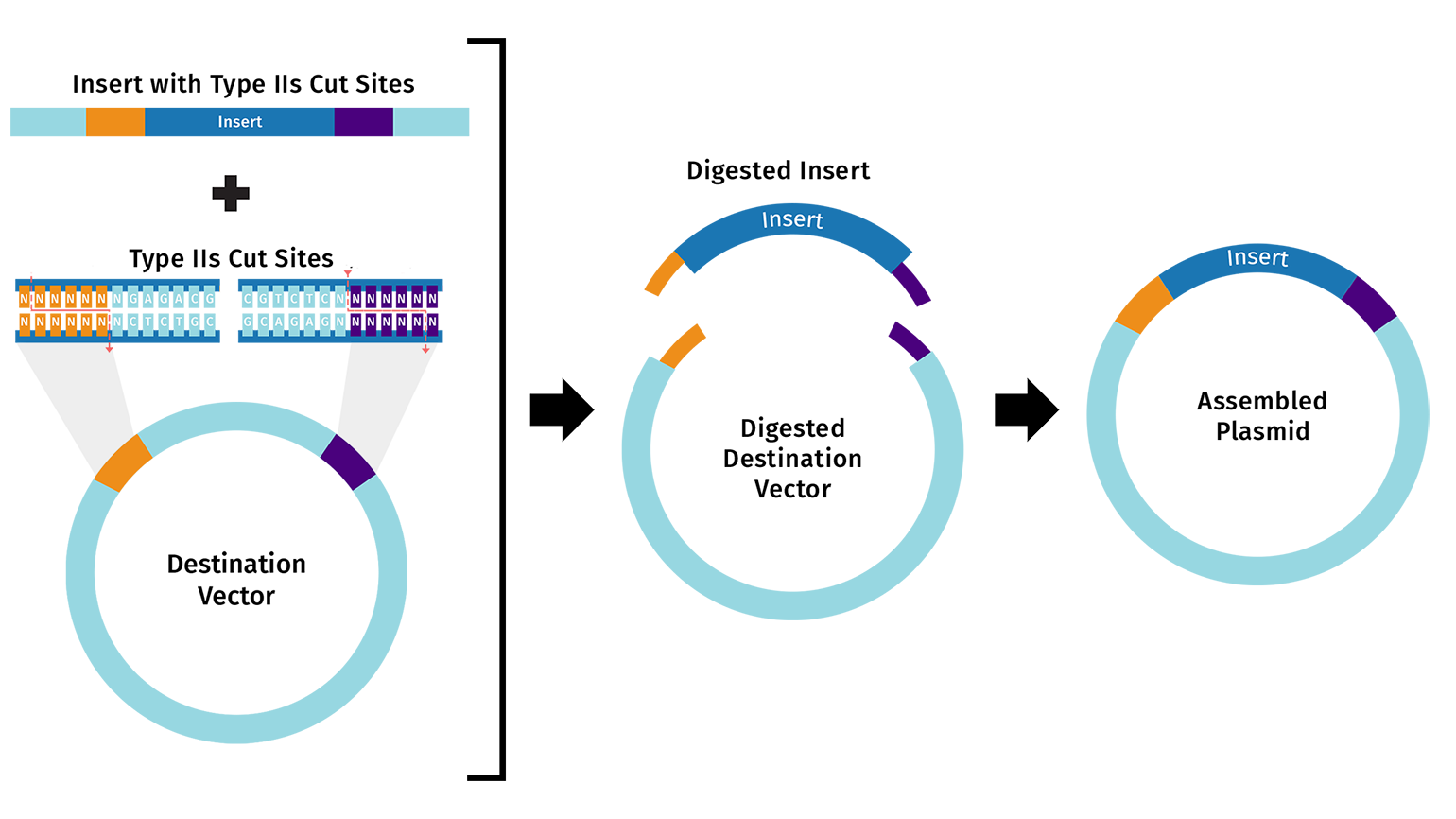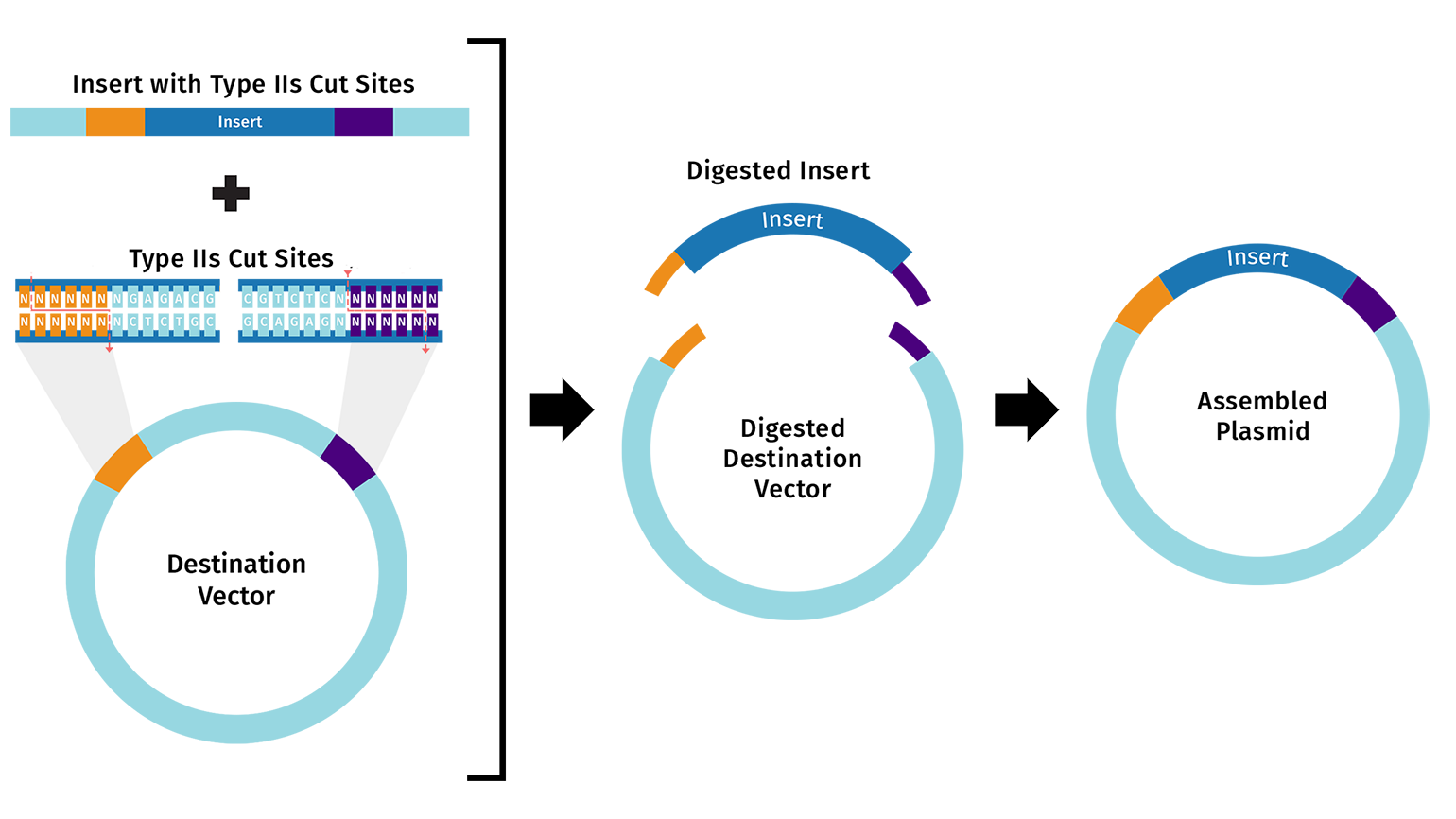Week 6: GENETIC CIRCUITS PART I
What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose? the Phusion High-Fidelity PCR Master Mix contains Phusion DNA polymerase, nucleotides, and an optimized reaction buffer including MgCl2
Phusion DNA polymerase is the enzyme that copies the DNA template, with high proofreading fidelity for accurate amplification.
dNTPs (nucleotides) is the building blocks the polymerase incorporates into the new DNA strand.
Optimized reaction buffer keeps the reaction conditions suitable for efficient polymerase activity, including pH and salt conditions.
MgCl2 provides magnesium ions, which are required as a cofactor for DNA polymerase function.
HF Buffer is the default high-fidelity buffer, and GC buffer is used for GC-rich or hard-to-amplify templates.
What are some factors that determine primer annealing temperature during PCR? Primer annealing temperature is mainly determined by the melting temperature of the primers: their length, GC content, and sequence composition. In practice, it is usually set a few degrees below the lower primer Tm so the primers can bind the template efficiently.
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR makes a specific DNA fragment by copying a template with primers and a DNA polymerase. The usual workflow is template DNA, forward and reverse primers, dNTPs, polymerase, buffer, and thermocycling through denaturation, annealing, and extension. This is best when you want a fragment defined by a sequence and not just by available cut sites. or if you want to introduce mutations, tags, overlaps or add tails in the primers.
Restriction digest A restriction digest makes fragments by using restriction enzymes that cut DNA at specific recognition sites. The workflow is usually DNA plus enzyme, buffer, incubation at the enzyme’s optimal temperature, then gel purification if needed. This is ideal when you want precise cuts at known sites already present in the DNA and have compatible ends for cloning into a vector.
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning? For Gibson cloning, the DNA fragments you generate by PCR need to have matching homologous overlaps at their ends, and the final construct has to be designed so adjacent pieces can anneal seamlessly. In practice, that means PCR primers should add the overlap sequences, and any restriction-digested backbone should be cut so the exposed ends correspond to the intended junctions in the assembly.
