Week 6 HW: Genetic Circuits Part I: Assembly Technologies
Part 1: DNA Assembly
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Some of the components include the Phusion DNA Polymerease (key to PCR), dNTPs (these are used to synthesize new DNA), and buffering materials to stabilize the environmental conditions during the raction.
- What are some factors that determine primer annealing temperature during PCR?
Some of these include buffer conditions, primer melting temperature, primer length, and GC content.
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR uses primers, dNTPs, buffers and polymerase to amplify DNA. Restriction enzyme digests are very precise in that they cut genomic material at precise locations, but they require restriction sites.
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
They need to be optimally designed for the system. This requires special attention primer design and their parameters.
- How does the plasmid DNA enter the E. coli cells during transformation?
After a shock is induced, plasmid DNA can enter E. Coli cells through pores that emerge in their membrane.
- Describe another assembly method in detail (such as Golden Gate Assembly)
6.1 Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Golden gate assembly is a method for efficient, seamless assembly that uses Type IIS restriction endonucleases to cleave DNA that exists outside of recognition sequences. Designs for inserts and cloning vectors place the Type IIS recognition sites distal to the cleavage site, and this allows efficient removal. NEB states 3 main advantages for GGA. One is that no scar sequence is introduced. Another is that the overhangs can allow for multiple simulaneous assemblies. Further, ligation and digestion can be done at the same time. A truncated example of a successful case is diagrammed below, with inspiration from a figure on NEB’s website.
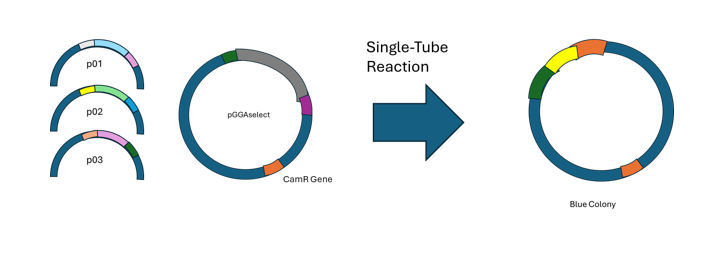

Source/Reference: https://www.neb.com/en/applications/cloning-and-synthetic-biology/dna-assembly-and-cloning/golden-gate-assembly
6.2 Model this assembly method with Benchling or Asimov Kernel!
Route chosen: Asimov Kernel
Owing to the brilliance of the Kernal from Asimov, redesigning the wheel was unnecessary.
I found the “pPD005 pcDNA Golden Gate” under the Addgene Plasmid Repository detailing a construct that was already created It is shown below in circular form.
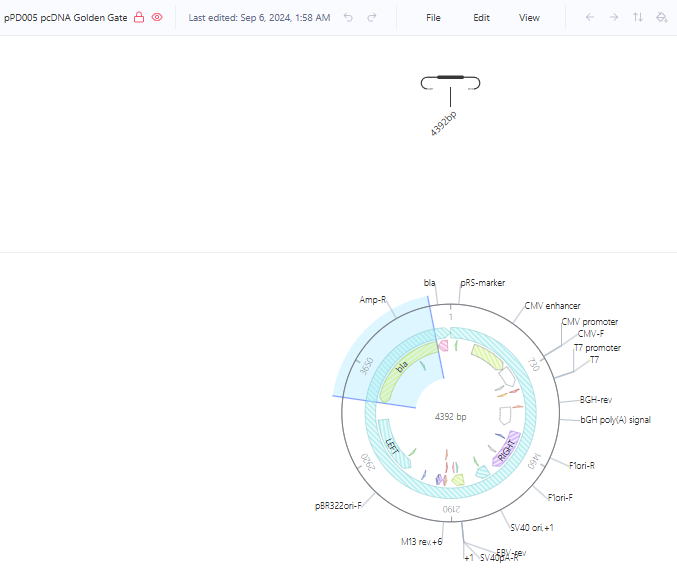

