplay between genetic information, living matter, and human creativity. The synthesized DNA becomes a medium of artistic expression, illustrating how genetic code can be “written” and interpreted beyond scientific applications.
To synthesize a gene like GFP for a bioart project, I would use oligonucleotide synthesis combined with gene assembly and order it through commercial DNA synthesis services (e.g., Twist Bioscience, Integrated DNA Technologies).
This method allows you to “write” DNA sequences precisely, including codon optimization for the host organism (e.g., E. coli or yeast). You can control the exact sequence, introducing mutations or color variants for artistic effect.
5.3 If I could edit DNA, I would focus on enhancing stress resilience in crop plants, such as wheat or rice. The goal would be to make plants more resistant to drought, extreme temperatures, and disease, improving global food security in the context of climate change.
DNA Editing Technologies The most precise and widely used technology today is CRISPR-Cas9, a genome-editing tool. For crops, CRISPR allows targeted edits such as:
knocking out genes that make plants sensitive to stress,
inserting or modifying genes that confer drought tolerance, disease resistance, or improved nutrient use.
CRISPR is favored because it is efficient, precise, and relatively easy to program for specific DNA sequences.