Week 2 HW: Read, Write, and Edit DNA
Part 1: Benchling & In-silico Gel Art
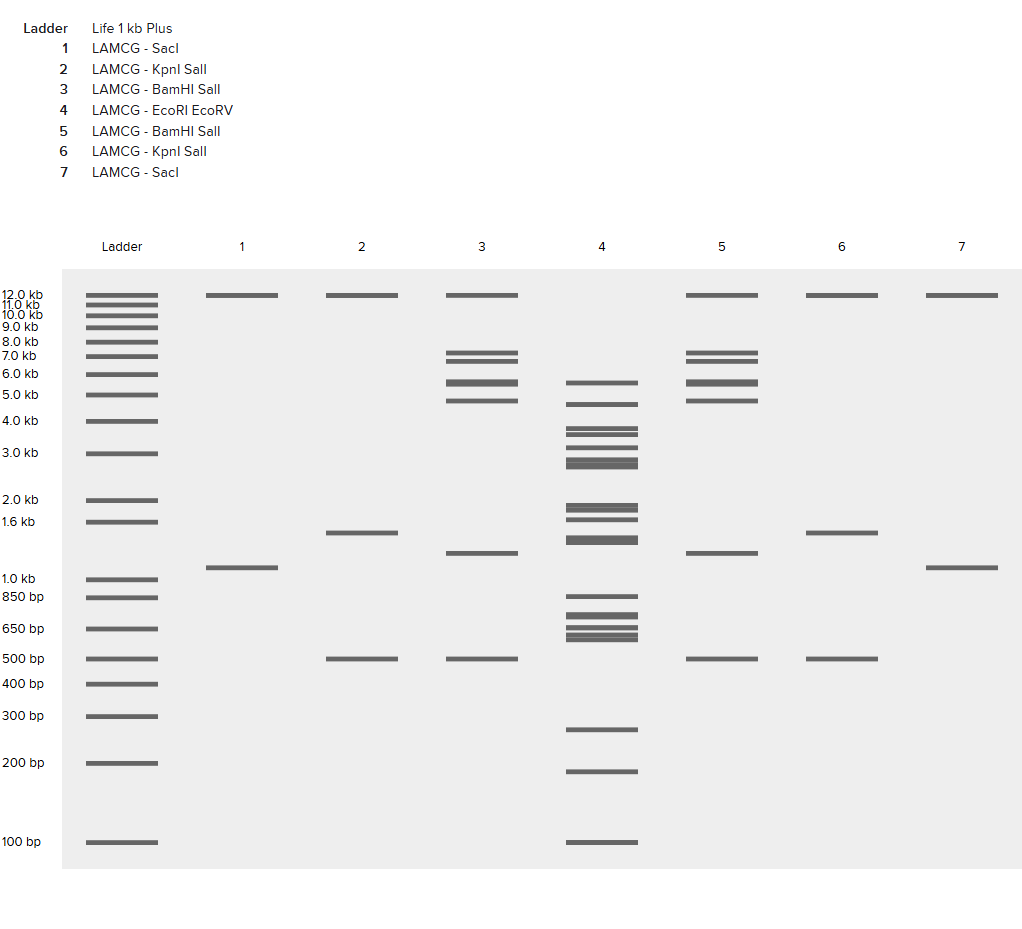

Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
See Week 2 Lab.
