Week 2 HW: DNA Read,Write and Edit
Part 1: Benchling and In-silico Gel Art
I followed the instructions and made a free account using benchling.com - a website used to design experiments, manage biological data, and collaborate on research where in our case for the purpose of this homework we will be using it edit DNA plasmid structures with different restriction enzyme digests.
The next step was to import Lambda DNA into the website as a new project, this DNA would act as the template for which we would perform restriction site digests with different enzymes.
I was a little confused as to why the link for Lambda DNA redirected me to an ordering page and I tried searching through it to see if it had a download link but couldnt find anything so I then directed my search to google and found it on the following page:
https://www.ncbi.nlm.nih.gov/nuccore/NC_001416.1?
Next I uploaded the .fasta file I was able to downloaded as a linear topology, in metadata there was an error to identify what schema was being used which was confusing as I uploaded itas a DNA sequence already.
Ignoring that error, I moved ontto simulation of restriction Enzyme Digestion for the following enzymes: EcoRI HindIII BamHI KpnI EcoRV SacI SalI
After performing the digest with all the above enzymes I recieved the following results:


And then to add a bit of an artistic approach to the gel lanes i tried to make a mirror image stair case which ended up looking like this:
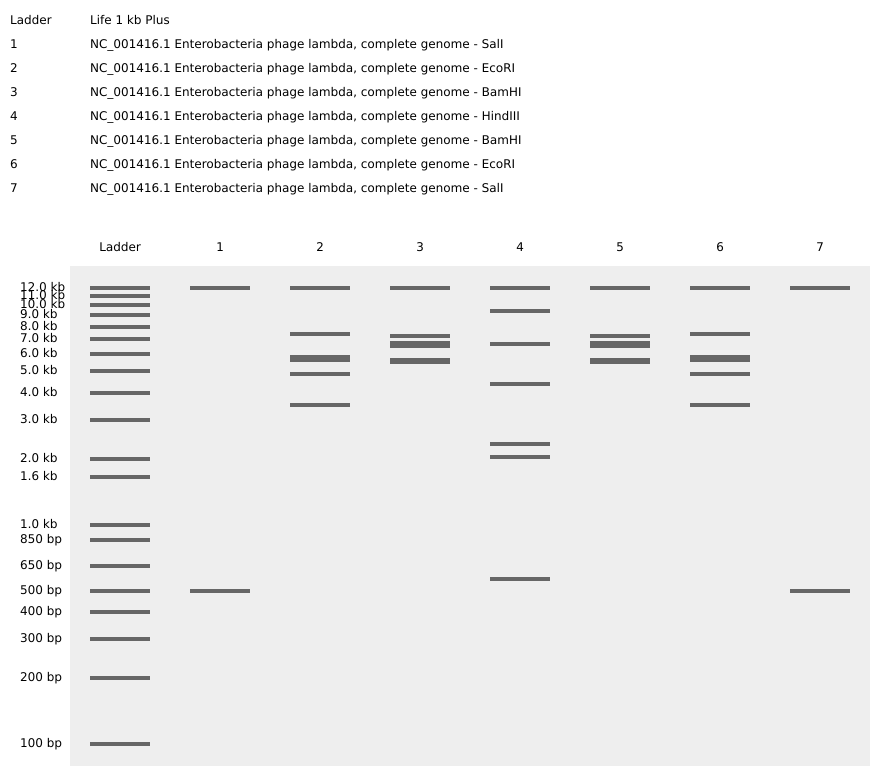


PART 3: DNA Design Challenge
Protein chosen: Retatrutide. An emerging game-changer in obesity pharmacotherapy, retatrutide represents the forefront of a broader shift toward the normalised use of peptide-based therapeutics, not only for treating metabolic diseases but also for expanding applications in cosmetic and lifestyle medicine.
Biological assembly of Retratrutide Structure:
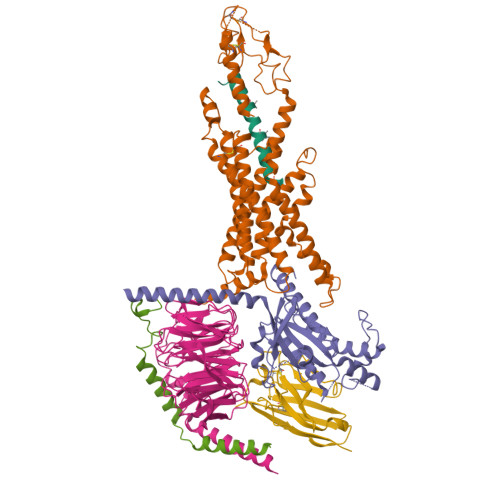

As retatrutide is a synthetic peptide drug rather than a naturally occurring protein, it is not listed as a canonical entry in UniProt. Therefore, we instead reference one of the endogenous hormones it is derived from, GLP-1.
GLP-1 amino acid sequence:
HAEGTFTSDVSSYLEGQAAKEFIAWLVKGRG
Using a reverse translation the following possible DNA sequence is obtained:
CATGCTGAAGGTACCTTTACTTCTGATGTCTCTTCTTATCTGGAAGGTCAAGCTGCTAAAGAATTTATTGCTTGGCTGGTCAAAGGTCGTGGT
Codon optimisation is necessary because the genetic code is degenerate, meaning that most amino acids can be encoded by multiple different codons. Although these codons specify the same amino acid, they are not used equally across all organisms. Different species show codon bias, where certain synonymous codons are preferred over others. This preference is largely linked to the availability of corresponding tRNAs in the host organism, which can strongly influence the efficiency and accuracy of translation.
In this project, codon optimisation was used to adapt the DNA sequence so that it matches the codon usage preferences of the chosen expression host. This helps to improve protein expression by reducing the likelihood of slow or inefficient translation.
In our case, E. coli was selected as the host organism because of its rapid growth rate, ease of transformation, low cost, and widespread use in recombinant protein expression. These features make it a practical and efficient system for producing the target peptide.
And so following codon optimization suited for E. coli the following is obtained:
CATGCGGAAGGTACCTTTACCTCTGATGTGTCTTCTTATCTGGAAGGTCAAGCGGCGAAAGAATTTATTGCGTGGCTGGTGAAAGGTCGTGGT