Week 2: DNA Read/Write/Edit :
1. Benchling & In-silico Gel Art
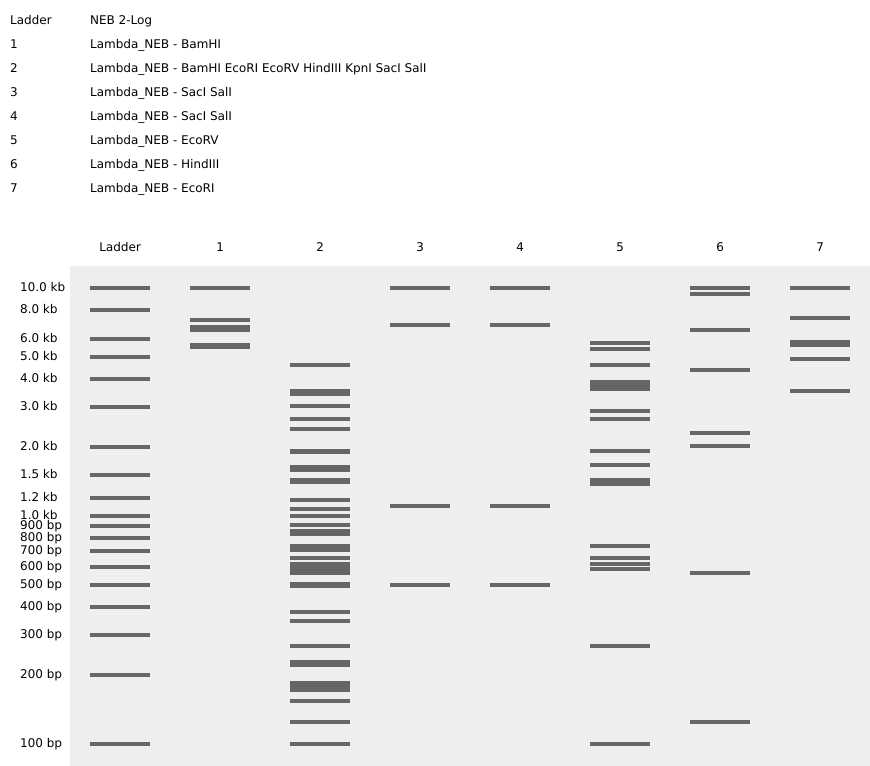


3. Protein Design Challenge
3.1 Protein Choice
pHRed is a red fluorecent protein used to detect the pH in live cells. CO2 acts as an acid in the blood and forms carbonic acid (Garcia and Ramirez) and reduces the pH, so this can be used to create a biosensor to detect the pH of blood. Using the of pH is a more accurate data point to understand the condition of the blood and is currently done throught an artiel blood gas test (Cleveland Clinic).
The protein sequence is blow
NSRIATEGRIDTRFRGSIKNVTVSGENHETFDLINIFRDKSSVGIVKNLGEATITEKTSYDYTHTGDYLISATPVPFTFTEFDKLVRFSPXDFEQFESIEIQFRTKEPRGVLLFVGPDNAHTDYVCLEFYDRNLYLAFGIDGKDYRKQMNPKGTFVTTGNFHTIFIKRDRNHKFTAKFENVEVDIGDQSGHQREFGSYTYIGGIDNPSRLPWYVWSREGFVGCINYMRVNEDKYIDPRGKNNQYSGDIAGVDIGKCLNDVRHCTASHCEG
3.2 Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
Using bioinformatics.com i reversed the protein into the below DNA sequence
aacagccgcattgcgaccgaaggccgcattgatacccgctttcgcggcagcattaaaaacgtgaccgtgagcggcgaaaaccatgaaacctttgatctgattaacatttttcgcgataaaagcagcgtgggcattgtgaaaaacctgggcgaagcgaccattaccgaaaaaaccagctatgattatacccataccggcgattatctgattagcgcgaccccggtgccgtttacctttaccgaatttgataaactggtgcgctttagcccgnnngattttgaacagtttgaaagcattgaaattcagtttcgcaccaaagaaccgcgcggcgtgctgctgtttgtgggcccggataacgcgcataccgattatgtgtgcctggaattttatgatcgcaacctgtatctggcgtttggcatt
gatggcaaagattatcgcaaacagatgaacccgaaaggcacctttgtgaccaccggcaactttcataccatttttattaaacgcgatcgcaaccataaatttaccgcgaaatttgaaaacgtggaagtggatattggcgatcagagcggccatcagcgcgaatttggcagctatacctatattggcggcattgataacccgagccgcctgccgtggtatgtgtggagccgcgaaggctttgtgggctgcattaactatatgcgcgtgaacgaagataaatatattgatccgcgcggcaaaaacaaccagtatagcggcgatattgcgggcgtggatattggcaaatgcctgaacgatgtgtcgccattgcaccgcgagccattgcgaaggc
The concensus sequence
aaywsnmgnathgcnacngarggnmgnathgayacnmgnttymgnggnwsnathaaraaygtnacngtnwsnggngaraaycaygaracnttygayytnathaayathttymgngayaarwsnwsngtnggnathgtnaaaayytnggngargcnacnathacngaraaracnwsntaygaytayacncayacnggngaytayytnathwsngcnacnccngtnccnttyacnttyacngarttygayaarytngtnmgnttywsnccnnnngayttygarcarttygarwsnathgarathcarttymgnacnaargarccnmgnggngtnytnytnttygtnggnccngayaaygcncayacngaytaygtntgyytngarttytaygaymgnaayytntayytngcnttyggnathgayggnaargaytaymgnaarcaratgaayccnaarggnacnttygtnacnacnggnaayttycayacnathttyathaarmgngaymgnaaycayaarttyacngcnaarttygaraaygtngargtngayathggngaycarwsnggncaycarmgngarttyggnwsntayacntayathggnggnathgayaayccnwsnmgnytnccntggtaygtntggwsnmgngarggnttygtnggntgyathaaytayatgmgngtnaaygargayaartayathgayccnmgnggnaaraayaaycartaywsnggngayathgcnggngtngayathggnaartgyytnaaygaygtnmgncaytgyacngcnwsncaytgygarggn
3.3 Codon optimization
Optimisation is necessary because genetic code is degenrate so the same amino acid can come from multiple codons. Optimising ensures the codons used are the most compatible for the host. Prokayotes also would not need post transcriptional modifications like eukrayotes cells because DNA in prakoyotes cells are within the same cell walls as ribosomes, it does not require the 5’ capping,3 Poly-A tail and splicing modifications. Using envectorbuilder.com i optimised this for yeast.
GCTGCTTGTGCTGGTTGTTGTGGTTGTGCTACAACTGGTTGTGGTGCTTGTTGTGGTGCAGCTGGTGGTTGTTGTGGTTGTGCTACTACCGGTGCAACTGCTTGTTGTTGTGGTTGTACTACTACTTGTGGTTGTGGCGGCTGTGCAGGTTGTGCAACTACTGCAGCTGCAGCAGCTTGTGGTACCGGTGCATGTTGTGGTACTGGTGCTGGTTGTGGAGGTTGCGGTGCTGCAGCCGCCTGTTGTGCTACAGGTGCAGCTGCATGCTGTACAACTACTGGTGCTACCTGTACAGGTGCTACAACTGCTGCTTGTGCAACAACAACTACTACATGTGGTTGTGGTGCAACTGCCGCTGCTGCAGGTTGTGCTGGTTGTGGTACAGGTGGTGGTTGTGCTACTACTGGTACTGGTGCTGCTGCCGCTGCTTGTTGTACTGGTGGAGGTTGTGGTGCTGCAGGTTGTGGTGCATGTTGTGCTACAACCGCATGTTGCGGTGCAGCCGCTGCTGCAGCATGTTGTGCAGGTTGTACAGCTACAGGTGCAACAACAGCAACTGCTTGTTGTTGTGCTACTGCTTGTTGTGGTGGTTGTGGTGCAACTACAGCTACATGTACTGGCGCTACAACTGCTGGTTGTGGTTGTGGTGCTTGTTGTTGTTGTGGTGGTACCGGTTGTTGTGGTACCACTACTGCATGTTGTACTACTACTGCTTGTTGTGGTGCTGCAACAACTACTGGTGCTACCGCTGCTGCTTGTACAGGTGGTACAGGATGTGGTTGTACAACTACTGCTGGTTGTTGCTGTGGTGGTGCTACAACTACAACTGGCGCTGCTTGTGCAGGTACTACTACCGGTGCTGCAGCTGGTTGTGCTACTACTGGTGCAGCCGCAACTACATGTGCTGGTACTACTACTTGTGGTTGTGCTTGTTGTGCAGCTGCTGGTGCTGCATGTTGCGGTTGTGGTTGTGGTGGTTGTGGTACCGGCTGCACTGGTTGTACAGGTACTACTACGGGTACAGGTGGTGGTTGTTGCTGTGGCGGTGCAACTGCTGCATGTGGTTGTGGTTGTGCTACCGCATGTTGCGGTGCTACTACTGCTACAGGTACAGGTACCGGTTGCTGTACCGGTGGTGCTGCTACAACAACTACAGCTACAGGTGCCACATGTGGTTGTGCTGCTTGTTGTACTGGTACTGCAACGTGCACAGGTGGTTGTGGTACAACTACTGGCGGTTGTGCTACTACTGGTGCTACTGGTGGTTGCGCAGCTGCCGGTGCAACTACAGCTACATGTGGTTGTGCTGCTGCATGTGCAGGTGCTACTGGTGCAGCATGCTGTTGTGGTGCTGCAGCTGGTGGTTGTGCTTGTTGTACAACTACAGGTACTGGTGCATGTTGCGCTTGTTGTGGTGGTTGTGCAGCTTGTACAACTACTTGTGCTACAGCGTGCTGTGCAACAACTACTACAACAGCTACTACTGCTGCAGCTTGTGGTTGTGGTGCAACTTGTGGTTGTGCTGCGTGTTGCGCTACTGCTGCTGCTACTACTACAGCATGTTGTGGTTGTGGTGCAGCAGCTACTACAACTGGTGCTGCTGCTGCTTGTGGAACAGGTGGTGCTGCCGGTACCGGTGGTGCGACAGCTACTACTGGTGGTTGCGGTGCTACCTGTGCTGGTGCTGGCTGTGGTGGTTGTTGCGCCACCTGTGCTGGTTGTGGTTGCGGTGCAGCCACTACAACAGGTGGTTGTGCAGGTTGTACTGCCACTGCTTGTTGTACGGCCACTGCCACTACTGGTGGTTGTGGTGGTTGTGCTACAACCGGTGCTACAGCTGCATGTTGTTGTGGTGCTGGTTGTTGTGGTTGTTGTACTGGTTGTTGTGGTACAGGTGGTACTGCTACTGGTACTGGCACCGGTGGTGCTGGTTGTTGCGGTTGTGGAGCTGCTGGTGGATGTACTACAACTGGTACAGGTGGTGGTTGCACTGGTTGTGCTACTACTGCTGCTTGTACCGCCACTGCTACAGGTTGTGGTTGTGGTACAGGTGCTGCATGTGGTGCAGCCGGTGCTACTGCAGCAGCCACTGCCACTGCAACAACTGGTGCTACTTGTTGCGGTTGTGGTTGTGGTGGTTGTGCAGCTGCTGCAGCTTGCGCAGCTTGTTGTGCCGGTACTGCAACAGCTGGTTGTGGCGGTTGCGGTGCAACAGCTACTACAGGTTGCGGTGGTGGTTGTGGTACAGGTGGTGCTACTGCTACAACAGGTGGATGTGCTGCCGCTACTGGTTGTTGTACTGGTGCAGCATGTGGTGCAACTGGTACTGGAACTTGTGGTTGTTGTGCCACTACTGGTTGTGCTTGTTGCGGTTGCGGAGCTGGATGTTGTGCTACTACTGGTTGTGGTGCAGCTGGTGGTTGTTGA
3.4 Protien Sythesis
Next step would require turning sythesising this DNA, using twist, to produce a tube of the sythesised DNA. For cell dependant sythesis, this has been optimised for yeast so the most cost effect method would be to use chemical transformation can be done to get the DNA into the yeast. This works by mixing the DNA with calcium chloride to neutralise the negative charges in the plasmid in an ice bath. This is then moved from the ice bath to 42º for 45s and then back to ice(A and Je). The change in tempreture creates a pressurised enviroment that pulls the DNA into the yeast. Once the DNA is in the yest, it begins transcription and enzymes create the matching mRNA which ribosomes read to assemble the amino acids to fold the protein. Cell free production would use ribosomes, enzymes and ATP’s in a test tube. The ribosomes should immedietly start tranlating and transcibing the DNA into protein.