Part 2: Asimov Kernel
- Create a Repository for your work
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Create a blank Notebook entry to document the homework and save it to that Repository
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Explore the devices in the Bacterial Demos Repo to understand how the parts work together by running the Simulator on various examples, following the instructions for the simulator found in the “Info” panel (click the “i” icon on the right to open the Info panel)
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Create a blank Construct and save it to your Repository
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Recreate the Repressilator in that empty Construct by using parts from the Characterized Bacterial Parts repository
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Search the parts using the Search function in the right menu
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Drag and drop the parts into the Construct
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Confirm it works as expected by running the Simulator (“play” button) and compare your results with the Repressilator Construct found in the Bacterial Demos repository
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Document all of this work in your Notebook entry - you can copy the glyph image and the simulator graphs, and paste them into your Notebook
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Build three of your own Constructs using the parts in the Characterized Bacterials Parts Repo.
10.1. Explain in the Notebook Entry how you think each of the Constructs should function 10.2. Run the simulator and share your results in the Notebook Entry 10.3. If the results don’t match your expectations, speculate on why and see if you can adjust the simulator settings to get the expected outcome
10.1-10.3 These actions were all performed. I exported the images and markdown as below. However, they can be found under W&M_Whale_Test_Rep_Notebook_Entry To explain constructs – these were recreations and thus were expected to act as the demos, prior to the run of their simulations. The success is in part a testament to effect design of the Kernal software.
What Follows is a markdown export of my Notebook Entry in Ansimov Kernal under W&M Whale Test Rep:
This entry starts with a general summary of the use of Asimov’s Kernel thus far. The user interface takes a little getting used to but is easy to learn.
I started by using the Bacterial Demos Repo as per the homework to understand how parts work together.
A simulation was ran which ran similarly to others other before it. I did not expect difference given that these were demos.
The Repressilator was the first, which was selected by starting from a blank construct using parts from the Characterized Bacterial Parts.
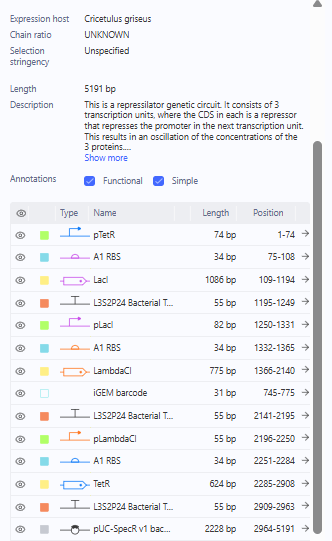

Characterized Bacterial Parts Examination for Repressilator


Represillator Recreation
This was then simulated with results expected from a Demo.
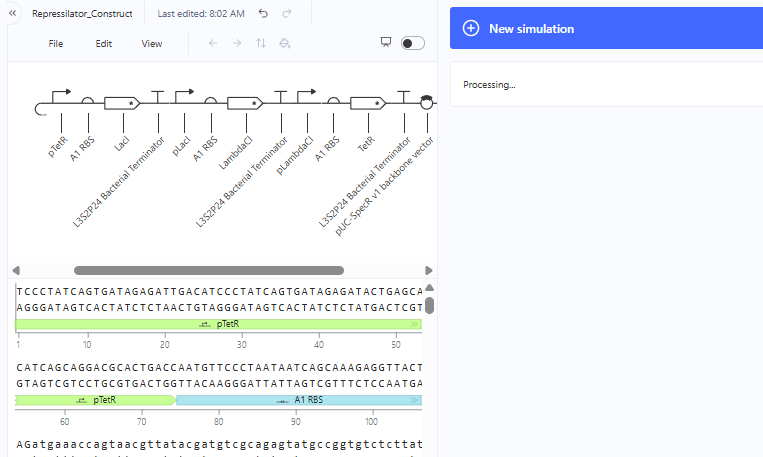

Represillator Recreation Simulation Start
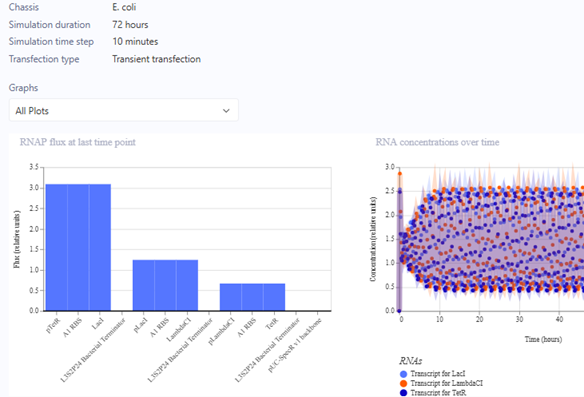

Represillator Recreation Preview
For the Homework, we were asked to create 3 more constructs. I chose recreations of the J23117 Promoter, Self-Replicating Promoter, and Circuit 3. Simulations accompanied them. I expected them to all run similar to the demos, which they did.
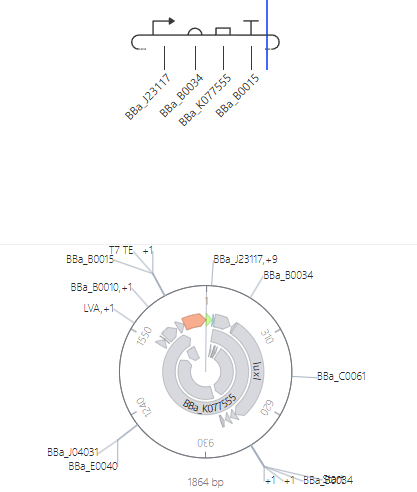

