Week 5 HW: Protein Design Part II
This week’s homework was divided into 3 parts.
Part A involved SOD1 Binder Peptide Design. That was broken into 3 parts: Part 1: Part 1: Generate Binders with PepMLM
A) Begin by retrieving the human SOD1 sequence from UniProt (P00441) and introducing the A4V mutation. Done B) Using the PepMLM Colab linked from the HuggingFace PepMLM-650M model card: Done C) Generate four peptides of length 12 amino acids conditioned on the mutant SOD1 sequence.
AHWGVYTGVKKAKRX 15.649067 AWVPPYAVVYALRAX 18.835777 SRWPPYAARVEWAKA 19.332172 SRYDEVVGVKKLRKX 14.812749
D) To your generated list, add the known SOD1-binding peptide FLYRWLPSRRGG for comparison.
Done
Record the perplexity scores that indicate PepMLM’s confidence in the binders.
Recorded
Part 2: Evaluate Binders with AlphaFold3
A) Navigate to the AlphaFold Server: alphafoldserver.com
Done
B) For each peptide, submit the mutant SOD1 sequence followed by the peptide sequence as separate chains to model the protein-peptide complex.
Submitted:
Original: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
1st: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQAHWGVYTGVKKAKRA
2nd: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQAWVPPYAVVYALRAA
3rd: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQSRWPPYAARVEWAKA
4th: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQSRYDEVVGVKKLRKA
5th (with mutant): MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQFLYRWLPSRRGG
All Xs were converted to As
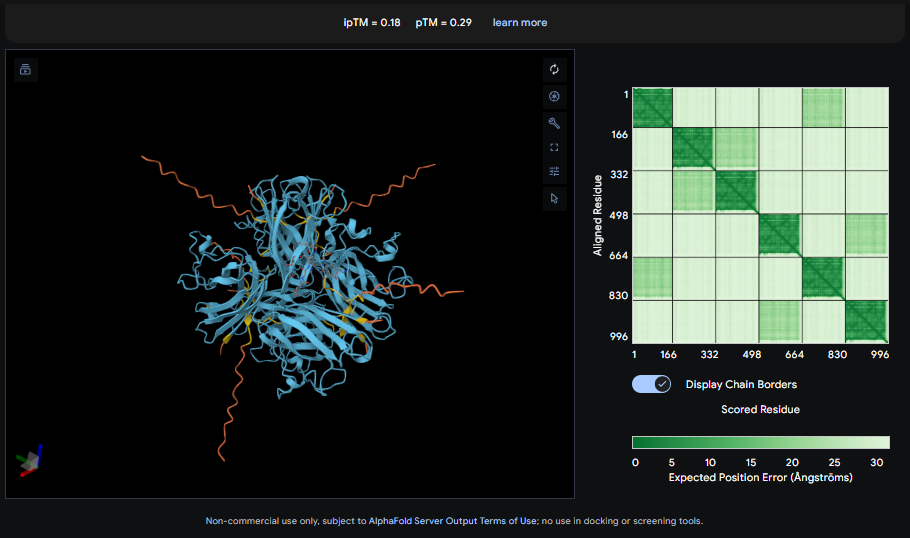
C/D) Record the ipTM score and briefly describe where the peptide appears to bind. Does it localize near the N-terminus where A4V sits? Does it engage the β-barrel region or approach the dimer interface? Does it appear surface-bound or partially buried? D) In a short paragraph, describe the ipTM values you observe and whether any PepMLM-generated peptide matches or exceeds the known binder.
The IpTM score is .18. This does not appear to indicate high confidence in binding. The lack of clear localiztion, strong β-barrel engagement, meaningful Dimer interfacing, combined with surface bounding of the peptides does not suggest strong binding interactions.

Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
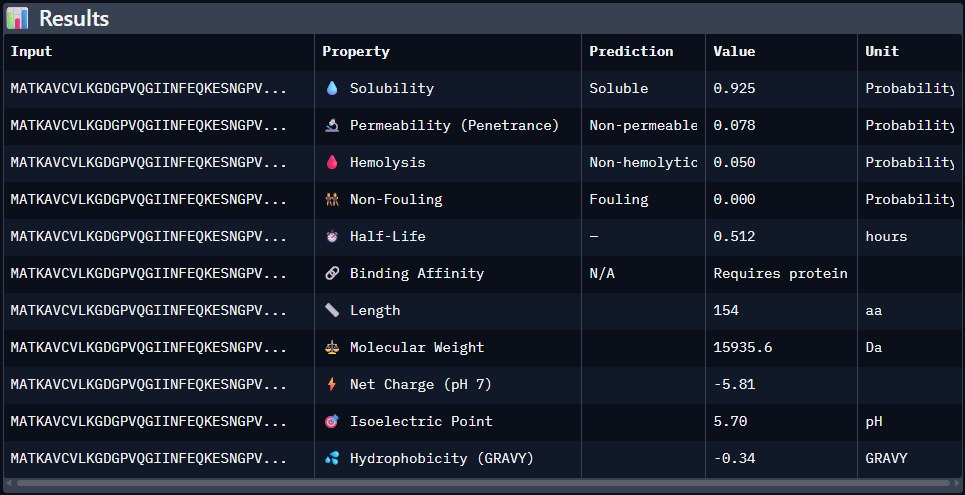
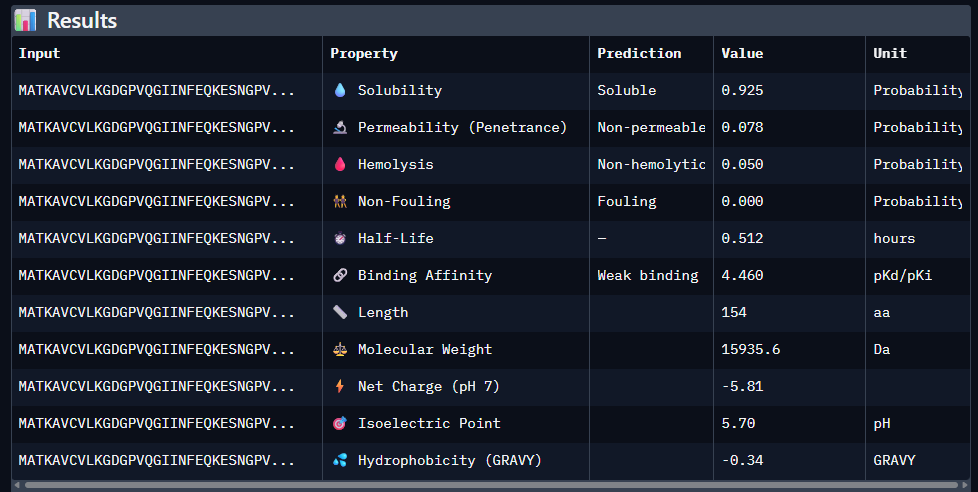
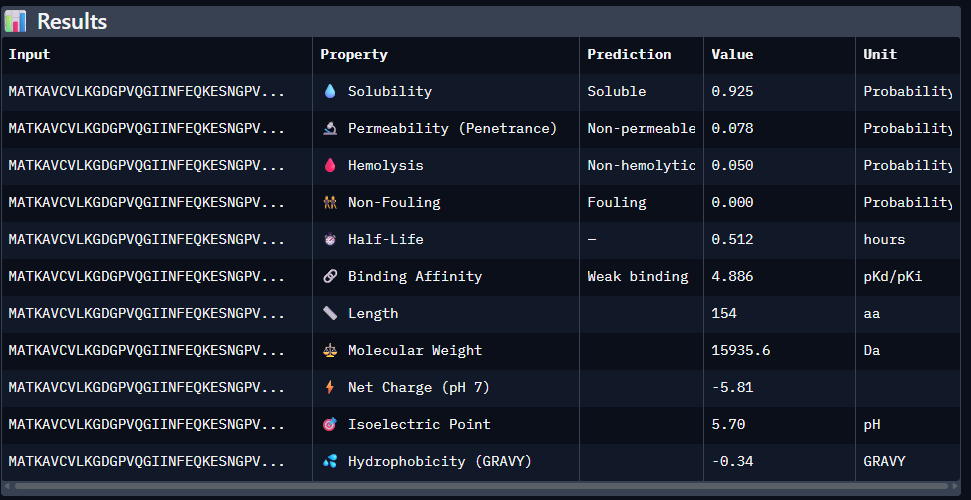
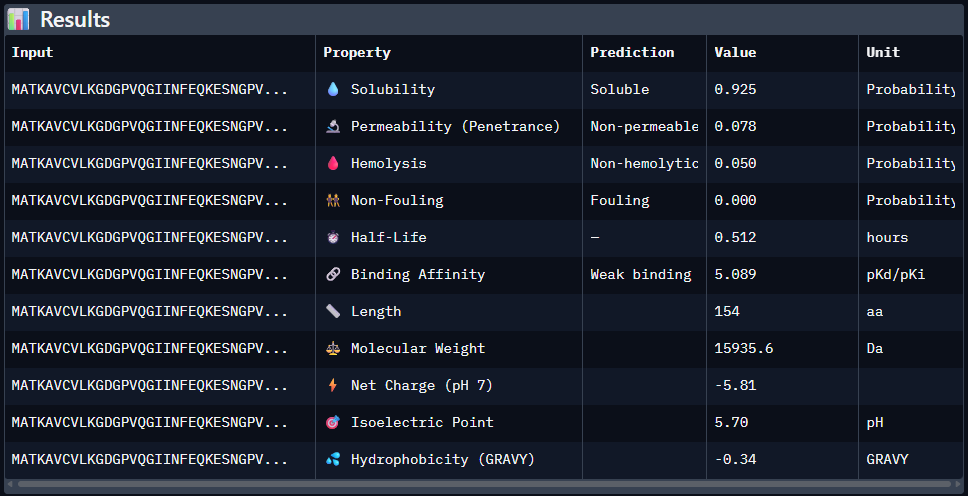
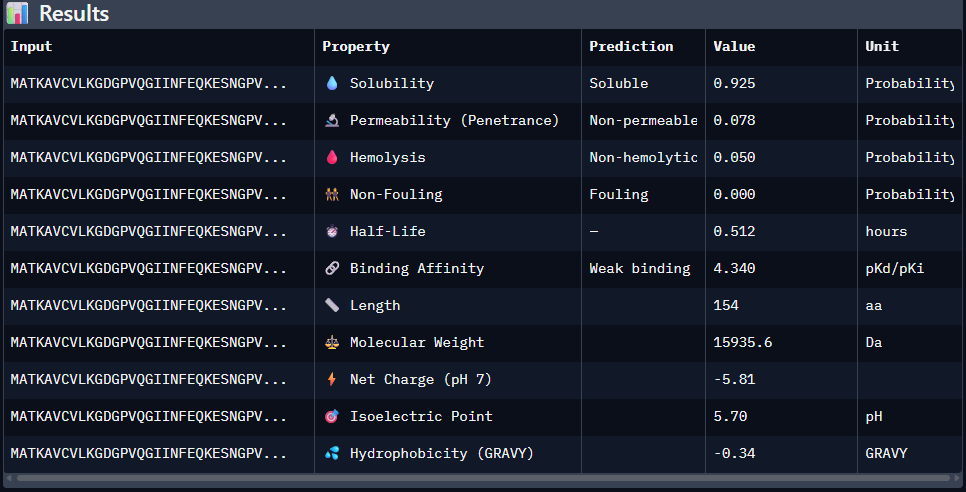
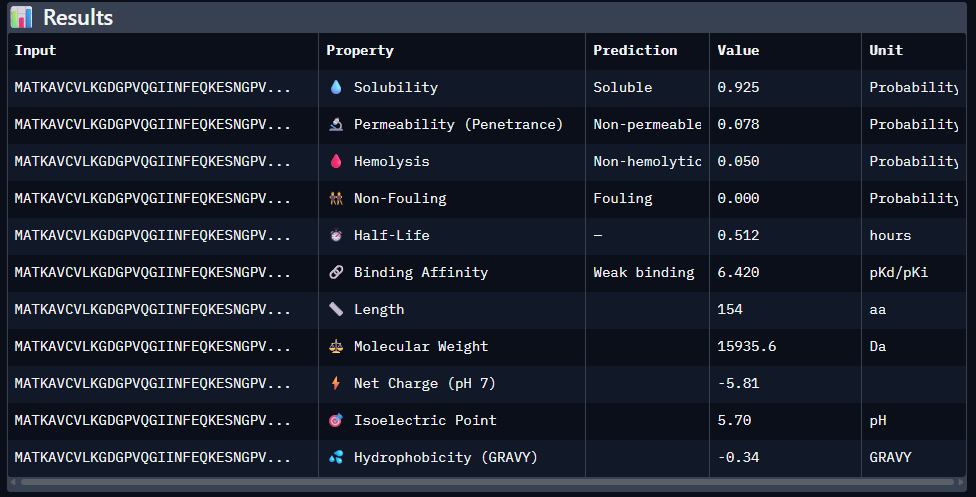
Paste the peptide sequence: Paste the A4V mutant SOD1 sequence in the target field. Check the boxes Predicted binding affinity Solubility Hemolysis probability Net charge (pH 7) Molecular weight
Data for such were pasted as pictures below, in this order:
Original 1st Sequence: AHWGVYTGVKKAKRX 2nd Sequence: AWVPPYAVVYALRAX 3rd Sequence: SRWPPYAARVEWAKA 4th Sequence: SRYDEVVGVKKLRKX 5th Sequence: Mutant
Part 4: Generate Optimized Peptides with moPPIt
Open the moPPit Colab linked from the HuggingFace moPPIt model card