4. Twist Orders
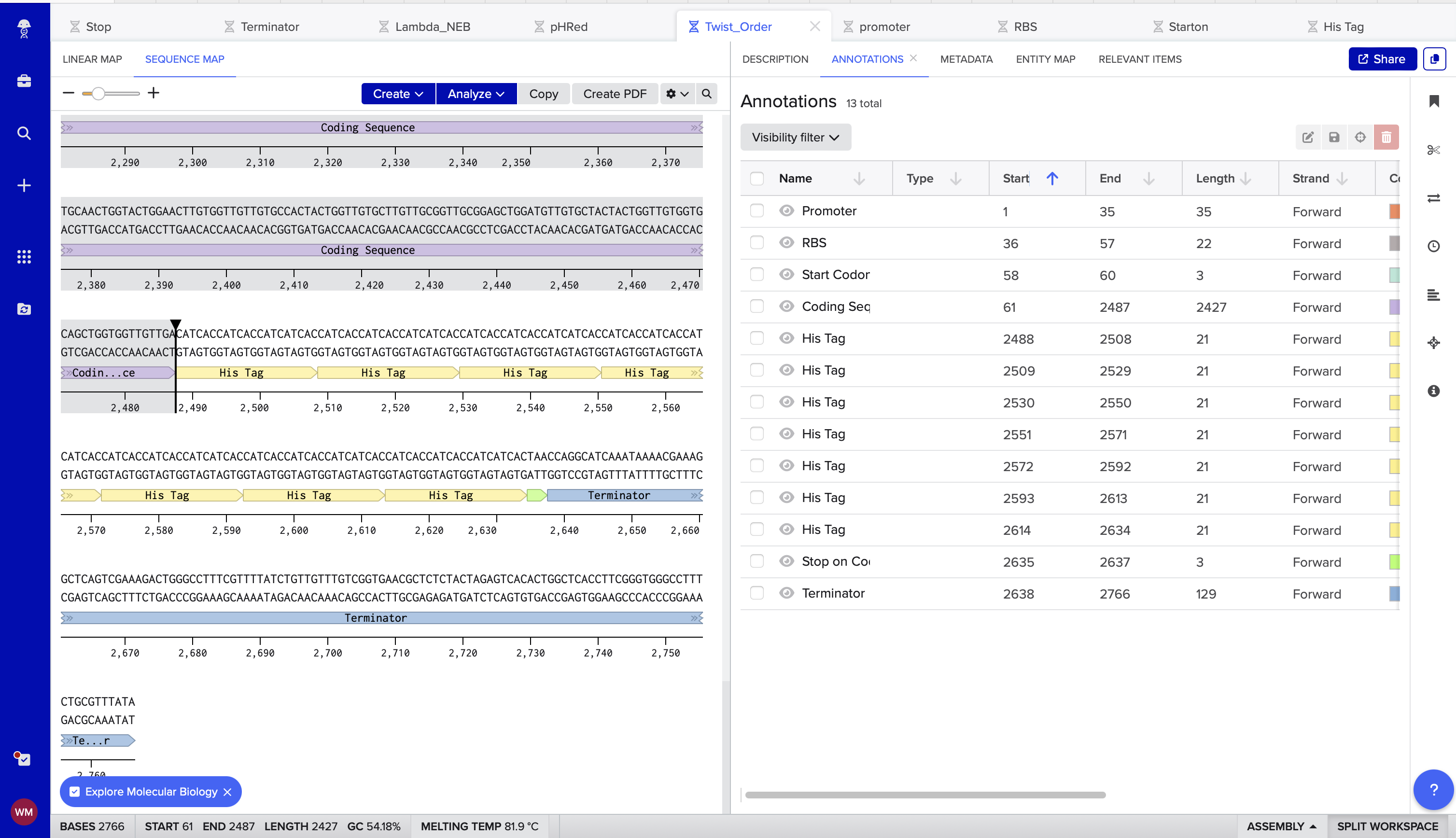
 Link to Benchling click [here] (https://benchling.com/s/seq-4NHKPQMzr1XF9FPeLKLG?m=slm-T6K9Zspozrn2EITqbIXm)
Link to Benchling click [here] (https://benchling.com/s/seq-4NHKPQMzr1XF9FPeLKLG?m=slm-T6K9Zspozrn2EITqbIXm)
For the vectors i will use pTwist PIC9K which has ampicilin resistance. I couldnt create a twist account as it said “Registration cannot be completed at this time”