Week 10 HW: Advanced Imaging & Measurement Technology
Homework: Final Project
Fluorescent Output (GFP Expression)
Intensity of fluorescence (proxy for stress-response activation. Microfluidic imaging Important:
- Direct readout of stress pathway activation
- Allows spatial mapping across salinity gradient
Gene Expression Levels (mRNA)
Real-time PCR systems. Important:
- Confirms that fluorescence reflects transcriptional activity
DNA Sequence Verification
DNA sequencing Important:
- Ensures mutations and reporter constructs are correct
Protein Expression and Stability
Gel electrophoresis Important:
- Confirms protein is produced and not degraded
Ion Transport Activity
Ion-sensitive fluorescent dyes Important:
- Direct indicator of osmotic stress response
Cell Morphology (Frustule Structure)
Microscopy imaging over time Important:
- Links genetic response to physical adaptation
Homework: Waters Part I — Molecular Weight
- Theoretical pI/Mw: 5.90 / 27875.41
2.1. m/zn = 903.7148; m/zn+1 = 933.7349 z= 933.7349/(903.7148 - 933.7349); z= -31.1 - take a module - z= 31
2.2. MW = z × (m/z) − z × H;
MW = 31 × 903.7148 − 31;
MW = 27984 Da
2.3. Theoretical molecular weight eGFP: 27875.41 Da
Accuracy = |MWexp − MWtheory| / MWtheory;
Accuracy = |27984 − 27875.41| / 27875.41;
Accuracy = 0.0039;
0.39 %
- No. To determine the state of charge (z) in LC-MS, several peaks are needed, such as (z), (z+1), (z+2)
Homework: Waters Part II — Secondary/Tertiary structure
1.
In its native state, a protein like eGFP is properly folded into a compact 3D structure stabilized by noncovalent interactions (hydrogen bonds, hydrophobic interactions, ionic interactions). This folded conformation is what gives the protein its biological function.
In the denatured state, the protein is unfolded. These stabilizing interactions are disrupted (e.g., by organic solvent, acid, heat), causing the structure to lose its compact shape and become extended and flexible. Importantly, the primary structure (amino acid sequence) remains intact—only the higher-order structure is lost.
Proteins are typically analyzed using electrospray ionization (ESI), which produces multiply charged ions.
2.
The peak at ~2800 m/z has a +10 charge state, determined from the ~0.1 m/z spacing between isotopic peaks (Δm/z = 1/z)
Homework: Waters Part III — Peptide Mapping - primary structure
1.
| peptide sequence |
|---|
| HNIEDGSVQLADHYQQNTPI GDGPVLLPDNHYLSTQSALS K |
| DHMVLLEFVTAAGITLGMDE LYK |
| GEELFTGVVPILVELDGDVN GHK |
| LPVPWPTLVTTLTYGVQCFS R |
| LEYNYNSHNVYIMADK |
| FSVSGEGEGDATYGK |
| SAMPEGYVQER |
| LEHHHHHH |
| FEGDTLVNR |
| EDGNILGHK |
| QHDFFK |
| YPDHMK |
| FICTTGK |
| DDGNYK |
| TIFFK |
| DPNEK |
| GIDFK |
| VNFK |
| IELK |
K = 15 R = 3
2.
19 peptides
3.
22 chromatographic peaks
4.
No. There are 3 more peaks in the chromatogram.
5.
The isotope spacing is= 1/z; the peaks are spaced ~0.5 m/z apart (525.76712, 526.25918), therefore z = +2.
M=(m/z×z)−z×1.007
M=(525.767×2)−(2×1.007)
M≈1051.534−2.014=1049.52 Da
[M+H]⁺= = M+1.0073 = 1,049.51964 + 1.0073 = 1,050.52694 Da
6.
Accuracy = |MWexp − MWtheory| / MWtheory;
Accuracy = |1,050.52694 − 1050.5214| / 1050.5214;
Accuracy = 5.27ppm
7.
The percentage of the sequence that is confirmed by peptide mapping is 88%
Homework: Waters Part IV — Oligomers
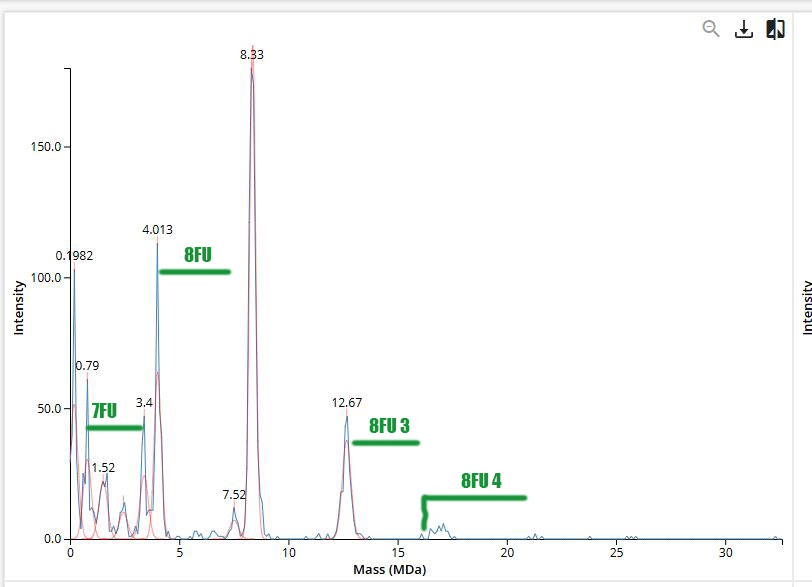

Homework: Waters Part V — Did I make GFP?
| – | Theoretical | Observed/measured on the Intact LC-MS | PPM Mass Error |
|---|---|---|---|
| Molecular weight (kDa) | 28.01 | 27.98 | 906 ppm |