Week 02 HW: DNA Read, Write, & Edit


Table of Contents
0. Basics of Gel Electrophoresis
Gel electrophoresis is a fascinating process that allows DNA fragments to be separated according to size. The migration of fragments through agarose reveals invisible molecular differences as visible banding patterns. What interests me most is how information encoded in DNA becomes a spatial structure that can be interpreted visually.
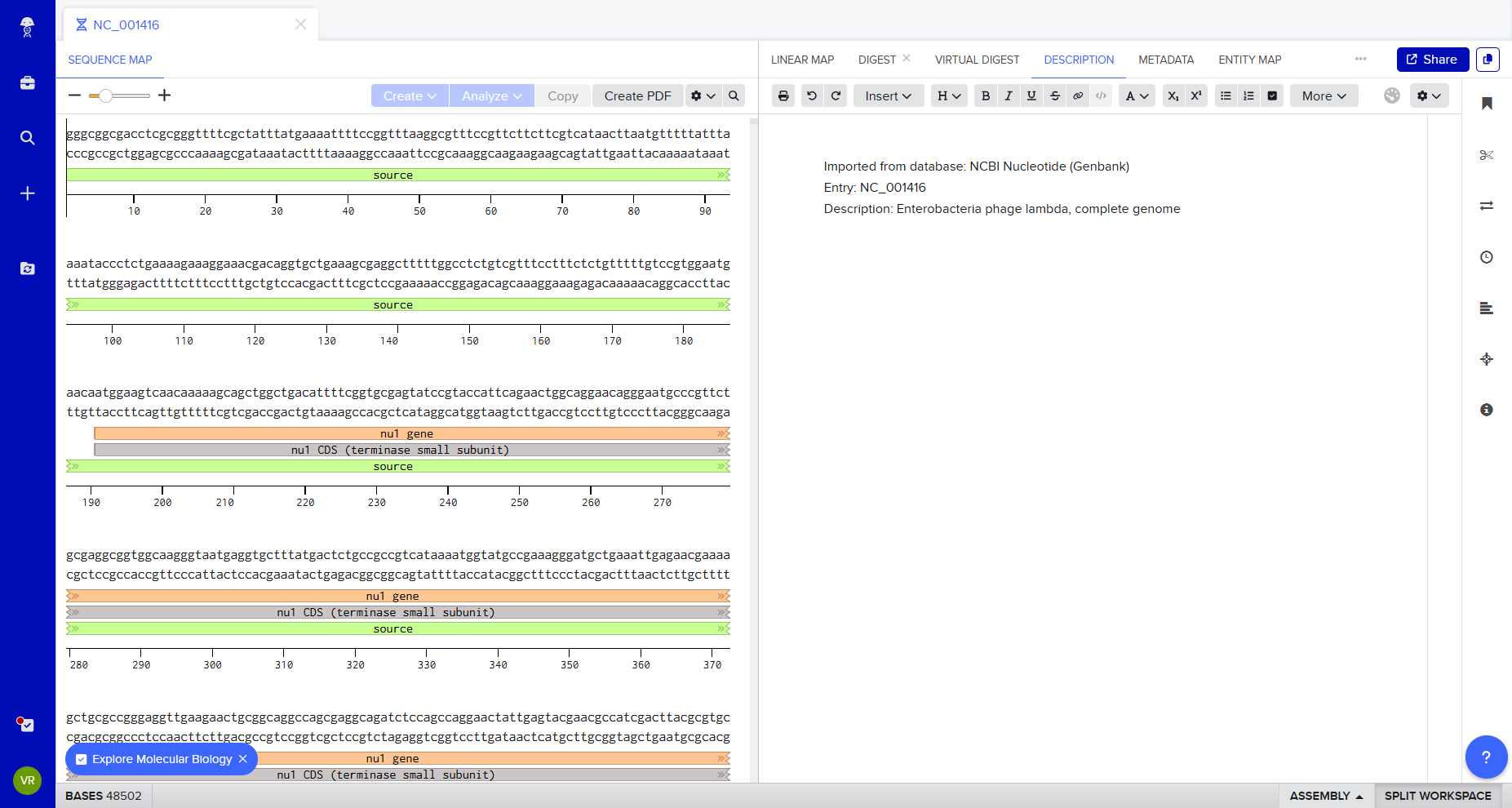
1. Benchling and In-silico Gel Art
Following the protocol Gel Art: Restriction Digests and Gel Electrophoresis, I explored Benchling to simulate restriction enzyme digestion.
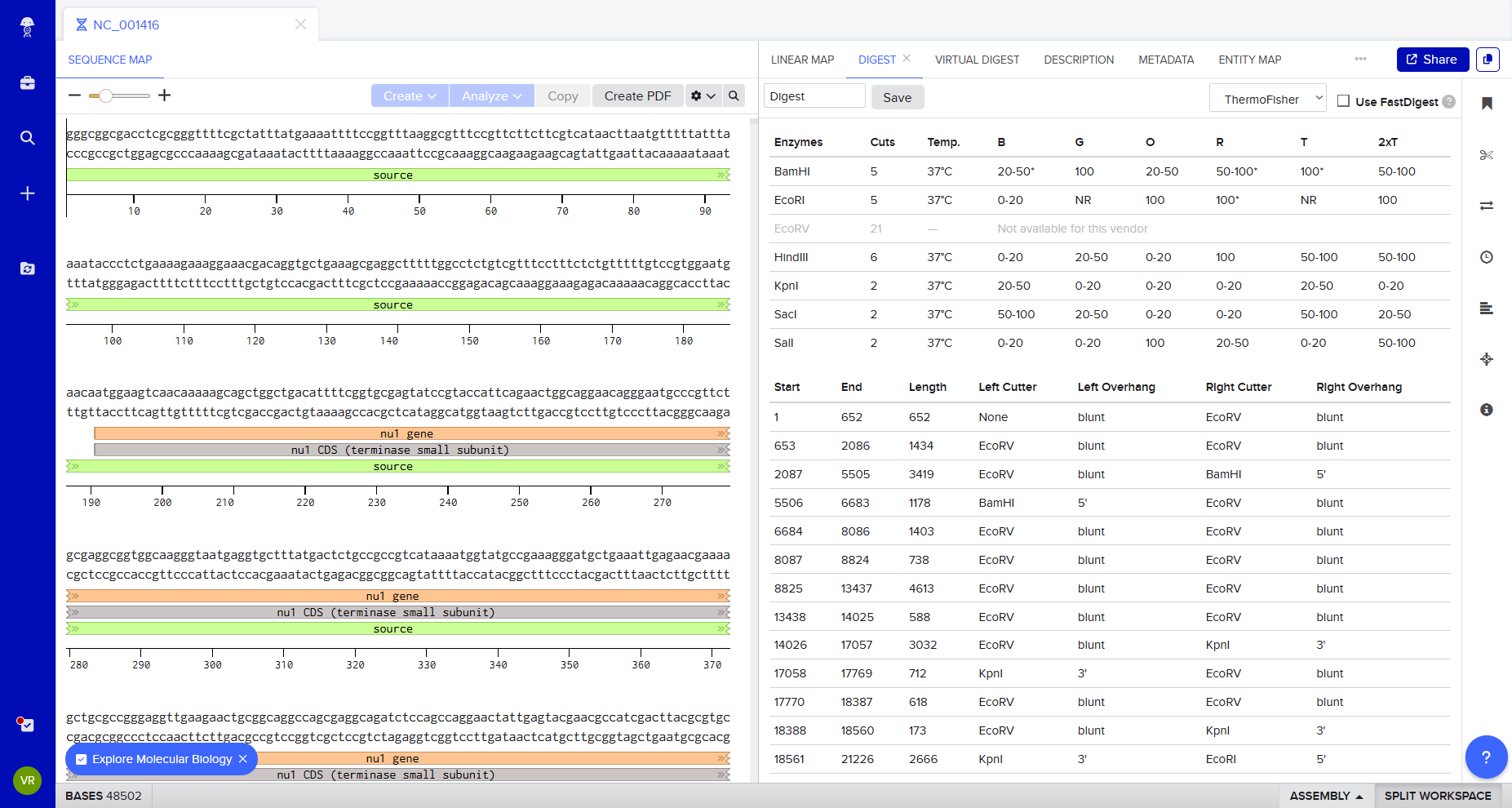
I imported Lambda DNA and simulated restriction digests using:
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
At first, I was not entirely sure what I was doing — I explored the platform experimentally, clicking through the interface to understand how the enzymes cut and how the DNA fragments were segmented.
How it could/ should looks on gel.
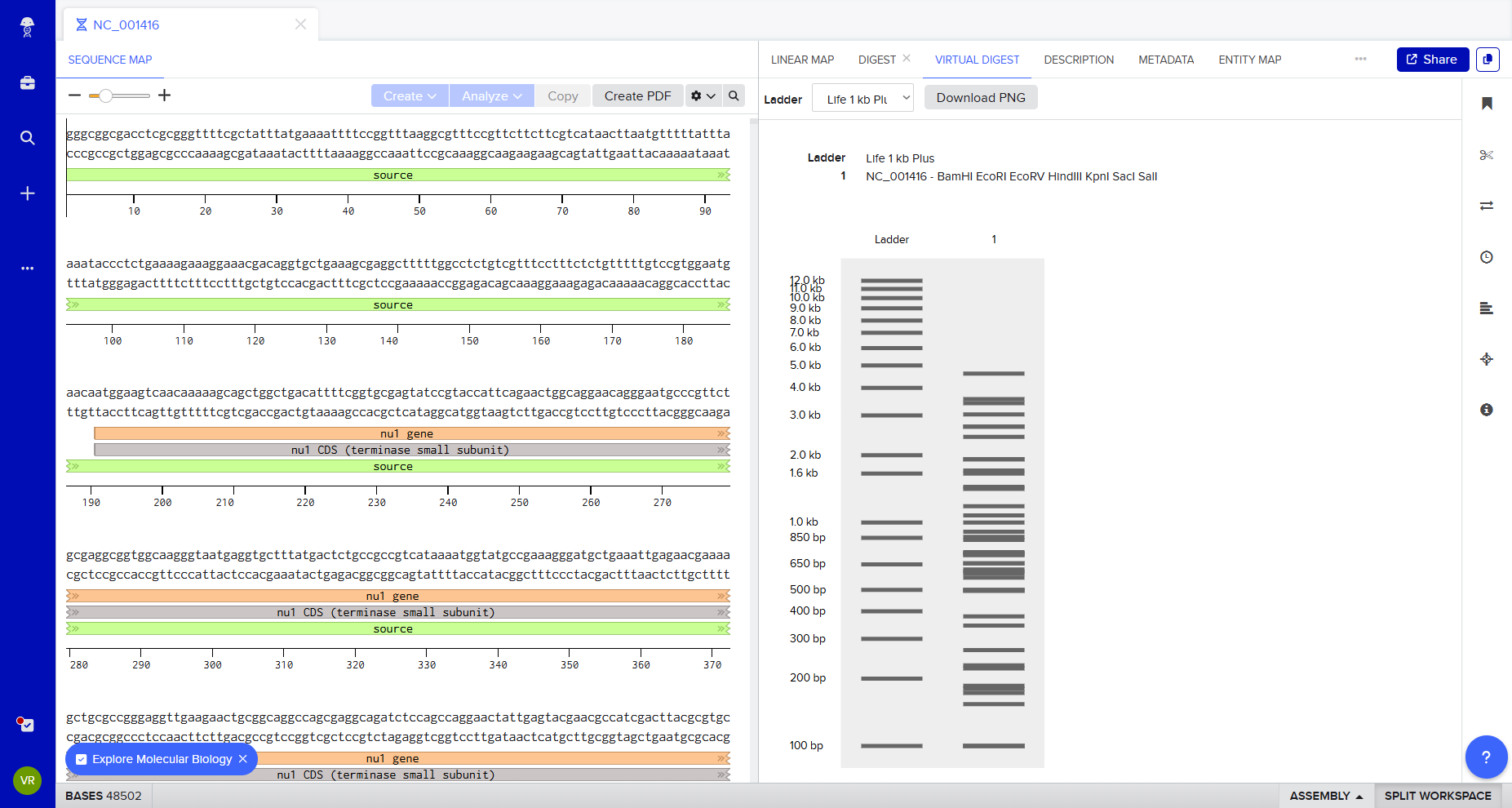

After generating fragment sizes, I moved to Ronan Donovan’s gel art tool:
👉 https://rcdonovan.com/gel-art
There, I entered fragment sizes from Benchling to simulate gel patterns. Interestingly, some of the same enzymes were already available in the tool. I experimented with fragment combinations, spacing, and contrast to create structured visual patterns. I tried composing a geometric “invader”-like form inspired by Paul Vanouse’s aesthetic approach.
This exercise made clear how restriction logic can generate latent images embedded in genomic structure.
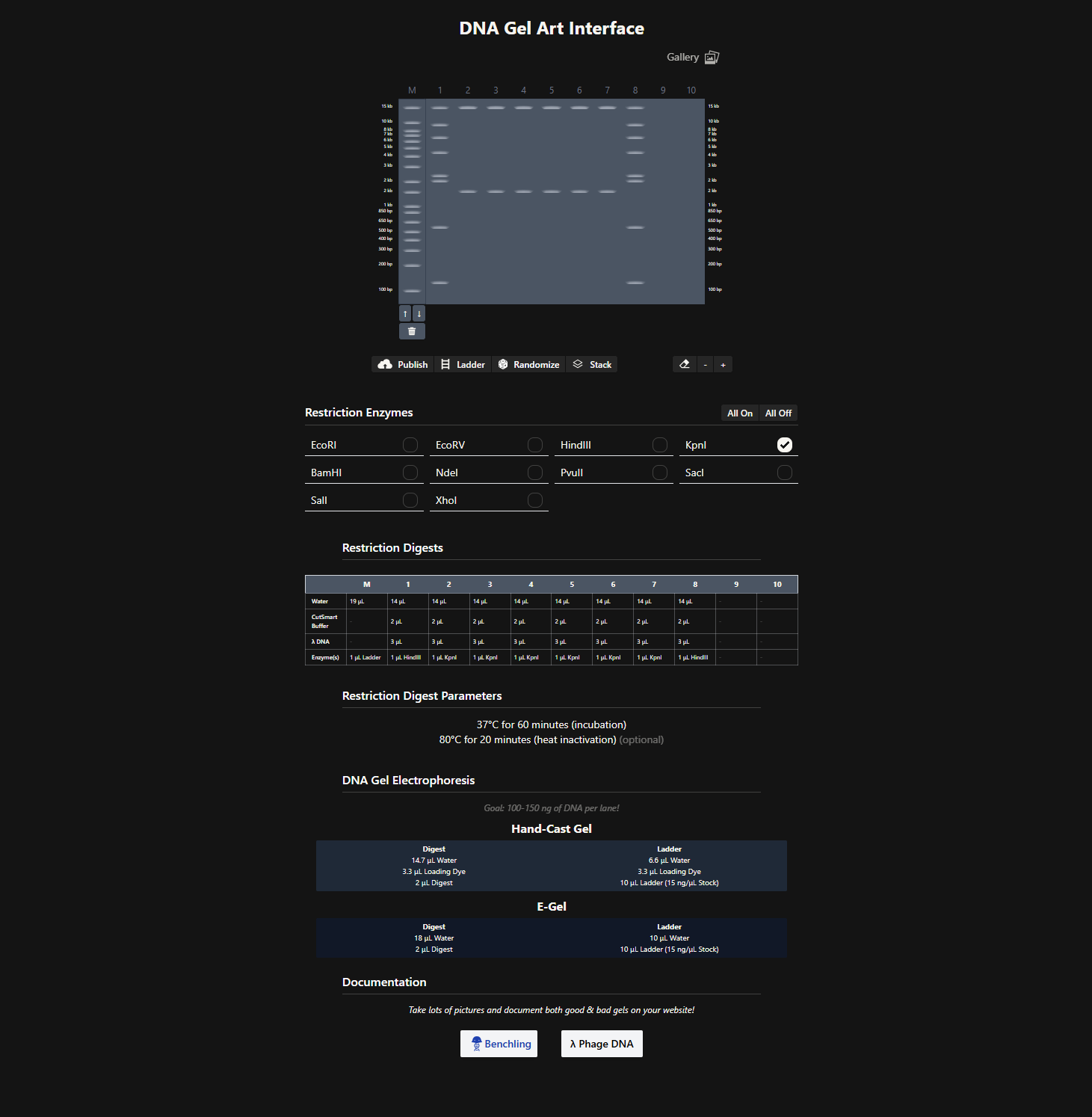

2. Gel Art Restriction Digests and Electrophoresis
This tutorial was particularly helpful:
👉 https://www.youtube.com/watch?v=TIZRGt3YAug
Although I do not currently have access to a laboratory to run physical gels, I have previously seen Paul Vanouse’s Latent Figure Protocol artworks in person. A curator friend owns one and showed it to me during a bioart conference. Seeing the material presence of gel-based imagery deeply informed how I approached this in-silico exercise.
3.1 Choose Your Protein
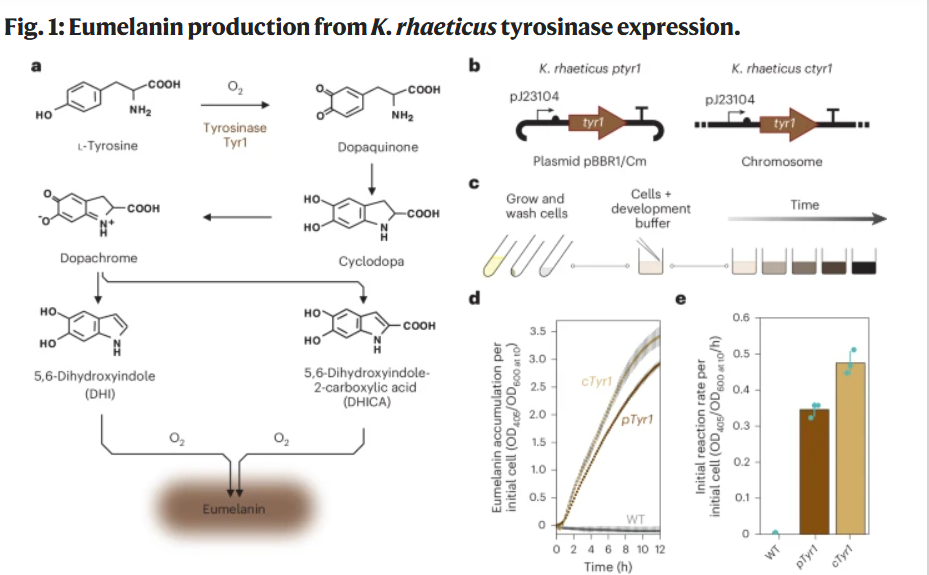

Protein chosen: Tyrosinase (Tyr1) from Bacillus megaterium
I chose the tyrosinase Tyr1 enzyme from Bacillus megaterium because it directly enables the biosynthesis of eumelanin, a redox-active polymer with electronic and material properties.
