week-06-hw-genetic-circuits-part-i
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix contains several key components that enable accurate DNA amplification. It includes Phusion DNA polymerase, a high-fidelity enzyme with proofreading activity that reduces errors during DNA synthesis. The mix also contains dNTPs, which serve as the building blocks for new DNA strand formation. An optimized reaction buffer provides the correct pH and ionic conditions, including magnesium ions (Mg²⁺), which are essential cofactors for polymerase activity. Additionally, stabilizers and enhancers are included to improve enzyme performance and allow efficient amplification of difficult templates such as GC-rich regions. Some versions also contain a tracking dye, enabling direct loading of PCR products onto a gel for analysis.
2. What are some factors that determine primer annealing temperature during PCR?
Several factors determine the optimal primer annealing temperature in PCR, as it must allow specific binding without nonspecific interactions. The most important factor is the melting temperature of the primers, which depends on their length and nucleotide composition. Primers with a higher GC content have higher Tm values because G–C base pairs form three hydrogen bonds, making them more stable than A–T pairs. Primer length also influences Tm, with longer primers generally requiring higher annealing temperatures. The sequence composition matters as well, since secondary structures like hairpins or primer dimers can affect binding efficiency. Additionally, the salt concentration and buffer conditions in the reaction can influence primer-template interactions and stability. Typically, the annealing temperature is set a few degrees (about 3–5°C) below the primer Tm to ensure specific and efficient binding.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Though they both fragment of DNA, PCR creates a ver large scale volume of repliucants of the exact same linear fragment and when it does this its joining fresh nucleotides together forming new DNA, whereas with restriction enzyme digests they are acting as molecular scissors and are break a pre existing DNA molecule into seperate fragments at specific restriction sites.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
Gibson Assembly is a cloning method that joins DNA fragments with overlapping ends in a single isothermal reaction. It uses an exonuclease to create single-stranded overhangs, allowing complementary fragments to anneal. And so the important part is to ensure that the adjacent fragments contain matcvhing overlapping end sequences, usually around 20-40 base pairs long - these overlaps must be designed so that each fragment can anneal specifically to the next fragment or to the linearized vector.
5. How does the plasmid DNA enter the E. coli cells during transformation?
Plasmid DNA enters E. coli cells during transformation by first making the cells competent, meaning their membranes are temporarily permeable to DNA. In chemical transformation, cells are treated with calcium chloride, which helps neutralize the negative charges on both the DNA and the cell membrane. A brief heat shock (e.g., 42 °C) then creates a thermal imbalance that allows the plasmid DNA to pass through the membrane and into the cell.
6. Describe another assembly method in detail (such as Golden Gate Assembly)
Golden Gate Assembly is another molecular cloning method that allows multiple DNA fragments to be joined together in single reaciton using special Type IIS estriction enzymes (such as BsaI) and DNA ligase,. The Type IIS restriction eznymes cut DNA outside their recognition sequence and leave a specific custome designed sticky ends that are unique for each fragment and so this allows the DNA fragments to be assembled in a specific order and orientation.
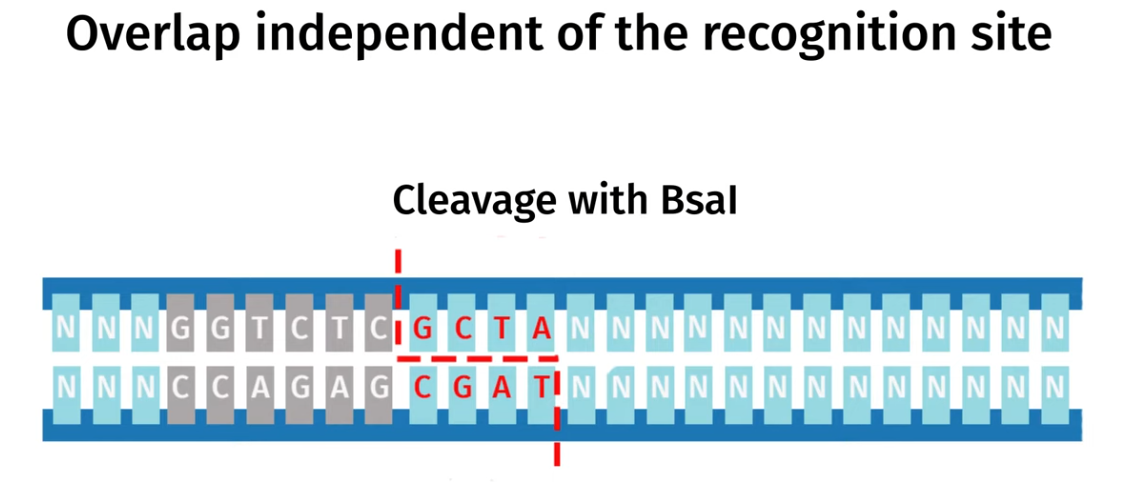
 DNA ligase simultaeneuosuly joins these fragments together as they are being cut and once the fragments are correctly assembled the recognition sites are removed so the final construct doesnt get cut again.
DNA ligase simultaeneuosuly joins these fragments together as they are being cut and once the fragments are correctly assembled the recognition sites are removed so the final construct doesnt get cut again.
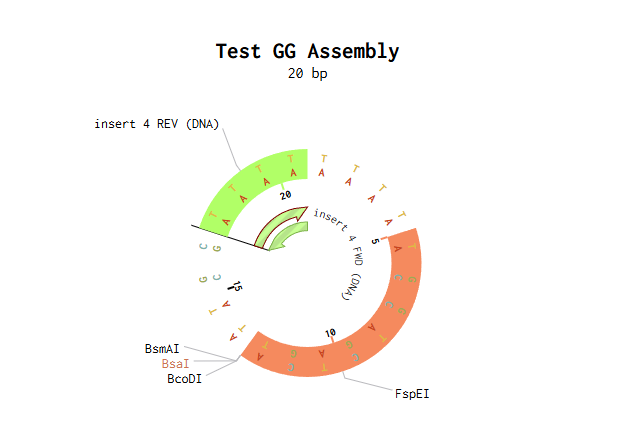
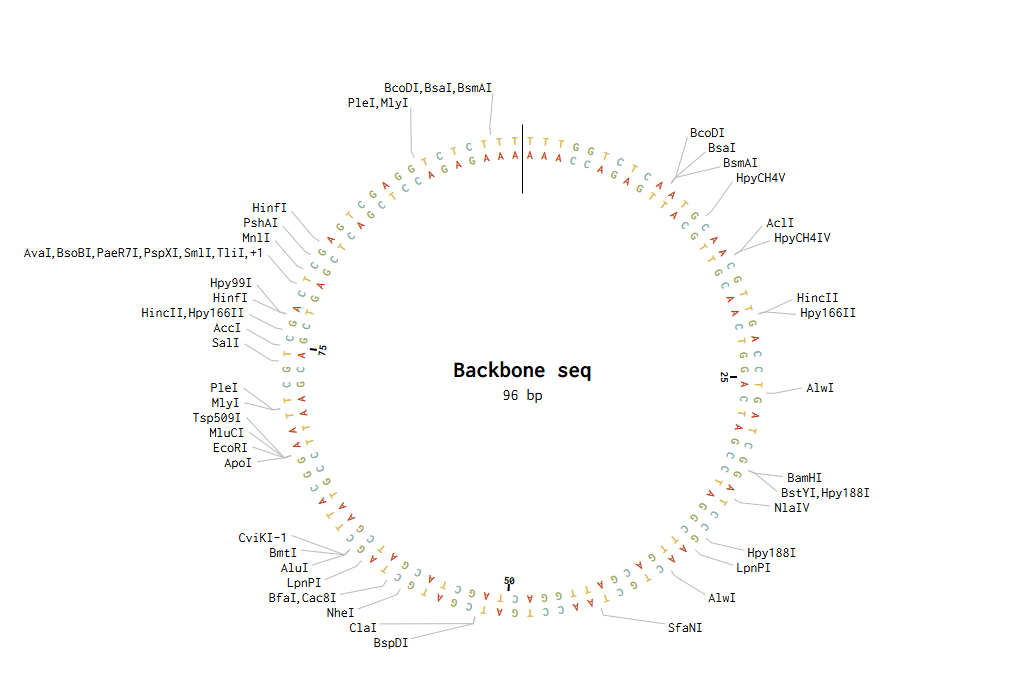
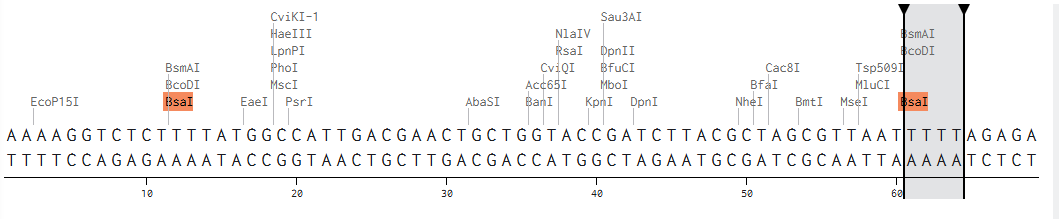
I modelled Golden Gate Assembly in Benchling by creating a test backbone and insert sequences and adding restriction sites for a Type IIS enzyme, BsaI. Unlike normal restriction enzymes, BsaI cuts outside of its recognition site, which allows custom overhangs to be generated. I first entered the DNA sequences into Benchling, making the backbone circular and the insert linear. I then checked the restriction sites and used the assembly tool to simulate how the fragments would join. At first, the assembly did not work because some of the sticky ends did not match, so the fragment sequences had to be corrected and reset. After adjusting the design so that the overhangs were compatible, the assembly worked and Benchling produced a final construct map. This showed how Golden Gate Assembly can join DNA fragments in a specific order and orientation.
Backbone and insert fragment structures:
And this was the final product after running the Golden Gate Assembly simulation via benchling: