<Tehseen Rubbab > — HTGAA Spring 2026
About me
Contact info
Homework
- Week 1 HW: Principles and Practices
- Homework 2 - DNA READ,WRITE,& EDIT
- Homework 4 – Protein Design I
- Week 3 Homework
Week 1 HW: Principles and Practices
I am curious about exploring the potential of HTGAA utilising the Design Build Test and Learn strategy. I have recently done an Innovate UK project in which I used MoClo kits to design the synthesis of xanthone in yeast. I am curious about exploring the potential of the phenylpropanoid pathway and the plethora of useful chemicals that could be produced.
Homework 2 - DNA READ,WRITE,& EDIT
Part 1: Benchling & In-silico Gel Art Simulate Restriction Enzyme Digestion Part 3: DNA Design Challenge 3.1. Choose your protein Retrieved Germin like protein sequence located on chromosome 8, important in plants biotic and abiotic stresses.
Part A. Conceptual Questions 1. How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons) From 500 g of meat, you ingest approximately 1 mole of protein, which equals 6 × 10²³ molecules according to Avogadro’s constant.
You can view/download my presentation here: Week 3 Homework PPT

I am curious about exploring the potential of HTGAA utilising the Design Build Test and Learn strategy. I have recently done an Innovate UK project in which I used MoClo kits to design the synthesis of xanthone in yeast. I am curious about exploring the potential of the phenylpropanoid pathway and the plethora of useful chemicals that could be produced.
Hence, I am proposing this study “Engineered Naringenin production through Design Build Test and Learn Strategy”, as a committed listener, for which I would be interested to work in Victoria,Genspace or Designer Cells.I am currently based in UK and hence intereted in working on nodes in UK.
The study involving the metabolic engineering of Naringenin will be conducted under approved BSL-1 laboratory conditions, in accordance with institutional and biosafety guidelines. For in vitro cell cultures, commercially available human-derived cell lines will be used. No primary human subjects will be directly involved. All data collection, analysis and reporting will adhere to the principles of ethical research conduct.
Naringenin (C15H12O5) is a novel bioactive compound within the flavonoid group, found naturally in the skin of citrus fruits. This citrus flavonoid is well known for its anti-inflammatory, antioxidant, and anticarcinogenic properties, and can support skin against the detrimental impacts of inflammaging. Outside skin care research, Naringenin has shown promising results in treating issues like obesity and cardiovascular diseases.
The primary source of Naringenin is citrus fruits, hence consuming a lot of natural resources and labour. What if we make the process economical and produce naringenin at lab scale? Sounds great and doable through synthetic biology.
I propose engineering the phenylpropanoid pathway in S. cerevisiae to produce naringenin, utilizing the Design Build Test and Learn (DBTL) strategy of MoClo kits.
The proposed study holds the potential to revolutionise the cosmetic industry by providing an economical source of Naringenin without relying on natural resources, hence a step towards preventing the depletion of natural resources.
The study requires step-by-step, careful pathway optimisation through omics approaches to identify any bottlenecks in the engineering strategy in a heterologous host. If not followed accurately, the pathway will not lead to Naringenin production.
With attention to detail and proper record keeping, the project holds the potential to revolutionize the cosmetic industry.
| Consideration | Option 1 | Option 2 | Option 3 |
|---|---|---|---|
| Enhance Biosecurity | |||
| By preventing incidents | ✓ | ||
| By helping respond | ✓ | ||
| Foster Lab Safety | |||
| By preventing incidents | ✓ | ||
| By helping respond | ✓ | ||
| Protect the Environment | |||
| By preventing incidents | ✓ | ||
| By helping respond | |||
| Other Considerations | |||
| Minimizing costs and burdens to stakeholders | ✓ | ||
| Feasibility | ✓ | ||
| Not impede research | ✓ | ||
| Promote constructive applications | ✓ |
I will propose constructive applications of the proposed research in the healthcare and beauty sectors, and not impede research and researchers. The governance action to be addressed should focus on the rights of the worker/researcher. As many projects are led by industry, researchers must not be treated as paid labour but are provided fair recognition, intellectual property rights, and equal participation in patent ownership.
I am extremely thankful to be a part of HTGAA-2026, looking forward to learning from world leading researchers in the field of synthetic biology.