This protein is particularly relevant to my project, which explores the biological functionalization of bacterial cellulose. By expressing Tyr1 in Komagataeibacter rhaeticus, the cellulose-producing bacterium can generate eumelanin within or around the extracellular matrix, thereby modifying the optical and potentially electroactive properties of the material.
Tyr1 can encode a macroscopic material property in the cell of Komagateibacter Rhaeticus.
This strategy has been experimentally validated in:
Walker et al., 2024 — Self-pigmenting textiles grown from cellulose-producing bacteria with engineered tyrosinase expression The authors engineered K. rhaeticus strains expressing Tyr1 and demonstrated eumelanin production under buffered neutral pH conditions (Supplementary Data 1, 41587_2024_2194_MOESM1_ESM).
The paper is here 👉 https://www.nature.com/articles/s41587-024-02194-3 Details are in Supplementary information / Supplementary information / Supplementary Figs. 1–5, Tables 1–3 and Supplementary Data 1 and 2.
Or here : 📄 Supplementary Walker et al
The principale idea :
DNA (tyr1) ↓ transcription mRNA ↓ translation Tyrosinase enzyme ↓ catalytic activity Melanin polymer ↓ integration into cellulose matrix Modified impedance ↓ Human interaction (touch sensing)
Rather than maximizing conductivity, the goal is to explore how living metabolism can reorganize electrochemical properties within a self-grown material matrix. Maybe a hybrid approach combining enzymatic melanin production with controlled ionic modulation of the growth medium may provide a balance between biological emergence and measurable electrical response.
Impedance vs Conduction (for BC-based “electronic” biofilms)
Quick intuition
Conduction (DC): current flows through a continuous conductive path (like a wire).
→ You usually need a percolating network (CNT/graphene/metal paths).Impedance (AC): the material reacts to an alternating signal.
→ It mixes resistance + capacitance + ionic diffusion, and is extremely sensitive to: hydration, ions, porosity, and microstructure.
My goal is a tactile / capacitive surface then impedance is usually the most realistic target.
Concept map (what you measure defines what you build)
flowchart TB
A[BC biofilm / pellicle]--> B{What do I measure}
B-->|DC or very low frequency| C[Conduction]
B-->|AC multi-frequency| D[Impedance]
C-->C1[Electronic transport]
C-->C2[Needs a percolating path]
C2-->C3[CNT graphene Ag nanowires PEDOT]
D-->D1[Resistance R]
D-->D2[Capacitance C double layer]
D-->D3[Ionic diffusion]
D-->D4[Strongly affected by water ions porosity]
D4-->D5[Good for touch pressure hydration sensing]Protein Sequence (from Supplementary Data 1)
In the app bencling ==> https://benchling.com/s/seq-cSFfevSwjHFxf6KwwXGx?m=slm-D8VDordW2oyuqdI6bH4r
Structural Micro-Analysis: Copper Coordination and Catalysis
Tyrosinases are type-3 copper enzymes. Their catalytic activity depends on two copper-binding sites (CuA and CuB), each coordinated by conserved histidine residues. These histidine-rich motifs enable:
- Binding of molecular oxygen
- Oxidation of L-tyrosine to L-DOPA
- Conversion to dopaquinone
- Spontaneous polymerization into eumelanin
The presence of conserved histidines in the Tyr1 sequence reflects this copper-dependent catalytic architecture. The structure of the enzyme directly determines its ability to generate a redox-active polymer. Here, protein folding and metal coordination translate genetic code into material transformation.
Why This Protein Matters
1️⃣DNA → enzyme → material property The tyr1 gene encodes an enzyme. The enzyme catalyzes a redox reaction. The reaction produces a polymer. The polymer modifies material properties.
Thus: Genetic sequence → enzyme structure → catalytic activity → polymer formation → macroscopic material pigmentation and electroactivity This directly embodies the theme of “Reading & Writing Life.”
2️⃣ Minimal synthetic biology intervention Unlike multi-gene systems (e.g., the curli operon), Tyr1: Requires only one coding sequence Does not modify the cellulose biosynthesis operon (bcsABCD) Adds a new functional layer to the extracellular matrix This makes it an elegant example of additive biological functionalization, where a single gene expands the material phenotype of a living system.
3️⃣ Connection to biofabrication and living materials
Walker et al. demonstrate patterned melanin production using optogenetic control of tyr1 expression. This shows that: Gene expression can spatially control material coloration Biological programming can encode textile-level patterning Material properties can be written through genetic regulation This aligns directly with my research interest in growing functionalized 3D artifacts and interactive living interfaces.
Beyond color: material programming
In this project, the goal is not simply to encode color in DNA. The visible pigmentation is only a surface effect. What is actually encoded is a redox-active polymer production pathway inside living matter.
Melanin modifies:
- local redox properties,
- hydration-dependent conductivity,
- and potentially the impedance behavior of the cellulose matrix.
So pigmentation here is not aesthetic encoding, but electrochemical encoding. The gene does not just change appearance — it changes how the material behaves electrically. In that sense, DNA is not only encoding a protein, but indirectly programming a material state.
3.2 Reverse Translate
Protein (amino acid) sequence → DNA (nucleotide) sequence
Because of codon degeneracy, there is not a single unique DNA sequence for a given protein sequence. Many different nucleotide sequences can encode the exact same amino acid sequence. To reverse-translate Tyr1, I used a reverse-translation approach (equivalent to common online reverse-translation tools) by selecting one plausible codon per amino acid (a “standard/high-frequency codon” strategy).
The protein sequence used is Tyr1 from Bacillus megaterium as reported in Walker et al. 2024 (Supplementary Data 1). :contentReference[oaicite:1]{index=1}
Reverse-translated nucleotide sequence (one valid coding DNA sequence)
Degeneracy and probabilistic encoding
During this process, I realized that reverse translation is not deterministic. The genetic code is degenerate: multiple DNA sequences can encode the same protein. Choosing a DNA sequence is therefore not neutral. Codon usage can influence expression level, folding efficiency, and metabolic burden in the host organism. I am only beginning to understand how codon bias shapes expression outcomes. What appears to be a simple reverse translation step is in reality a probabilistic optimization problem shaped by cellular context.
3.3 Codon Optimization
Why codon optimization is necessary
Although multiple DNA sequences can encode the same protein due to codon degeneracy, not all synonymous codons are used equally in different organisms.
If a gene uses codons that are rare in the host organism:
- Translation can be slow
- Ribosomes may stall
- Protein yield can decrease
- Misfolding may increase
Codon optimization ensures that the nucleotide sequence:
- Matches the host organism’s codon usage bias
- Improves translation efficiency
- Reduces metabolic burden
- Avoids problematic motifs (e.g., strong secondary structures, cryptic promoters, restriction sites)
Thus, codon optimization improves its expression in a chosen host.
Organism chosen for optimization
I chose to optimize the tyr1 gene for expression in: Komagataeibacter rhaeticus
This organism is the cellulose-producing bacterium used in Walker et al. (2024) and is central to my broader project on biologically functionalized bacterial cellulose.
Because K. rhaeticus has a relatively high GC content and a codon usage profile distinct from Bacillus megaterium, optimization is required to:
- Improve translation efficiency
- Ensure stable protein production
- Support effective melanin biosynthesis
Additional design constraints
Following best practices (e.g., Twist Bioscience optimization tools), the optimized sequence was also designed to:
- Avoid Type IIS restriction enzyme recognition sites:
- BsaI
- BsmBI
- BbsI
- Avoid internal repetitive motifs
- Maintain balanced GC distribution
Codon-Optimized DNA Sequence (for K. rhaeticus)
3.4 You have a sequence / Now what?
Now that we have a codon-optimized DNA sequence encoding Tyr1, the next step is to express the protein.
How can this protein be produced?
There are two main strategies:
A. Cell-dependent expression (in vivo)
In this approach, the DNA sequence is inserted into a plasmid vector containing:
- A promoter (constitutive or inducible)
- A ribosome binding site (RBS)
- The coding sequence (our optimized tyr1)
- A transcription terminator
The plasmid is then introduced into a host organism (e.g., Komagataeibacter rhaeticus).
What happens biologically?
1️⃣ Transcription
RNA polymerase binds to the promoter and transcribes the DNA sequence into messenger RNA (mRNA).
2️⃣ Translation
Ribosomes bind to the mRNA at the ribosome binding site.
The mRNA codons (triplets) are read.
tRNAs deliver amino acids corresponding to each codon.
The ribosome assembles the amino acids into the Tyr1 protein.
3️⃣ Protein folding and function
The translated Tyr1 protein folds into its 3D structure.
Copper ions bind to the active site.
The enzyme becomes catalytically active.
In the case of Tyr1, the enzyme then catalyzes: L-tyrosine → L-DOPA → dopaquinone → eumelanin Thus, genetic information becomes a catalytic material transformation.
B. Cell-free expression (in vitro)
Alternatively, the DNA sequence can be expressed using a cell-free transcription-translation system.
These systems contain:
- RNA polymerase
- Ribosomes
- tRNAs
- Amino acids
- Energy sources
The optimized DNA is added directly to the mixture.
Transcription and translation occur outside living cells.
Advantages:
- Rapid prototyping
- No need for transformation
- Easier control of conditions
For exploratory material functionalization, cell-free systems can serve as a fast validation step before in vivo implementation.
From DNA to Protein: Summary
DNA (double-stranded)
→ transcription →
mRNA (single-stranded)
→ translation →
Protein (amino acid chain)
This is the operational flow of the Central Dogma.
From DNA to material interface
If we follow the chain of transformations:
DNA (tyr1 gene)
→ mRNA
→ Tyrosinase enzyme
→ Melanin polymer formation
→ Integration into the cellulose matrix
→ Modified impedance behavior
→ Human touch interaction
This illustrates that writing DNA is not only about modifying organisms. It is about programming how matter self-organizes over time.
My interest is not genetic novelty for its own sake, but how cellular metabolism reorganizes matter into functional biofilms.
4. Prepare a Twist DNA Synthesis Order
4.1. Create a Twist account, and Benchling account
To prepare the DNA synthesis order, I created:
- A => Twist Bioscience account https://ecommerce.twistdna.com/app
- A => Benchling account https://benchling.com
Benchling was used to design and annotate the DNA construct.
Twist was used to validate and prepare the sequence for synthesis.
The idea is to make a manufacturable artifact, marking the transition from theory to industrial bioengineering.
4.2. Build Your DNA Insert Sequence
The expression cassette was assembled in Benchling as a linear DNA construct including:
- Promoter
- RBS
- Coding sequence (Tyr1)
- Stop codon
- Terminator
Benchling file:
ZIP version here : 📄 Tyr1 ZIP
Annotated sequence overview
Benchling allows visualization of the construct with annotated features and automatic translation of the coding sequence.
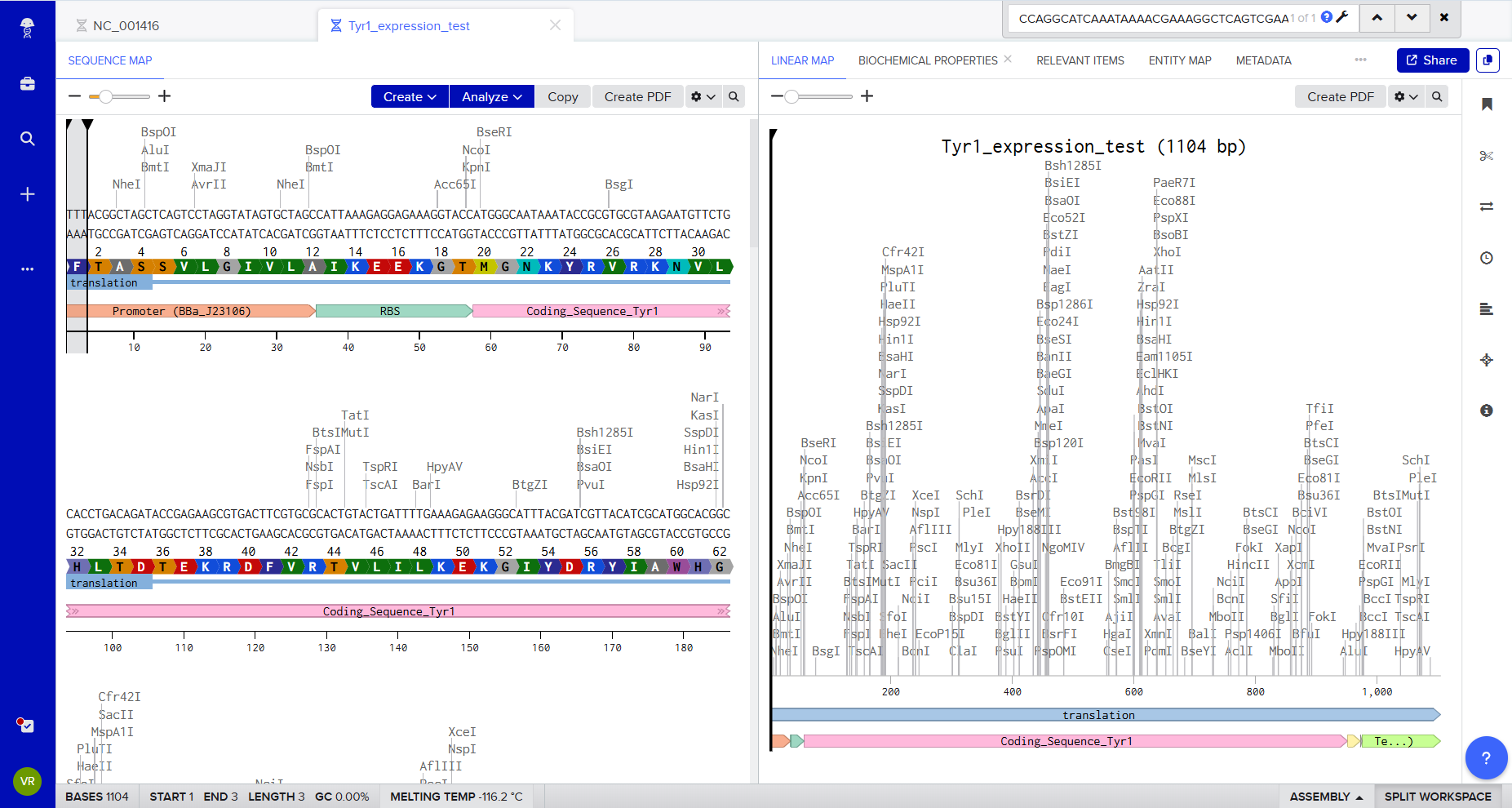

This confirms:
- Correct open reading frame
- Proper annotation of CDS
- Expected protein translation
Twist synthesis validation
The sequence was uploaded to Twist Bioscience for synthesis validation and plasmid build preparation.
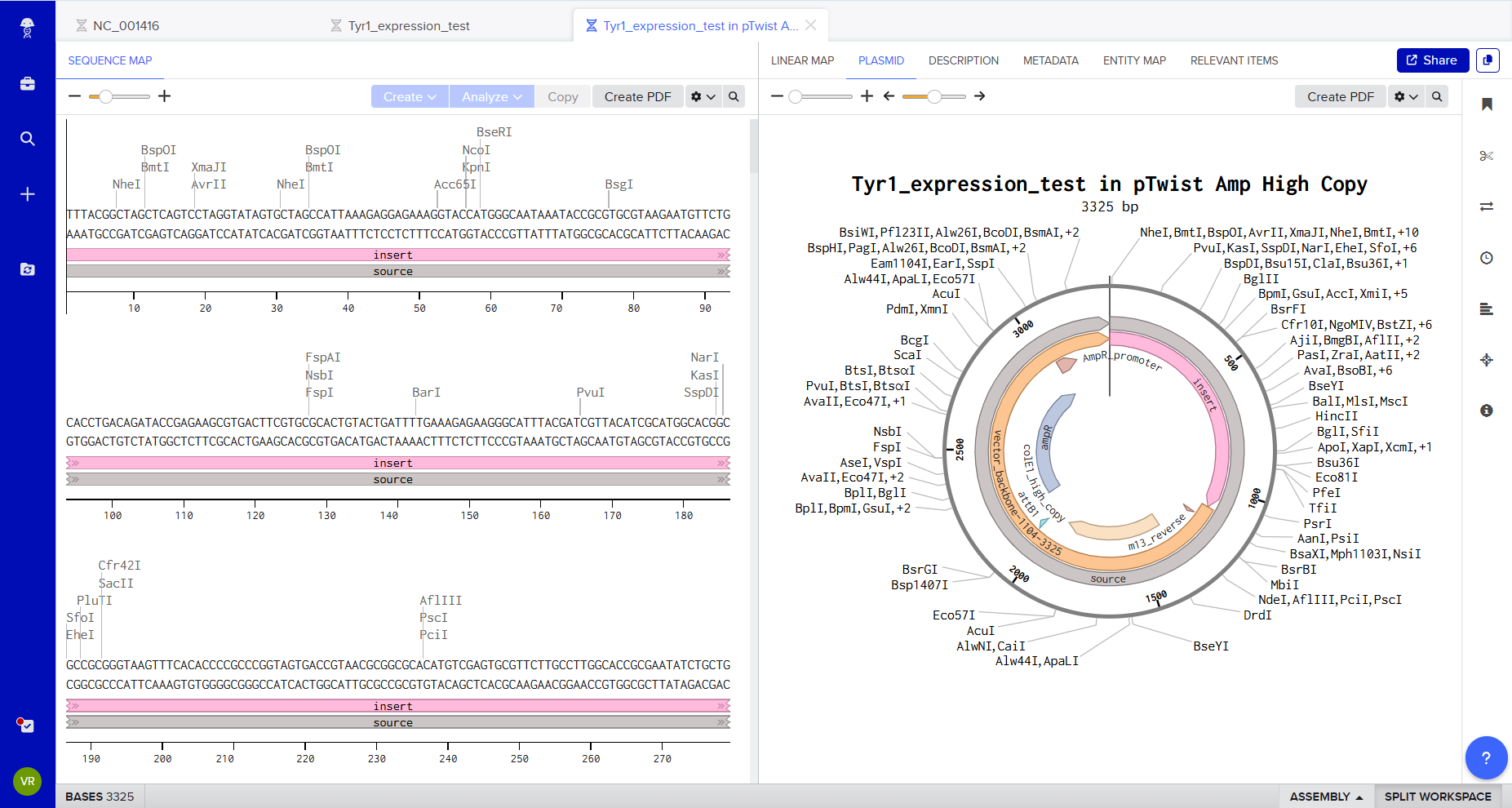

Twist checks for:
- Sequence length constraints
- GC content
- Secondary structures
- Restriction site conflicts
- Synthesis compatibility
This step transforms a conceptual genetic design into a manufacturable DNA construct.