Part 3: DNA Design Challenge
3.1. Choose your protein.
- Skeletal aspartic acid-rich protein 1 Acropora millepora (Staghorn coral)
-sp|B3EWY6|SAAR1_ACRMI Skeletal aspartic acid-rich protein 1 OS=Acropora millepora OX=45264 PE=1 SV=1 MAFVSCFHLRLLFLCLALFMAAECRPDELNKKVDSDETISDDDVSARVQPNGGKIMIVRD NDYDASDDNDNDNDDDDNNDNDNDNDDDNDVDRDNDNDDDDFDDSNDDMLSFELDSIEEK DSDGNDVGSTEGHSVESFEDRPFSLSSVDRNSNALGVAAINVNLSTKLEDSNADVDIMLY LFREDGTISFGNETFDVQAGTVKFNIKISNWDFCDGSAQDCSEAKAGEYLDVNIKFKSKD TPIEVTDEERKSQNKPAVCKDKDTPDTDSDPDDSSDNANDGDDDDDDDCPHIYNMGGDSE MLLNRGVMNGDTYTAMPFGFPKVEIEDGEKKIKFRVPKFDDNVNIDPSVTPGRVPKNASP SPALCLKIHILFIALLQAVTLFINSW
- I chose skeletal aspartic acid rich protein 1 (SAARP1) because it is directly involved in coral calcification and biomineralization, linking a specific protein to the physical formation of the coral skeleton. Skeletal aspartic acid rich protein 1 (SAARP1) is a major component of the skeletal organic matrix (SOM) in the staghorn coral Acropora millepora. It is a highly acidic protein with roughly 20 percent aspartic acid residues and is involved in coral calcification and biomineralization. SAARP1 belongs to a conserved family of acidic proteins that likely helps regulate the formation, structure, and deposition of aragonite crystals in the coral skeleton.
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
- NCBI Reference Sequence: XM_029346345.2
- acaccggaaa ccagagattt gcagcagaaa tgtgacataa ggctgctttc cgagctagca cttatcagcg tccattcgcc atttggcagc attgtttatg gccgctgaat gtcgcccaga cgagctgaac aaaaaagtgg acagtgatga aacaattagc gatgatgacg tctccgcaag agttcagccc aatggcggga aaatcatgat cgtccgcgac aatgactacg atgcctccga cgataatgat aatgatgatg atgacaataa cgataacgat aacgataacg atgatgataa tgatgtcgat cgtgataatg ataatgatga cgacgatttt gacgacagta acgacgatat gctctcgttt gagttggata gcatagaaga gaaggactcg gatggaaatg acgtagggag cactgaagga cattctgtag aatcatttga agatcgacca ttttctctgt cttccgttga tcgcaacagc aatgctttag gtgtcgcggc catcaatgtc aacctgtcaa cgaaacttga ggattctaac gcagacgtag acatcatgtt gtacctgttt cgcgaagatg gaaccatttc ctttggtaac gaaacatttg atgtacaagc tggaactgtt aagtttaata ttaagattag caactgggat ttctgtgatg gttcagcgca agactgcagc gaggcaaaag caggcgaata ccttgatgta aacatcaaat tcaagagtaa agacacacca atcgaagtaa cagacgagga acgaaaaagt cagaataaac cggcagtgtg caaggacaaa gacacaccag atactgacag cgatccagat gatagcagtg ataatgccaa tgatggggat gacgatgacg atgacgactg tcctcacatc tataacatgg gtggagattc agagatgtta ctcaacagag gggtaatgaa tggtgacacc tacactgcta tgccgtttgg attccctaaa gtcgaaattg aggatggaga gaagaaaata aagtttcgcg ttcccaagtt tgacgataac gtaattatag accctagcgt gaccccagga agagtgccaa agaatgcttc gccttcacct gccctttgcc tgaaaattca catcctcttc attgcactac ttcaagctgt taccctattt atcaacagtt ggtaacactc aaggggtttt aaacattact atgtggaatt tgaggcgaaa ataagcgaag gcaatacttg cttcgattcc tctttgcact gaattgccat ctgtaatttt gacagagaat agcatcgtta acaccacgtg ttaaagtata caccatagtc tcgagcaatt gtcgtaacag gagataacaa agttatcaac gaaacgttta cactcaggaa tgacatcagt attagtttca gttttgagaa tgaacagacg cgctctccat ggaaaccatg gtgttgttag ggtttattta aattttaaat aaagtaatgc taaccaccaa cctacctacc tacctacatt cgttttacaa ttcagaatga gttttctttt taccatggct caacctccat gcgccaccat tgctaggctt gctacacaca tcgcgcgttt attcagtatc tttcttatta ccgtatatgg tgtttcacca aaatagttga agatcgaagg ccactatcga agattaagtt gatattatgc aaaacaggtc attcactaat taagagtctt gtttgatgtg aatccctacc gcaagctaaa actaatcttg tcatgaactt ttaggcaatg tattgctgaa gattggagaa agcagcgtag ggattatgct ccaaatgtca cctttgcaat tttcttacgg ttgtaattta accctgttat gtttaatatc aaattttgtt atccaattaa attaattgaa acactgcc