J23117 Promoter
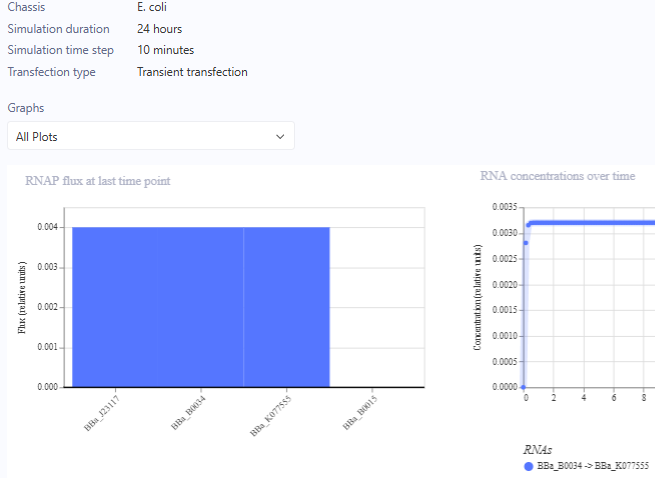

J23117 Promoter Simulation Preview
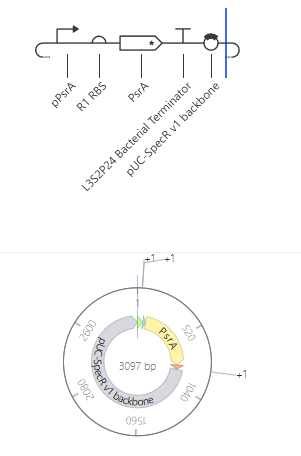

Self-Replicating Promoter
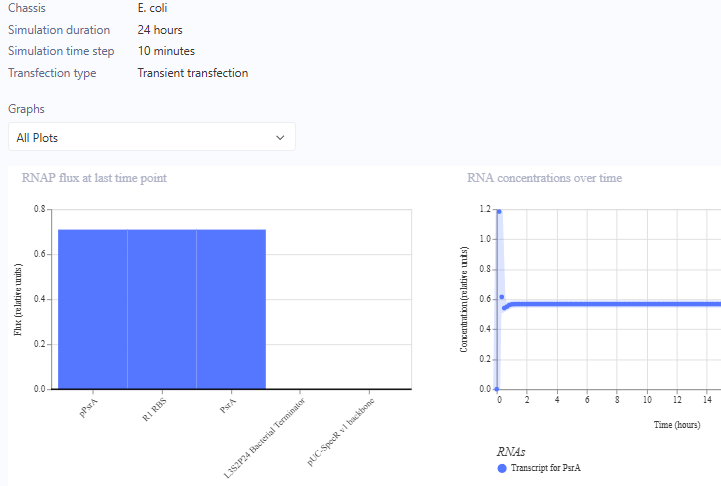

Self-Replicating Promoter Preview
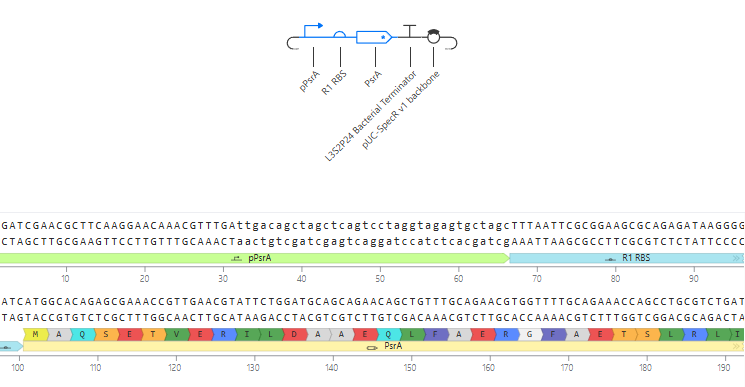

Circuit 3
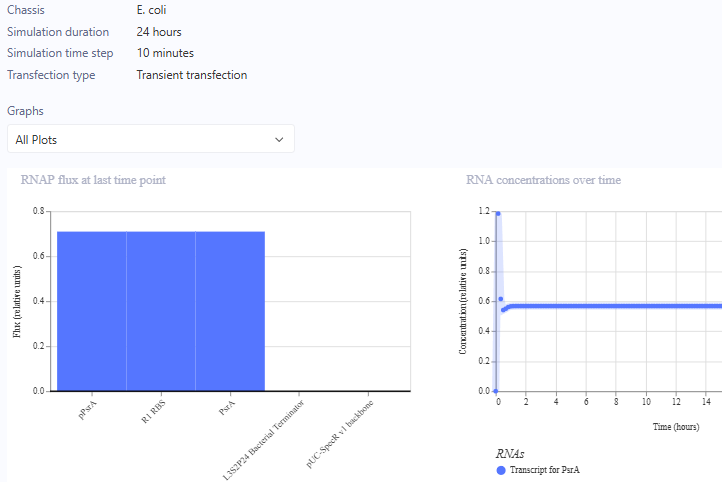

Circuit 3 Preview