Kind Regards,
Tehseen
Simulate Restriction Enzyme Digestion
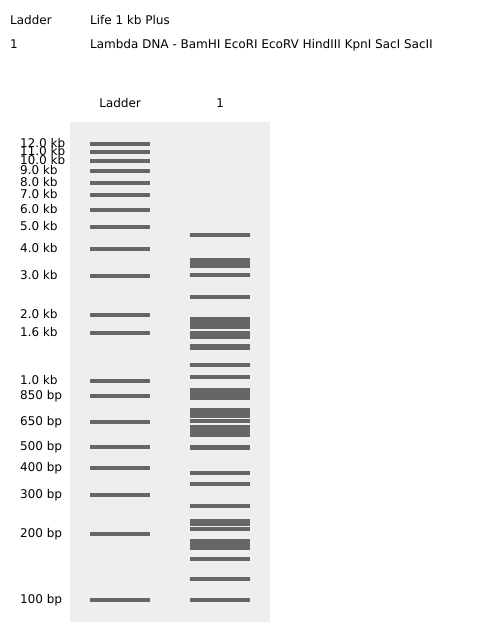

Retrieved Germin like protein sequence located on chromosome 8, important in plants biotic and abiotic stresses.
Done using IDT tool as Twist Bioscience link provided was not working, optimised for expression in S. cerevisiae, removed type II restriction enzymes BsaI, BsmBI and BpsI.
The sequence entered of OsRGLP1 having start codon and a stop codon:
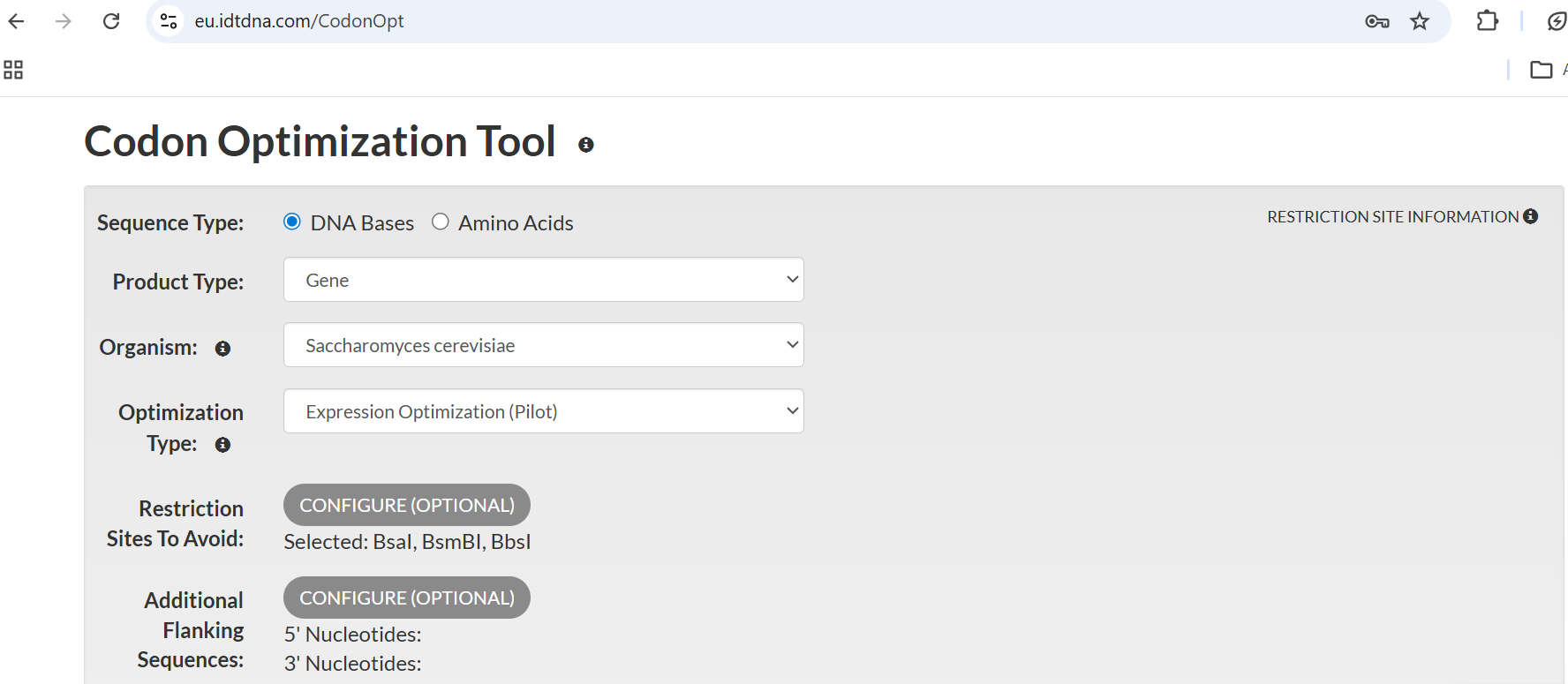

These are the parameters used on the IDT codon optimisation tool; restriction sites to avoid were entered. However, when checking the sequence on Benchling for the absence of these enzymes, BsmBI and BsaI are not present, but one restriction site of BbsI does exist. The codon optimisation might have been more accurate if Twist or GenWiz software had been used.