Done
Make a copy and switch to a GPU runtime.
Done
In the notebook: Paste your A4V mutant SOD1 sequence. Choose specific residue indices on SOD1 that you want your peptide to bind (for example, residues near position 4, the dimer interface, or another surface patch). Set peptide length to 12 amino acids. Enable motif and affinity guidance (and solubility/hemolysis guidance if available). Generate peptides.
Done
After generation, briefly describe how these moPPit peptides differ from your PepMLM peptides. How would you evaluate these peptides before advancing them to clinical studies?
Erros were received and will be debugged.
Update:
This readout was given after fixing.
Binder Hemolysis Non-Fouling Solubility Half-Life Motif Specificity TEVEEQEDRQHH 0.932494111 0.910815179 0.916666687 1.714286923 0.001987276 0.934523821 LAAGQALGITTA 0.918551639 0.573852718 0.416666627 6.734929085 0.006364056 0.886904776 ARKLTPEDQKQG 0.961578194 0.903518379 0.75 2.08972311 0.080991976 0.803571403
moPPit peptides appear better to develop. I would evaluate these peptides further computationally before advancing them to clinical studies. This would involve ranking by various qualities and further optimization. If successful, in vitro studies would follow.
Part B: BRD4 Drug Discovery Platform Tutorial (Gabriele)
This assignment was optional. For the sake of priorities, it will not be posted here.
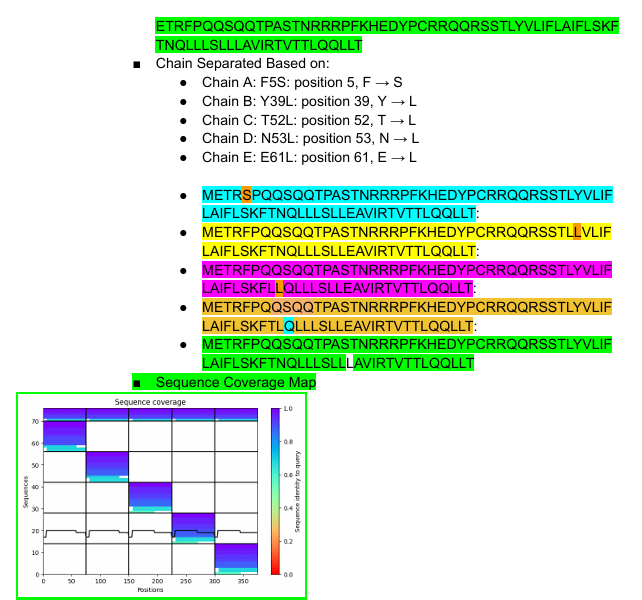
Part C: Final Project: L-Protein Mutants
This was held between of 5 members, 3 of whom were able to provide their results jointly. We persued the Option 1: Mutagenesis.