Files
📄 Download: Tyr1 synthesis package (ZIP)
Optional: SBOL visualization
To visually represent the genetic architecture, the construct can also be recreated using:
🔗 https://sbolcanvas.org
SBOL Canvas enables graphical design of:
Promoter → RBS → CDS → Terminator
This helps communicate the logic of the construct clearly and aligns with synthetic biology design standards.
5. DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would I want to sequence and why?
I would sequence synthetic DNA used for digital data storage. Instead of reading DNA from living organisms, I am interested in DNA that has been artificially synthesized to encode digital information (text, images, archives). In this case, DNA is not used as biology — it is used as a storage material.
Why this matters
DNA can store enormous amounts of information in a very small volume and can remain stable for long periods of time. Unlike hard drives or servers, it does not depend on electronic infrastructure. Sequencing such DNA means reading information written into molecules. It turns a biological technology into a digital decoding tool. This connects biology with computation and raises important questions about how information can be stored, preserved, and accessed in the future.
(ii) Which sequencing technology would I use and why?
I would use Illumina sequencing.
Why Illumina?
DNA used for digital storage is usually made of short fragments. Illumina sequencing works very well for short pieces of DNA and provides very high accuracy, which is essential when decoding stored data. Small sequencing errors could corrupt the recovered file, so accuracy is more important than read length.
What generation is it?
Illumina is considered a second-generation sequencing technology because it reads millions of DNA fragments at the same time using amplified clusters and fluorescent signals.
What is the input and preparation?
Input:
Synthetic double-stranded DNA fragments (around 100–200 base pairs).
Basic preparation steps:
- Add short adapter sequences to the DNA fragments
- Amplify the fragments
- Load them onto the sequencing machine
How does Illumina read DNA?
The machine copies the DNA one base at a time.
Each base (A, T, C, G) produces a specific fluorescent signal.
A camera records the signal after each step, and software converts these signals into a DNA sequence.
What is the output?
The output is a digital file containing:
- The DNA sequences
- A quality score for each base
In DNA data storage, these sequences are decoded back into binary information using error-correction algorithms.
Why not Nanopore?
Nanopore sequencing can read longer DNA fragments, but it generally has higher error rates. For digital data storage, accuracy is more important than read length, so Illumina is currently more suitable.
Conceptual note
In this case:
DNA is not a gene.
It is a storage medium.
Sequencing is not diagnosing life.
It is decoding information.
5.2 DNA Write
(i) What DNA would I want to synthesize and why?
I would synthesize a genetic construct enabling melanin production in cellulose-producing bacteria.
More specifically, I would synthesize a codon-optimized version of the tyrosinase gene (tyr1) from Bacillus megaterium, placed inside an expression cassette designed for bacterial hosts.
The goal is to enable engineered Komagataeibacter strains (cellulose-producing bacteria) to produce eumelanin, a dark redox-active polymer, directly within the growing cellulose matrix.
This would allow:
- Pigmented living materials
- Potentially electroactive cellulose composites
- A direct link between gene expression and macroscopic material properties
In this case, DNA is used not only to encode a protein, but to program a material property.
Example genetic construct (simplified expression cassette)
Promoter → RBS → tyr1 CDS → Stop → Terminator This DNA construct would enable constitutive expression of tyrosinase, leading to melanin formation when L-tyrosine is available.
(ii) What DNA synthesis technology would I use and why?
I would use phosphoramidite-based chemical DNA synthesis, the standard method used by companies like Twist Bioscience. This is currently the most reliable and scalable way to synthesize custom genes.
Essential steps of DNA synthesis
- Chemical synthesis of short oligonucleotides
- Assembly of oligos into full-length genes
- Error correction and amplification
- Cloning into a plasmid vector
- Sequence verification
Limitations
- Error rates increase with sequence length
- GC-rich or repetitive sequences are harder to synthesize
- Cost increases with size
- Large constructs require assembly from smaller fragments
- Scalability
Scaling biological material production is not a simple matter of increasing volume. In living systems, morphology, oxygen gradients, metabolic stress, and contamination risks can fundamentally alter structural outcomes. A construct that works at lab scale does not automatically behave the same way at industrial scale. Understanding small-scale control is therefore a necessary first step before considering scalability.
Despite these limits, gene-scale synthesis (1–3 kb) is highly robust today.
5.3 DNA Edit
My ambition is not maximal intervention, but controlled understanding. Before redesigning entire biosynthetic networks, I prefer mastering one enzymatic layer and observing its material consequences. Editing cellulose crystallinity, secretion pathways, or c-di-GMP regulation may be possible in the future. But at this stage, focusing on a single added function (tyr1 expression) allows a clearer understanding of cause and effect.
(i) What DNA would I want to edit and why?
I would want to edit the genome of a cellulose-producing bacterium (Komagataeibacter) to integrate a functional gene such as tyr1 directly into its chromosome.
Rather than keeping the gene on a plasmid, chromosomal integration would:
- Increase genetic stability
- Reduce reliance on antibiotic selection
- Make the engineered trait more sustainable
More broadly, DNA editing could be used to:
- Improve cellulose yield
- Modify crystallinity or fiber structure
- Control secretion of functional proteins
- Enable responsive or patterned material growth
The objective is material programming through living systems such as Komagataeibacter and raises technical and ecological questions regarding containment, stability, and unintended environmental interactions.
(ii) What technology would I use and why?
I would use CRISPR-Cas9 genome editing. CRISPR allows precise insertion or modification of DNA at specific genomic locations.
How CRISPR edits DNA
- Design a guide RNA (gRNA) targeting a specific genomic sequence
- Deliver Cas9 protein and guide RNA into the cell
- Cas9 creates a double-strand break at the target site
- Provide a donor DNA template containing the new gene
- The cell repairs the break using the donor template (homology-directed repair)
Required inputs
- Guide RNA sequence
- Cas9 enzyme (plasmid or ribonucleoprotein complex)
- Donor DNA template
- Competent cells
- Transformation system
Limitations
- Editing efficiency may be low in non-model organisms
- Off-target edits are possible
- Homology-directed repair is not always efficient
- Delivery systems can be challenging
Despite these challenges, CRISPR remains the most precise and flexible editing tool currently available.
Conceptual note
DNA writing defines what is possible.
DNA editing reshapes what already exists.
Together, they allow us to move from reading biology to programming living materials.
Personnal note
Writing DNA is not merely modifying organisms; it is programming how matter self-organizes over time.
6. Exploration of others strategies for my project
Compare 4 Conductivity Strategies (What They Really Do)
| Strategy | Main Effect | What Carries the Signal | Typical Ingredients | What Changes in the Material | Difficulty (1–5) |
|---|---|---|---|---|---|
| 1) Graphene/CNT in-situ | Strong conduction | Electronic percolation network | CNT, graphene/rGO, dispersion aid | Large drop in resistance, wire-like behavior | 2/5 |
| 2) PEDOT:PSS / Ag nanowires (post or in-situ) | Strong conduction | Polymer film or metallic network | PEDOT:PSS, AgNWs | High conductivity, less biological emergence | 3/5 |
| 3) Tyr1 → melanin (bio-made dopant) | Impedance / redox / protonic | Redox polymer + water/ions | Tyr1 + Cu²⁺ + tyrosine (neutral pH development step) | Response depends on hydration and pH; useful for sensing | 3/5 |
| 4) Curli-like programmable fibers | Structure + hybrid conduction | Protein fibers + mineralization or binding | Curli operon + metal-binding peptides | Programmable and patternable but multi-gene | 4/5 |
Reading this table
- If the goal is to create a wire-like conductive material, strategies 1–2 are the most effective.
- If the goal is to create a living sensing surface, strategy 3 (and potentially 4 later) is more coherent.
Is Tyr1 Enough for a Tactile Sensor?
What I am actually building with Tyr1 + BC
With Tyr1/melanin integrated into bacterial cellulose, I am not creating a high-performance conductor. Instead, I am most likely creating an impedance-based sensor.
Two realistic sensing behaviors:
Touch or pressure → impedance change
Mechanical compression modifies microstructure and redistributes water and ions.
→ Impedance changes, especially at specific AC frequencies.Hydro-tactile response (touch + humidity)
BC is highly sensitive to hydration.
→ Strong signal variation, but humidity must be controlled to avoid false readings.
Does the material need to remain alive?
Not necessarily. Two modes are possible:
Living mode
The material continues to grow and self-organize.
Advantage: conceptual strength, biological emergence.
Limitation: signal drift over time due to metabolism and morphology changes.Stabilized mode
Washed and partially dried but rehydratable.
Advantage: more stable electrical behavior and easier reproducibility.
For experimental validation, a stabilized but responsive material may be more reliable.
Is electrodes + amplifier + ADC sufficient?
Yes — provided that impedance (AC) is measured correctly.
Basic workflow:
- Inject a small, known AC excitation signal
- Measure amplitude (and ideally phase)
- Test at 2–3 different frequencies
Possible hardware:
- Dedicated impedance IC (e.g., AD5933 / AD5940)
- Or a simple bridge with sinusoidal excitation and amplitude measurement
Electrode configurations:
2-electrode setup
Simpler, but more sensitive to contact variability.4-electrode setup
Better separation of excitation and measurement, more stable results.See what I’ve already done with Bacterial cellulose and electrodes signals :
Minimal validation experiment
To test whether Tyr1 truly contributes:
- BC control (no Tyr1)
- BC + Tyr1 + Cu²⁺/tyrosine development step
- Compare impedance vs pressure and humidity
If Tyr1 produces a measurable difference compared to control, the biological contribution is validated.
Why Copper Matters and Hybrid Strategies
Do I need to add copper?
Yes.
Tyrosinase is copper-dependent.
Without sufficient Cu²⁺, enzyme activity drops significantly.
However:
- Too little copper → weak melanin synthesis
- Too much copper → toxicity and growth inhibition
Copper availability must therefore be optimized, not maximized.
Can other dopants improve impedance?
Yes, but carefully.
Impedance in BC systems is strongly influenced by:
- Hydration level
- Ionic concentration
- Microstructure and porosity
A mild hybrid strategy can be interesting:
- Tyr1 provides a biologically synthesized redox layer
- Controlled ionic tuning (salts, buffering, hydration control) shapes electrical response
This maintains biological emergence while improving signal stability.
Long-Term Direction: Redox → c-di-GMP → Cellulose Organization
c-di-GMP is a central intracellular regulator controlling biofilm formation and cellulose production.
In Komagataeibacter, cellulose synthesis is tightly linked to c-di-GMP levels.
A future direction could be:
- A redox or electrical stimulus alters intracellular c-di-GMP levels
- This modifies cellulose production or organization
- Which then changes macroscopic impedance
This would represent a deeper form of biological programming, where structure and electrical behavior emerge from intracellular regulation.
However, this approach is high complexity (4/5–5/5) and better suited as a long-term research direction.