The gene sequence can be recombinantly expressed in S. cerevisiae by using a protocol adapted from Gietz, R. D et al., 2007.
Added 5’ and 3’ sequences for Type II restriction digestion and have annotated the sequence 5’ end - CDS - 3’ end. Can be subsequently used for ordering through Twist Bioscience.
Used vectors of YTK kit and used the combinatorial assembly feature of Benchling to design the final vector: pYtk009 vector (pTDH3) promoter, pYTK051 vector (tENO1) terminator.
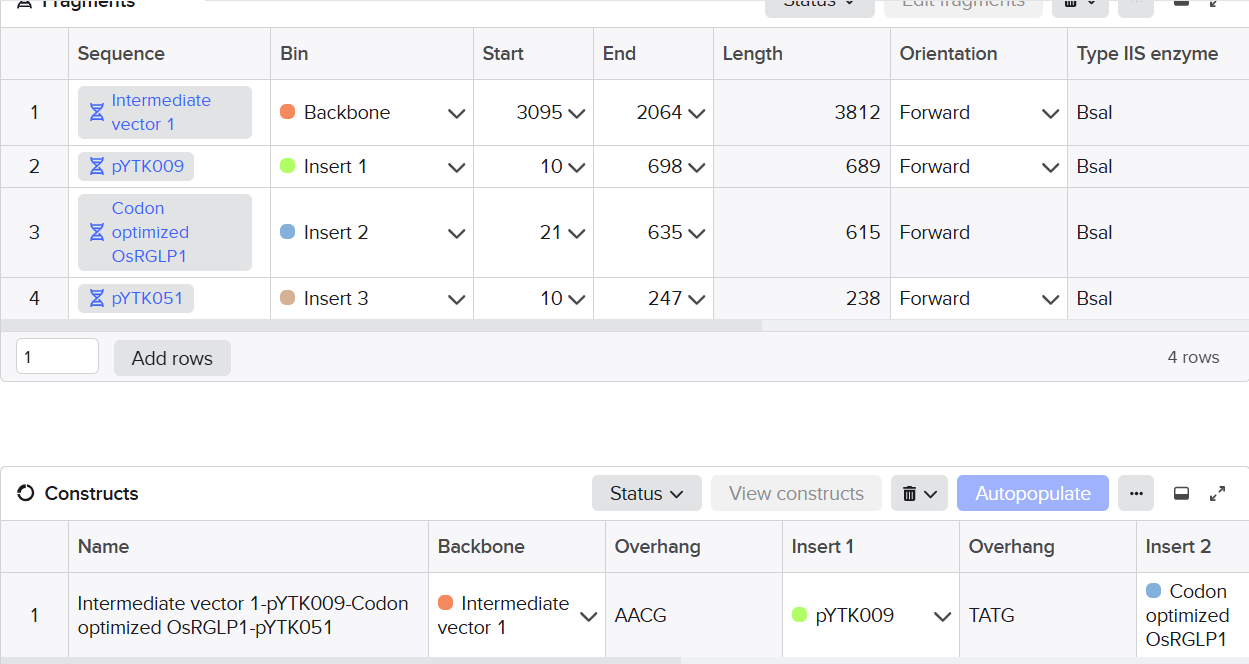

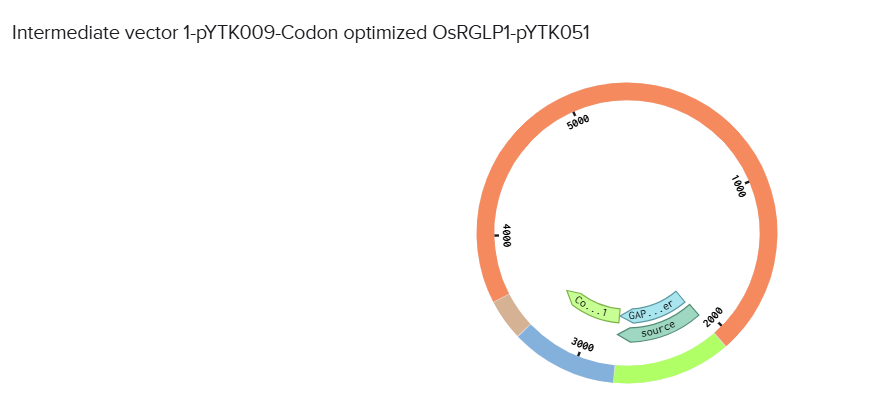

(on average an amino acid is ~100 Daltons)
From 500 g of meat, you ingest approximately 1 mole of protein, which equals 6 × 10²³ molecules according to Avogadro’s constant.
Humans do not become cows or fish because food is broken down during digestion into biomolecules such as carbohydrates, fats, proteins, and simple sugars. These molecules are then reassembled into human tissues according to our own genetic code.
The genetic code naturally fits 20 amino acids while providing enough functional diversity to encode proteins. Proteins are encoded by triplet codons composed of four nucleotides:
4³ = 64 codons
61 codons encode amino acids and 3 codons function as stop signals.
Yes, synthetic amino acids can be designed by maintaining the standard backbone:
NH₂–CH–COOH
while modifying the R side chain to provide new chemical properties.
Examples:
Fluorinated amino acids
Long hydrophobic amino acids
Redox-active amino acids
Before enzymes and life existed, amino acids likely formed through prebiotic chemical reactions on early Earth and in space.
Simple molecules such as:
could react under energy sources such as:
This concept was demonstrated in the Miller–Urey experiment (1953). Additionally, amino acids have been detected in meteorites, suggesting that space chemistry may have contributed to the origin of biomolecules used by early life.
An α-helix composed entirely of D-amino acids would form a left-handed helix, which is the mirror image of the right-handed helices formed by L-amino acids in natural proteins.
Proteins contain several types of helices besides the α-helix, including:
Most natural helices are right-handed because proteins are composed primarily of L-amino acids. The stereochemistry of L-amino acids makes the right-handed helix sterically and energetically favourable.
β-sheets aggregate because their backbone hydrogen bonds and side-chain interactions favour tight stacking, which can lead to structures such as amyloid fibrils.
Driving forces of β-sheet aggregation:
These forces stabilise the aggregated state.
Germin and germin-like proteins are ubiquitous plant proteins expressed during various biotic and abiotic stresses.
The OsRGLP1 protein has confirmed superoxide dismutase activity, although additional activities such as oxalate oxidase remain under investigation.
Sequence length: 224 amino acids
Most frequent amino acid: Alanine (A) — 23 occurrences
A: 23 L: 21 P: 18 V: 18 N: 17 S: 15 F: 15 G: 13 T: 12 D: 12 I: 12 K: 11 Q: 9 M: 6 H: 6 Y: 4 E: 4 W: 3 R: 3 C: 2
BLAST search parameters:
OsRGLP1 belongs to the germin-like protein (GLP) family. Germins share 30–70% sequence similarity with wheat germins. Protein sequence homologues
Reference:
Zhang, Y. et al. (2018). Overexpression of germin-like protein GmGLP10 enhances resistance to Sclerotinia sclerotiorum in transgenic tobacco. Biochemical and Biophysical Research Communications.
• Does your protein belong to any structure classification family Germin and GLPs are described as archetypal members of the cupin superfamily (Dunwell 1998). The cupin superfamily of proteins is named based on this conserved B barrel fold (Cupa; Latin word for barrel), discovered using a conserved motif found in germins.
Part C. Using ML-Based Protein Design Tools In this section, we will learn about the capabilities of modern protein AI models and test some of them in your chosen protein.
ii) Latent Space Analysis Scan
iii) Inverse fold modelling
iv. The heat map was generated in C1-protein protein language modelling, specifically from a deep mutational scan using the ESM-2 protein language model, which predicts the probability of every amino acid mutation at each position in the protein sequence
You can view/download my presentation here: Week 3 Homework PPT