3.3. Codon optimization.
- Even though multiple codons can encode the same amino acid, different organisms prefer different codons. That preference tracks with tRNA abundance and other expression constraints. If you use a DNA sequence with many codons that are “rare” in your expression host, translation can stall, causing lower yield, more misfolding/aggregation, and sometimes premature termination. The codon is optimized for E. Coli because we will be using it in labs as the host.
3.4. You have a sequence! Now what?
- Once I have a codon-optimized SAARP1 DNA sequence, I can produce the protein by placing the gene into an E. coli expression plasmid that includes a promoter, ribosome binding site (RBS), transcription terminator, origin of replication, and an antibiotic resistance marker, often plus a purification tag such as 6×His. The plasmid can be introduced into E. coli either via heat shock or electroporation. After the plasmid is inside E. coli, the gene is transcribed into mRNA when the promoter is active (often induced in expression systems). The mRNA is then translated by E. coli ribosomes, which read the mRNA codons and use tRNAs to assemble the SAARP1 amino-acid chain. Codon optimization improves this step by matching E. coli’s preferred codons, reducing ribosome stalling and increasing the likelihood of higher protein yield.
Part 4: Prepare a Twist DNA Synthesis Order
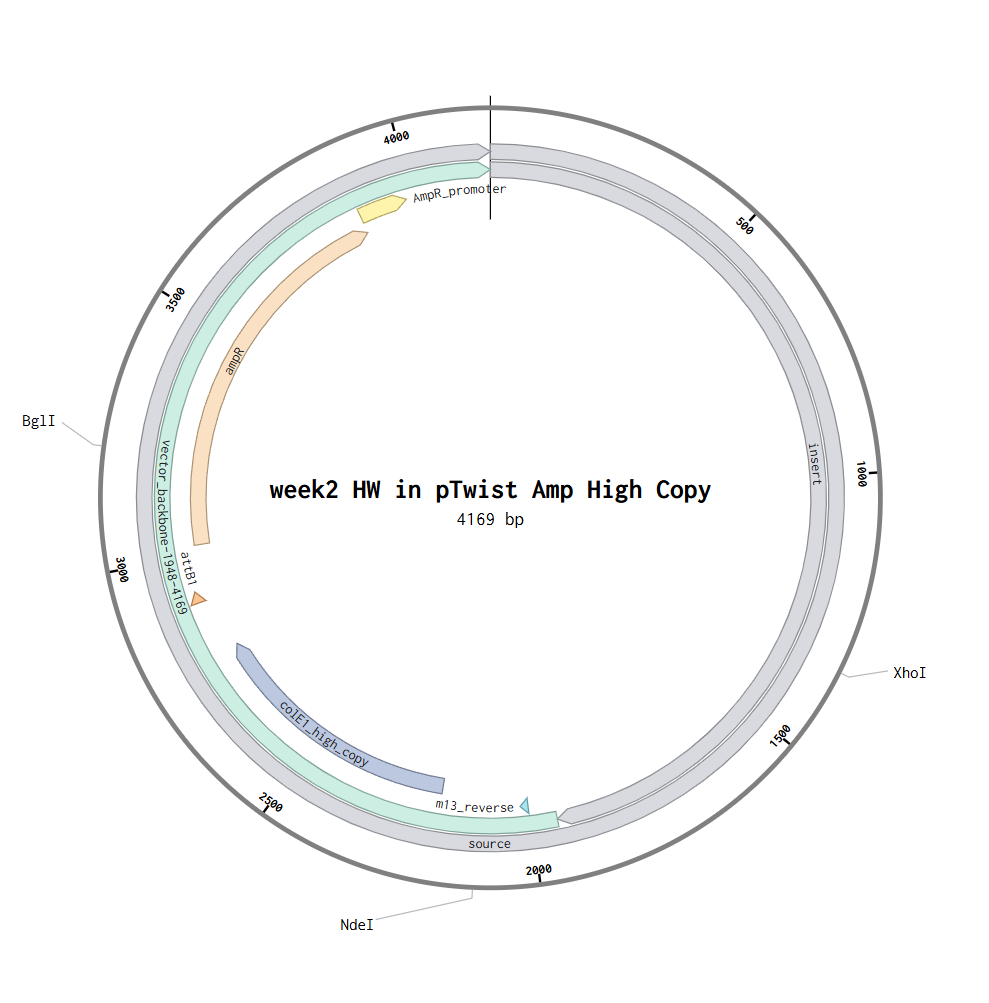

Part 5: DNA Read/Write/Edit
5.1 DNA Read (sequence)
- (i) What DNA would you want to sequence and why?
- I would sequence some of coral’s DNA like the SAARP1 (skeletal aspartic acid-rich protein 1) gene from Acropora millepora, because I want to explore coral’s potential as architectural material or learn about their calcification performance and stress response under warming and acidification to ensure green future shoreline development.
- (ii) What sequencing technology would you use and why?
- I would use Sanger sequencing to read the SAARP1 gene because it is accurate, cost-effective, and ideal for sequencing individual gene-length fragments amplified by PCR. PCR and synthetic oligos allow me to isolate the specific gene region, and Sanger sequencing provides precise base-level confirmation of the amplified or edited DNA. Sanger sequencing is considered first-generation sequencing, because it reads one DNA fragment at a time with high accuracy. The input is genomic DNA extracted from coral tissue. The essential preparation steps are: design oligo primers → PCR amplify the SAARP1 locus → purify the PCR product → add sequencing primers → load into the Sanger reaction. Sanger sequencing uses chain-terminating nucleotides labeled with fluorescent dyes. When DNA synthesis stops at each base position, the fragments are separated by size and read by a detector that converts fluorescence into a DNA sequence. The output is a chromatogram and a high-accuracy DNA sequence of the amplified SAARP1 fragment, typically in FASTA format.
5.2 DNA Write (synthesize)
- (i) What DNA would you want to synthesize and why?
- I would synthesize a codon-optimized SAARP1 coding sequence so I can express it in a chosen host (or produce peptides/biomineralization assays) and test how SAARP1 chemistry affects mineral deposition and coral-like calcification cues.
- (ii) What synthesis technology would you use and why?
- I would use phosphoramidite oligo synthesis plus assembly (the standard commercial approach) because it is fast, scalable, and ideal for making gene-length constructs via assembly from shorter oligos.
- Design sequence in silico → synthesize short oligos → assemble into the full gene (PCR/Gibson-like assembly) → clone into plasmid → sequence-verify.
- Long sequences accumulate errors and often need verification and rebuilding. Repetitive/low-complexity regions and extreme GC can reduce synthesis success. Scaling to many variants is feasible, but costs and validation time increase quickly.
5.3 DNA Edit (edit)
- (i) What DNA would you want to edit and why?
- I would edit the SAARP1 regulatory region (or specific coding residues) in coral to test whether higher or better-timed SAARP1 expression improves calcification under heat/acidification stress.
- (ii) What editing technology would you use and why?
- I would use CRISPR-based editing, ideally prime editing (or a high-fidelity Cas9 + HDR if prime editing isn’t feasible), because it can make targeted changes without needing large insertions.
- 1) How does it edit DNA, and what are the essential steps?
- CRISPR targets a specific locus with a guide RNA and edits it using an editor enzyme and a repair template or edit-encoding RNA. Key steps are guide design → delivery into cells/embryos → editing reaction → screening/validation.
- 2) What prep and inputs are needed?
- Inputs include a target sequence, guide RNA, editor protein (Cas9/prime editor), and (if needed) a repair template plus the recipient cells/embryos and delivery method (microinjection/electroporation/viral vectors depending on system).
- 3) Limitations (efficiency/precision)?
- Delivery and survival in coral embryos can be difficult and can cause mosaic edits. Off-target edits and unintended repair outcomes can occur. Even precise edits may have ecological and ethical constraints if used outside controlled research settings.