How does the plasmid DNA enter the E. coli cells during transformation?
The cell membrane is temporarily made permeable usually by heat shock or electroporation, which enables the plasmid to enter the E. coli and then the plasmid replicates and expresses its genes, including antibiotic resistance markers used for selection.
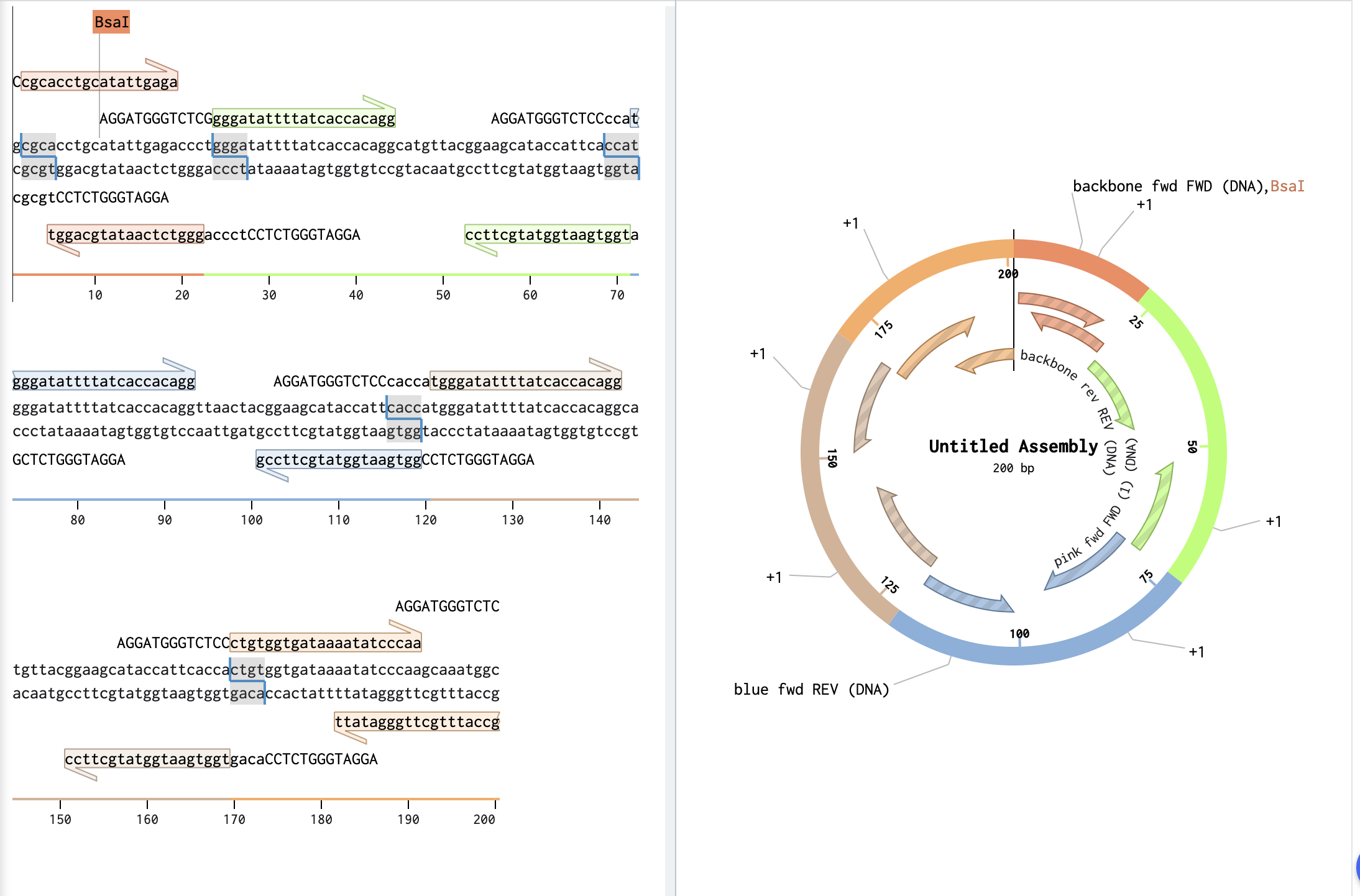
Describe another assembly method in detail (such as Golden Gate Assembly) Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online). Model this assembly method with Benchling or Asimov Kernel!
Golden Gate Assembly is a cloning method that combines restriction digestion and ligation in a single reaction. It uses a Type IIS restriction enzyme plus DNA ligase in one tube to cut DNA fragments and join them in an intended order, usually with scarless junctions. It is called scarless because the final construct does not retain the restriction sites at the junctions. Golden Gate works by designing each DNA part with unique overhangs so only the correct fragments join together. It is really useful for modular, multi-part cloning and for building larger or more complex constructs than a simple one-insert ligation.