To save space, given the large volume of images, a link to the inputs can be found in this google doc:
https://docs.google.com/document/d/1676c1tgFUlGaP-Bwp9_vDexbk3VsOJeuQeylNfvz76o/edit?usp=sharing.
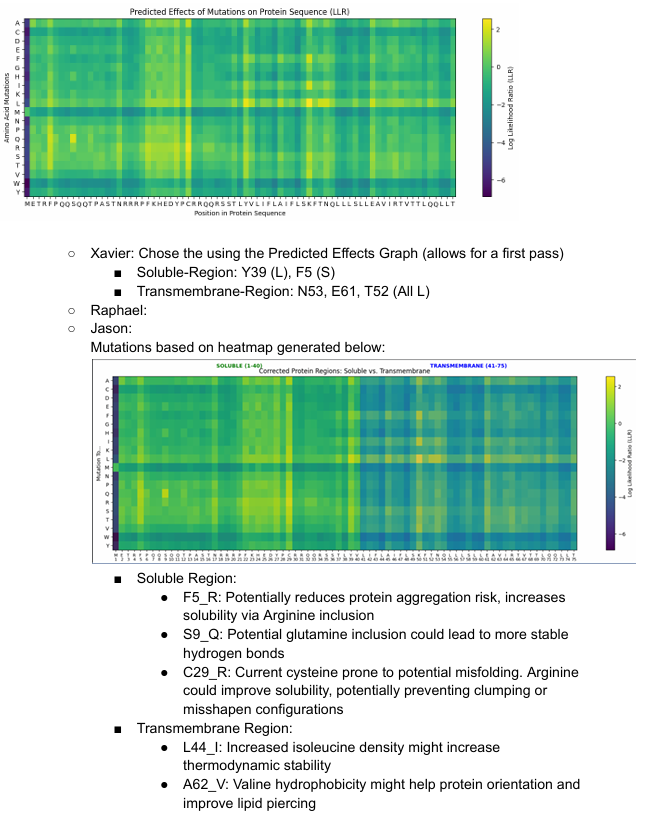
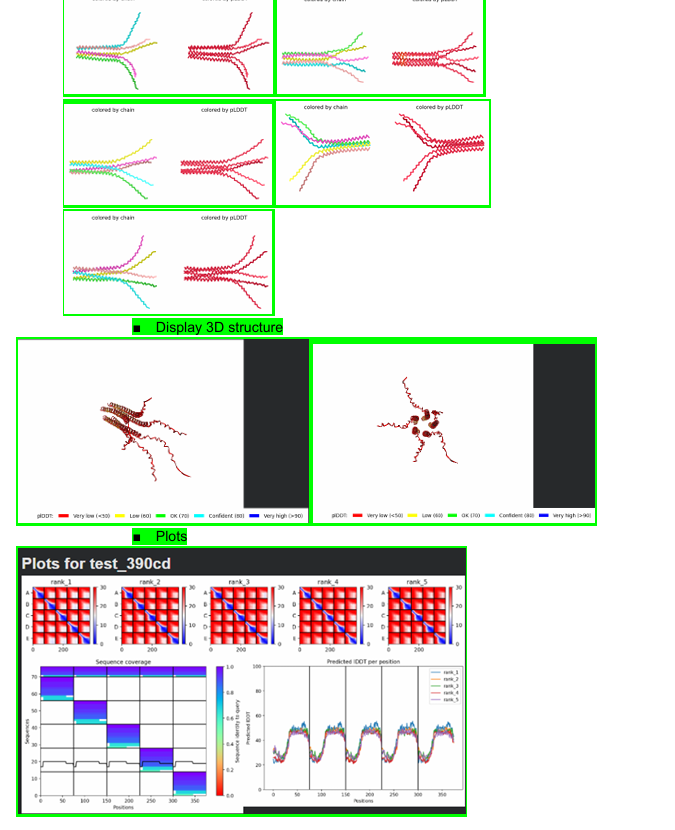
Some of the results will be shown below: