Subsections of Homework
Week 1 HW: Principles and Practices
![cover image]()
![cover image]()
Describe a biological engineering application or tool you want to develop and why.
One application that might be worth exploring would be in the realm of data storage for an imagined, hypothetical semi-exposed media. For this context special proteins could be designed in applications that bind to sequences that code for hidden malware or faulty sequences and or act for preserving media. I imagine that both this media design and this protein use would be niche but nonetheless fun to design for. Modalities could be as a gentle spray or as a settled solution that could be extracted.
Describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm).
A policy goal to ensure that this application contributes to an ethical future could be ensuring transparent design and standardization of these proteins (and their stats per context) so there is a way to validate their production and application prior to use. This could reduce the chance of bad actors sabotaging batches or distributing lower quality batches.
- Describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”)
| Governance Action | Purpose | Design | Assumptions | Failure and Success Risks |
|---|
| Standardized Libraries | Educates communities on proteins available and safe designs | Create a library to hold these designs, as well as protocols for their manufacture to spec | That communities will agree on these standards and that there are not unnecessary inequities holding these standards | Success means that manufacture paths are straightforward.
Failure means that design paths are too numerous to account for and fragmentation possibilities are higher |
| Blue-teaming Design | To build a community line of protection of designs and applications | Design blue teaming frameworks and recruit educators to practice | That there is sufficient interest for funding | Success means that it is easier for production and share. Failure means that this pipeline has an established line of people who |
| Red-teaming Design | To identify means of sabotaging to proteins to protect libraries and distributors | Design blue-teaming framework and recruit educators to practice | That there is sufficient interest for funding | Success means that it is easier for production and share. Failure means that this pipeline has less visibility on sabotage routes |
- Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals.
| Does the option: | Option 1 | Option 2 | Option 3 |
|---|
| Enhance Biosecurity | 3 | 3 | 3 |
| • By preventing incidents | 3 | 3 | 3 |
| • By helping respond | 3 | 3 | 3 |
| Foster Lab Safety | 3 | 3 | 3 |
| • By preventing incident | 3 | 3 | 3 |
| • By helping respond | 3 | 3 | 3 |
| Protect the environment | 3 | 3 | 3 |
| • By preventing incidents | 3 | 3 | 3 |
| • By helping respond | 3 | 3 | 3 |
| Other considerations | 3 | 3 | 3 |
| • Minimizing costs and burdens to stakeholders | 3 | 2 | 2 |
| • Feasibility? | 3 | 3 | 3 |
| • Not impede research | 3 | 3 | 3 |
| • Promote constructive applications | 3 | 3 | 3 |
- Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties.
I would prioritize library standardization. All operations, from that educators to users, can be facilitated from having a common reference frame. Further, among the options, this appears the most neutral. Overall, proper security requires practitioners from both sides of the spectrum: training in protecting and disrupting systems, and being willing to document for the good of the community.
Homework
Answers for Homework Questions from Professor Jabson:
- Error rate - 1: 10^6. This is dwarfed by the length of the human genome. The body has numerous selectivity and repair mechanisms to deal with mitigate issues from mutations.
- 64 / Some codes might not work due to chemical incompatibility and structural reasons.
Answers for Homework Questions from Dr. LeProust:
- Phosphoramidite DNA Synthesis Cycle
- Increasing inefficiency of reactions with greater length
- Extremely high inefficiency Error rates
Answers for Homework Question from George Church:
- Arginine, histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, and valine
I’m still forming my thoughts on how this affects my view of the Lysine Contingency.
Week 2 HW: DNA Read, Write, and Edit
Part 0: Basics of Gel Electrophoresis
[This was a pure watch session. Thus there’s nothing to add here.]
Part 1: Benchling & In-silico Gel Art
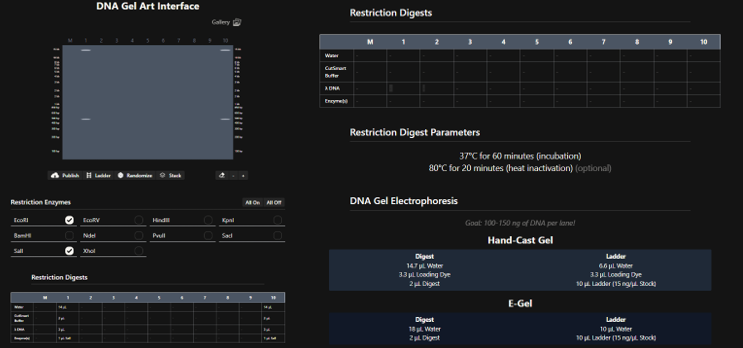
Restriction Enzymes Simulated on Lambda_NEB: EcoRI, HindIII, BamHI, KpnI, EcoRV, SacI, and SalI
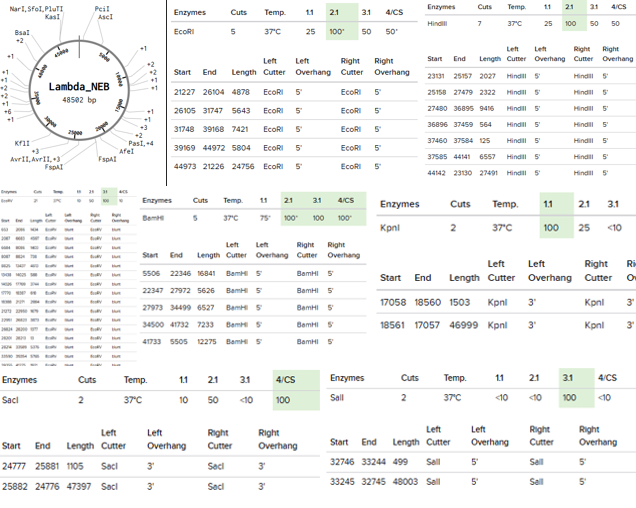

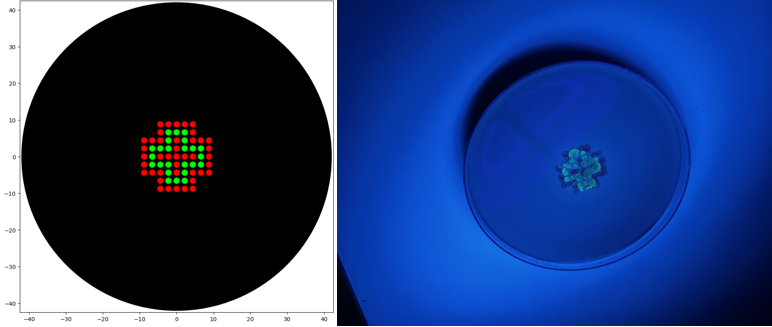
Simple Art produced by way of Paul Vanouse’s Latent Figure Protocol artworks through the use of RC Donovan’s Gel Art Iteration Tool (https://rcdonovan.com/gel-art):

“4 corners”, using EcoRI and SalI in Lanes 1 and 10.
Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
[This leaned on Wetlab Access. As a virtual student, this was not required.]
Part 3: DNA Design Challenge
3.1. Choose your protein.
Regarding proteins, I chose U-box domain-containing protein 12, also known as Plant U-box protein 12 or RING-type E3 ubiquitin transferase PUB12. I wanted to start with something and keep things relatively simple.
sp|Q9ZV31|PUB12_ARATH U-box domain-containing protein 12 OS=Arabidopsis thaliana OX=3702 GN=PUB12 PE=2 SV=1
MAKSEKHKLAQTLIDSINEIASISDSVTPMKKHCANLSRRLSLLLPMLEEIRDNQESSSE
VVNALLSVKQSLLHAKDLLSFVSHVSKIYLVLERDQVMVKFQKVTSLLEQALSIIPYENL
EISDELKEQVELVLVQLRRSLGKRGGDVYDDELYKDVLSLYSGRGSVMESDMVRRVAEKL
QLMTITDLTQESLALLDMVSSSGGDDPGESFEKMSMVLKKIKDFVQTYNPNLDDAPLRLK
SSLPKSRDDDRDMLIPPEEFRCPISLELMTDPVIVSSGQTYERECIKKWLEGGHLTCPKT
QETLTSDIMTPNYVLRSLIAQWCESNGIEPPKRPNISQPSSKASSSSSAPDDEHNKIEEL
LLKLTSQQPEDRRSAAGEIRLLAKQNNHNRVAIAASGAIPLLVNLLTISNDSRTQEHAVT
SILNLSICQENKGKIVYSSGAVPGIVHVLQKGSMEARENAAATLFSLSVIDENKVTIGAA
GAIPPLVTLLSEGSQRGKKDAATALFNLCIFQGNKGKAVRAGLVPVLMRLLTEPESGMVD
ESLSILAILSSHPDGKSEVGAADAVPVLVDFIRSGSPRNKENSAAVLVHLCSWNQQHLIE
AQKLGIMDLLIEMAENGTDRGKRKAAQLLNRFSRFNDQQKQHSGLGLEDQISLI
Site: https://rest.uniprot.org/uniprotkb/Q9ZV31.fasta
Base site: https://www.uniprot.org/uniprotkb/Q9ZV31/entry
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
Reverse Translation Tool – BCCM – GeneCorner (https://www.genecorner.ugent.be/rev_trans.html)
PUB12, a plant U-box–type E3 ubiquitin ligase DNA sequence
(1) >reverse translation of sp|Q9ZV31|PUB12_ARATH U-box domain-containing protein 12 OS=Arabidopsis thaliana OX=3702 GN=PUB12 PE=2 SV=1 to a 1962 base sequence of most likely codons.
atggcgaaaagcgaaaaacataaactggcgcagaccctgattgatagcattaacgaaatt
gcgagcattagcgatagcgtgaccccgatgaaaaaacattgcgcgaacctgagccgccgc
ctgagcctgctgctgccgatgctggaagaaattcgcgataaccaggaaagcagcagcgaa
gtggtgaacgcgctgctgagcgtgaaacagagcctgctgcatgcgaaagatctgctgagc
tttgtgagccatgtgagcaaaatttatctggtgctggaacgcgatcaggtgatggtgaaa
tttcagaaagtgaccagcctgctggaacaggcgctgagcattattccgtatgaaaacctg
gaaattagcgatgaactgaaagaacaggtggaactggtgctggtgcagctgcgccgcagc
ctgggcaaacgcggcggcgatgtgtatgatgatgaactgtataaagatgtgctgagcctg
tatagcggccgcggcagcgtgatggaaagcgatatggtgcgccgcgtggcggaaaaactg
cagctgatgaccattaccgatctgacccaggaaagcctggcgctgctggatatggtgagc
agcagcggcggcgatgatccgggcgaaagctttgaaaaaatgagcatggtgctgaaaaaa
attaaagattttgtgcagacctataacccgaacctggatgatgcgccgctgcgcctgaaa
agcagcctgccgaaaagccgcgatgatgatcgcgatatgctgattccgccggaagaattt
cgctgcccgattagcctggaactgatgaccgatccggtgattgtgagcagcggccagacc
tatgaacgcgaatgcattaaaaaatggctggaaggcggccatctgacctgcccgaaaacc
caggaaaccctgaccagcgatattatgaccccgaactatgtgctgcgcagcctgattgcg
cagtggtgcgaaagcaacggcattgaaccgccgaaacgcccgaacattagccagccgagc
agcaaagcgagcagcagcagcagcgcgccggatgatgaacataacaaaattgaagaactg
ctgctgaaactgaccagccagcagccggaagatcgccgcagcgcggcgggcgaaattcgc
ctgctggcgaaacagaacaaccataaccgcgtggcgattgcggcgagcggcgcgattccg
ctgctggtgaacctgctgaccattagcaacgatagccgcacccaggaacatgcggtgacc
agcattctgaacctgagcatttgccaggaaaacaaaggcaaaattgtgtatagcagcggc
gcggtgccgggcattgtgcatgtgctgcagaaaggcagcatggaagcgcgcgaaaacgcg
gcggcgaccctgtttagcctgagcgtgattgatgaaaacaaagtgaccattggcgcggcg
ggcgcgattccgccgctggtgaccctgctgagcgaaggcagccagcgcggcaaaaaagat
gcggcgaccgcgctgtttaacctgtgcatttttcagggcaacaaaggcaaagcggtgcgc
gcgggcctggtgccggtgctgatgcgcctgctgaccgaaccggaaagcggcatggtggat
gaaagcctgagcattctggcgattctgagcagccatccggatggcaaaagcgaagtgggc
gcggcggatgcggtgccggtgctggtggattttattcgcagcggcagcccgcgcaacaaa
gaaaacagcgcggcggtgctggtgcatctgtgcagctggaaccagcagcatctgattgaa
gcgcagaaactgggcattatggatctgctgattgaaatggcggaaaacggcaccgatcgc
ggcaaacgcaaagcggcgcagctgctgaaccgctttagccgctttaacgatcagcagaaa
cagcatagcggcctgggcctggaagatcagattagcctgatt
(2) >reverse translation of sp|Q9ZV31|PUB12_ARATH U-box domain-containing protein 12 OS=Arabidopsis thaliana OX=3702 GN=PUB12 PE=2 SV=1 to a 1962 base sequence of consensus codons.
atggcnaarwsngaraarcayaarytngcncaracnytnathgaywsnathaaygarath
gcnwsnathwsngaywsngtnacnccnatgaaraarcaytgygcnaayytnwsnmgnmgn
ytnwsnytnytnytnccnatgytngargarathmgngayaaycargarwsnwsnwsngar
gtngtnaaygcnytnytnwsngtnaarcarwsnytnytncaygcnaargayytnytnwsn
ttygtnwsncaygtnwsnaarathtayytngtnytngarmgngaycargtnatggtnaar
ttycaraargtnacnwsnytnytngarcargcnytnwsnathathccntaygaraayytn
garathwsngaygarytnaargarcargtngarytngtnytngtncarytnmgnmgnwsn
ytnggnaarmgnggnggngaygtntaygaygaygarytntayaargaygtnytnwsnytn
taywsnggnmgnggnwsngtnatggarwsngayatggtnmgnmgngtngcngaraarytn
carytnatgacnathacngayytnacncargarwsnytngcnytnytngayatggtnwsn
wsnwsnggnggngaygayccnggngarwsnttygaraaratgwsnatggtnytnaaraar
athaargayttygtncaracntayaayccnaayytngaygaygcnccnytnmgnytnaar
wsnwsnytnccnaarwsnmgngaygaygaymgngayatgytnathccnccngargartty
mgntgyccnathwsnytngarytnatgacngayccngtnathgtnwsnwsnggncaracn
taygarmgngartgyathaaraartggytngarggnggncayytnacntgyccnaaracn
cargaracnytnacnwsngayathatgacnccnaaytaygtnytnmgnwsnytnathgcn
cartggtgygarwsnaayggnathgarccnccnaarmgnccnaayathwsncarccnwsn
wsnaargcnwsnwsnwsnwsnwsngcnccngaygaygarcayaayaarathgargarytn
ytnytnaarytnacnwsncarcarccngargaymgnmgnwsngcngcnggngarathmgn
ytnytngcnaarcaraayaaycayaaymgngtngcnathgcngcnwsnggngcnathccn
ytnytngtnaayytnytnacnathwsnaaygaywsnmgnacncargarcaygcngtnacn
wsnathytnaayytnwsnathtgycargaraayaarggnaarathgtntaywsnwsnggn
gcngtnccnggnathgtncaygtnytncaraarggnwsnatggargcnmgngaraaygcn
gcngcnacnytnttywsnytnwsngtnathgaygaraayaargtnacnathggngcngcn
ggngcnathccnccnytngtnacnytnytnwsngarggnwsncarmgnggnaaraargay
gcngcnacngcnytnttyaayytntgyathttycarggnaayaarggnaargcngtnmgn
gcnggnytngtnccngtnytnatgmgnytnytnacngarccngarwsnggnatggtngay
garwsnytnwsnathytngcnathytnwsnwsncayccngayggnaarwsngargtnggn
gcngcngaygcngtnccngtnytngtngayttyathmgnwsnggnwsnccnmgnaayaar
garaaywsngcngcngtnytngtncayytntgywsntggaaycarcarcayytnathgar
gcncaraarytnggnathatggayytnytnathgaratggcngaraayggnacngaymgn
ggnaarmgnaargcngcncarytnytnaaymgnttywsnmgnttyaaygaycarcaraar
carcaywsnggnytnggnytngargaycarathwsnytnath
3.3. Codon optimization.
Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage.
- Optimizing codon usage can help account for host bias as well as reduce the pool of downstream errors to account for, as well as save materials and time.
Which organism have you chosen to optimize the codon sequence for and why?
Which: Arabidopsis thaliana
Why: I wanted to start somewhere, and this organism is familiar.
Optimization Tool Used: Vector Builder (https://en.vectorbuilder.com/tool/codon-optimization.html )
Chose to use the first sequence of the two. Enzyme recognition sites avoided included BsaI and BbsI.
Resulting Sequence is as follows:
PUB12, a plant U-box–type E3 ubiquitin ligase DNA sequence
ATGGCTAAATCTGAGAAGCATAAGTTGGCTCAGACTCTCATCGATTCTATAAATGAAATTGCTTCTATCTCAGATTCAGTTACTCCAATGAAGAAGCATTGTGCAAATTTGTCTAGGAGATTGTCACTTCTTCTTCCAATGTTGGAAGAGATTAGAGATAATCAAGAGTCTAGCTCTGAAGTCGTGAACGCTTTGCTCTCAGTTAAACAATCTTTATTACATGCTAAGGATCTCTTGTCTTTCGTCAGTCATGTGAGCAAGATATATCTTGTTCTTGAGAGAGATCAAGTGATGGTTAAGTTTCAAAAAGTTACTAGCCTTCTTGAGCAAGCTCTTTCTATAATCCCTTATGAGAATCTTGAAATTTCTGATGAATTGAAAGAACAAGTTGAATTGGTTCTTGTTCAACTTAGAAGATCTTTGGGTAAGAGAGGTGGTGATGTTTACGATGATGAACTTTATAAGGATGTTCTTTCACTTTACAGTGGAAGAGGATCAGTTATGGAGAGTGATATGGTTCGTCGAGTTGCCGAGAAATTGCAACTAATGACTATCACTGATTTGACACAAGAGTCTCTTGCTCTTCTTGATATGGTTTCTTCTAGTGGTGGAGATGATCCTGGAGAGTCATTCGAAAAGATGTCTATGGTTCTTAAGAAAATTAAGGATTTCGTTCAAACCTATAATCCTAACCTAGATGACGCTCCCCTTAGACTTAAATCATCATTGCCTAAATCGAGAGATGATGATCGTGATATGCTTATTCCACCTGAAGAATTCCGTTGTCCTATTTCGCTTGAGCTTATGACTGATCCTGTAATCGTTTCTTCAGGTCAAACCTATGAAAGAGAGTGTATTAAGAAGTGGCTTGAAGGAGGACATTTGACATGTCCTAAGACTCAAGAAACTTTGACATCTGATATCATGACCCCTAATTATGTTCTTAGATCTTTGATCGCTCAATGGTGTGAGTCGAATGGAATCGAGCCTCCAAAGAGGCCAAACATAAGTCAGCCTTCTAGTAAGGCTTCTTCATCATCTAGTGCTCCTGATGACGAACATAATAAGATCGAAGAATTGCTCTTGAAGTTGACTTCTCAGCAACCTGAAGATAGAAGATCCGCTGCTGGAGAGATCAGACTTTTGGCCAAACAAAACAACCATAACAGAGTTGCTATCGCTGCTTCAGGAGCTATTCCACTCTTGGTGAACCTTTTGACTATCTCAAACGATTCCAGAACACAAGAGCATGCTGTTACGTCTATCCTCAACCTTTCTATCTGCCAAGAAAATAAAGGTAAGATCGTTTATTCTAGTGGTGCAGTGCCTGGTATTGTTCATGTTTTGCAGAAGGGATCAATGGAGGCTAGAGAAAACGCTGCTGCTACTCTTTTCTCTCTTTCCGTTATAGATGAGAATAAGGTTACTATTGGAGCTGCTGGAGCAATTCCACCTTTGGTTACTCTCCTTTCTGAAGGATCACAGCGTGGAAAGAAGGATGCTGCTACTGCACTCTTCAACCTTTGTATCTTTCAGGGTAATAAAGGTAAGGCAGTTAGAGCAGGACTTGTGCCTGTGCTTATGAGGCTTTTGACTGAACCTGAATCTGGAATGGTTGATGAGAGCCTTTCTATTCTTGCTATTCTTTCTTCTCATCCAGACGGAAAGTCTGAAGTTGGAGCTGCTGATGCAGTTCCTGTTCTTGTTGATTTCATCAGATCTGGATCTCCTAGAAATAAGGAGAATTCTGCTGCAGTTCTTGTTCACTTGTGTTCATGGAATCAACAACATCTTATCGAAGCACAGAAGCTTGGAATCATGGATCTTCTCATCGAGATGGCTGAAAACGGAACTGATCGTGGTAAGAGAAAGGCCGCACAATTGCTTAATAGATTTTCTAGATTTAACGATCAGCAGAAGCAACACAGTGGTCTTGGTCTTGAAGATCAAATTTCATTGATT
Below is a printout contrasting the two:
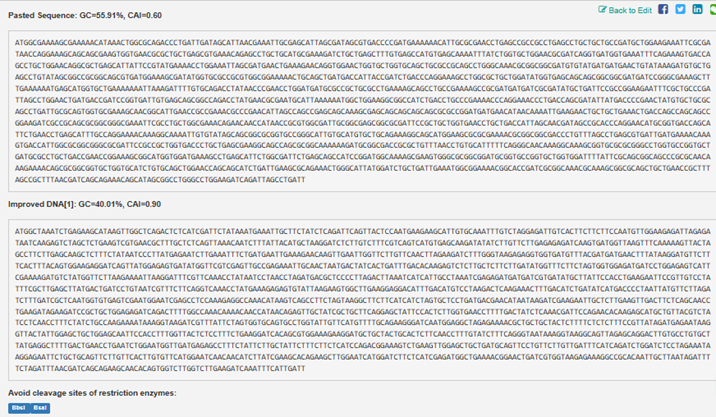

3.4. You have a sequence! Now what?
Recombinant DNA technologies could be utilized to make this protein from the DNA (Cell-free or Cell-dependent). Respectively they involve either special mix that can take place in a test tube or through using a live cell’s machinery to produce the protein.
Part 4: Prepare a Twist DNA Synthesis Order
Build Your DNA Insert Sequence
I prepared the above improved sequence as a test order. Below is the initial step through the creation of the DNA/RNA Sequence in Benchling with a Linear Topology.
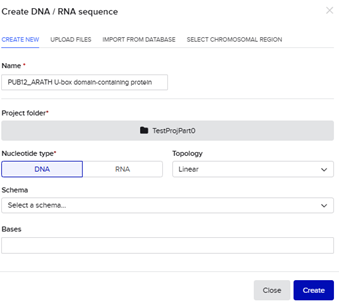

It was annotated below as such within Benchling before a linear map and file was constructed that could be uploaded to Twist Bio.
Start Codon: ATG
Coding Sequence:
ATGGCTAAATCTGAGAAGCATAAGTTGGCTCAGACTCTCATCGATTCTATAAATGAAATTGCTTCTATCTCAGATTCAGTTACTCCAATGAAGAAGCATTGTGCAAATTTGTCTAGGAGATTGTCACTTCTTCTTCCAATGTTGGAAGAGATTAGAGATAATCAAGAGTCTAGCTCTGAAGTCGTGAACGCTTTGCTCTCAGTTAAACAATCTTTATTACATGCTAAGGATCTCTTGTCTTTCGTCAGTCATGTGAGCAAGATATATCTTGTTCTTGAGAGAGATCAAGTGATGGTTAAGTTTCAAAAAGTTACTAGCCTTCTTGAGCAAGCTCTTTCTATAATCCCTTATGAGAATCTTGAAATTTCTGATGAATTGAAAGAACAAGTTGAATTGGTTCTTGTTCAACTTAGAAGATCTTTGGGTAAGAGAGGTGGTGATGTTTACGATGATGAACTTTATAAGGATGTTCTTTCACTTTACAGTGGAAGAGGATCAGTTATGGAGAGTGATATGGTTCGTCGAGTTGCCGAGAAATTGCAACTAATGACTATCACTGATTTGACACAAGAGTCTCTTGCTCTTCTTGATATGGTTTCTTCTAGTGGTGGAGATGATCCTGGAGAGTCATTCGAAAAGATGTCTATGGTTCTTAAGAAAATTAAGGATTTCGTTCAAACCTATAATCCTAACCTAGATGACGCTCCCCTTAGACTTAAATCATCATTGCCTAAATCGAGAGATGATGATCGTGATATGCTTATTCCACCTGAAGAATTCCGTTGTCCTATTTCGCTTGAGCTTATGACTGATCCTGTAATCGTTTCTTCAGGTCAAACCTATGAAAGAGAGTGTATTAAGAAGTGGCTTGAAGGAGGACATTTGACATGTCCTAAGACTCAAGAAACTTTGACATCTGATATCATGACCCCTAATTATGTTCTTAGATCTTTGATCGCTCAATGGTGTGAGTCGAATGGAATCGAGCCTCCAAAGAGGCCAAACATAAGTCAGCCTTCTAGTAAGGCTTCTTCATCATCTAGTGCTCCTGATGACGAACATAATAAGATCGAAGAATTGCTCTTGAAGTTGACTTCTCAGCAACCTGAAGATAGAAGATCCGCTGCTGGAGAGATCAGACTTTTGGCCAAACAAAACAACCATAACAGAGTTGCTATCGCTGCTTCAGGAGCTATTCCACTCTTGGTGAACCTTTTGACTATCTCAAACGATTCCAGAACACAAGAGCATGCTGTTACGTCTATCCTCAACCTTTCTATCTGCCAAGAAAATAAAGGTAAGATCGTTTATTCTAGTGGTGCAGTGCCTGGTATTGTTCATGTTTTGCAGAAGGGATCAATGGAGGCTAGAGAAAACGCTGCTGCTACTCTTTTCTCTCTTTCCGTTATAGATGAGAATAAGGTTACTATTGGAGCTGCTGGAGCAATTCCACCTTTGGTTACTCTCCTTTCTGAAGGATCACAGCGTGGAAAGAAGGATGCTGCTACTGCACTCTTCAACCTTTGTATCTTTCAGGGTAATAAAGGTAAGGCAGTTAGAGCAGGACTTGTGCCTGTGCTTATGAGGCTTTTGACTGAACCTGAATCTGGAATGGTTGATGAGAGCCTTTCTATTCTTGCTATTCTTTCTTCTCATCCAGACGGAAAGTCTGAAGTTGGAGCTGCTGATGCAGTTCCTGTTCTTGTTGATTTCATCAGATCTGGATCTCCTAGAAATAAGGAGAATTCTGCTGCAGTTCTTGTTCACTTGTGTTCATGGAATCAACAACATCTTATCGAAGCACAGAAGCTTGGAATCATGGATCTTCTCATCGAGATGGCTGAAAACGGAACTGATCGTGGTAAGAGAAAGGCCGCACAATTGCTTAATAGATTTTCTAG
Stop Codon: TAG
Linear Map:
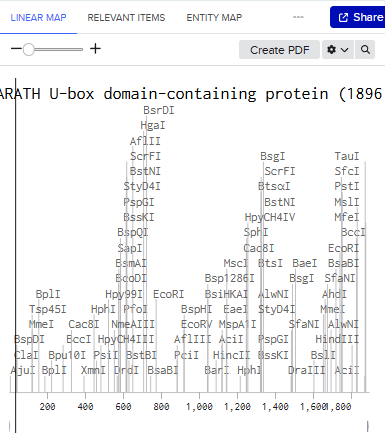

Here’s an example of what you just annotated in Benchling:
Sequence Import and Quote Obtained
The pTwist Amp High Copy: pTwist Amp Vector was chosen after the Clonal Gene Choice was pursued. The quote is to the left.. The annotated sequence page from TwistBio from which a GenBank construct file was downloaded is to the right.
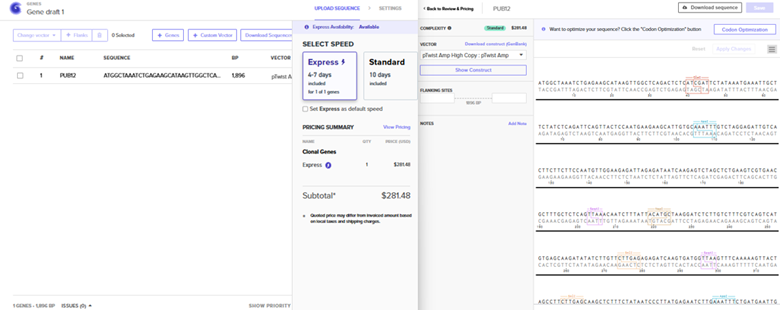

The construct was imported into Benchling to yield the plasmid below.
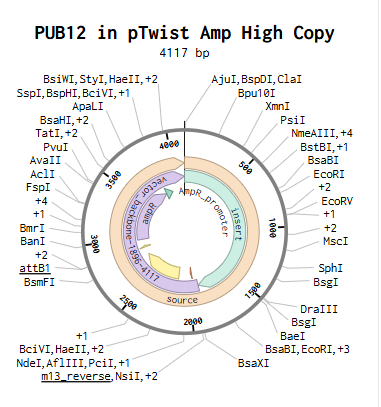


Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
- I might want to sequence large viruses, particularly out of sheer curiosity for the instructions that allow them to exist in their current forms.
Sources of interest:
Piacente, F., De Castro, C., Jeudy, S., Molinaro, A., Salis, A., Damonte, G., Bernardi, C., Abergel, C. and Tonetti, M.G., 2014. Giant virus Megavirus chilensis encodes the biosynthetic pathway for uncommon acetamido sugars. Journal of Biological Chemistry, 289(35), pp.24428-24439.
Legendre, M., Arslan, D., Abergel, C. and Claverie, J.M., 2012. Genomics of Megavirus and the elusive fourth domain of Life. Communicative & integrative biology, 5(1), pp.102-106.
Arslan, D., Legendre, M., Seltzer, V., Abergel, C. and Claverie, J.M., 2011. Distant Mimivirus relative with a larger genome highlights the fundamental features of Megaviridae. Proceedings of the National Academy of Sciences, 108(42), pp.17486-17491.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
- I would borrow from the methods used in the aforementioned literature, particularly “454-titanium and Illumina HiSeq approaches”. These methods appear adequate. Lack of a priori knowledge of the genome or genomic features not being required is helpful, in addition to single-nucleotide resolution, higher dynamic range, and less DNA/RNA needed.
https://www.ebi.ac.uk/training/online/courses/functional-genomics-ii-common-technologies-and-data-analysis-methods/next-generation-sequencing/
Also answer the following questions:
Is your method first-, second- or third-generation or other? How so?
- Second-generation. They engage massively parallel sequencing.
What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
- Library Preparation (fragmenting of DNA and adapters added to both ends of DNA for amplification)
- Sequencing
- Data Analysis and Cleanup
What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
Simplified:
- The DNA strand gets color coded with fluorescent terminators
- Images are taken of the flow cell after each letter is added
- Software determines the bases based on color intensities
- Calls are made, corrections are issued, and output is cleaned up
Source of interest: https://genohub.com/bioinformatics/10/base-calling
What is the output of your chosen sequencing technology?
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I am not sure yet, but I am leaning towards DNA origami art to experiment with the medium and explore versatility of applications.
Sources of interest:
Bush, J., Singh, S., Vargas, M., Oktay, E., Hu, C.H. and Veneziano, R., 2020. Synthesis of DNA origami scaffolds: Current and emerging strategies. Molecules, 25(15), p.3386.
Weck, J.M. and Heuer-Jungemann, A., 2025. Fully addressable designer superstructures assembled from one single modular DNA origami. Nature communications, 16(1), p.1556.
DNA origami by Paul W. K. Rothemund, California Institute of Technology, 2004. 100 nanometers in diameter.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
For validating the structures, if cost didn’t matter, I would consider using next generation sequencing (Illumina, for both sequences of the staples and scaffold) and Atomic Force Microscopy (Visual, especially confirming folds)
Also answer the following questions:
The essential steps of the chosen sequencing methods would be:
- Library Prep (DNA Fragmentation and Adapter ligation methods)
- Cluster Generation via amplification
- Sequencing and base calling
What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
- The major limitations would be cost, error rates, and short read lengths. This would not be efficient to scale as is.
Improvements would involve:
-Hand design of patterns (for ideating improvements)
-Computer design and optimization of material usage
-Production of material and strand-routing precision
5.3 DNA Edit
(i) What DNA would you want to edit and why?
DNA edits that I would like to perform would be those that allow for the minimization and or elimination of metabolic disease states. The why comes down to the quality-of-life improvements for all involved.
(ii) What technology or technologies would you use to perform these DNA edits and why?
Also answer the following questions:
How does your technology of choice edit DNA? What are the essential steps?
- That, I’m still thinking about. While some gene therapies have been successfully – my mind shifts to which exact technology is the “right” choice, along with the ethical hurdles needed to investigate such properly.
What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?-
What are the limitations of your editing methods (if any) in terms of efficiency or precision?
Week 3 HW: Lab Automation
- Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
DeRoo, J.B., Jones, A.A., Slaughter, C.K., Ahr, T.W., Stroup, S.M., Thompson, G.B. and Snow, C.D., 2025. Automation of protein crystallization scaleup via Opentrons-2 liquid handling. SLAS technology, 32, p.100268.
https://doi.org/10.1016/j.slast.2025.100268
This work describes an approach by which an Opentrons-2 liquid handling robot was used for automating sitting drop protein crystallization trials. This ability also improve comparability of products produced, improving studies that depend on their proper manufacture. An important detail is how the Opentrons-2 can prove a cost-effective option for laboratory operations. For example, at the time of writing, the Opentrons-2 can be purchased for around 13.5K USD vs that of a Gryphon machine at around 65K USD.
- Write a description about what you intend to do with automation tools for your final project.
I’m still forming my thoughts about how I want to effectively use automation tools for my final project.
So far, I am interested in branching off from example #2 given in the Homework and the above example, regarding screening an array of designed biosensor constructs.
One idea had in mind was towards a digital tracing project that revolves around said constructs used to track known entities.
Simply, products are given a unique ID with stored parameters. These are linked within a automation run so that each product is trackable as they are processed. One application that is probably already in use but would be fun to adapt towards something not already applied would be swappable combined wearable crystallized biosensors that are traded in daily for workers that are liable to be exposed to a particular organism and pollutant pairs.
I could use an Echo for transfer of nano-scale components.
The Bravo or Opentrons-2 could be used for precise, automated pipetting ,especially of the crystals.
The multiflow would be used to dispense the larger scale volume components.
The PlateLoc would be helful for sealing the plates.
The inheco could be used for controlled incubation.
The Xpeel would be used for careful desealing of the plates.
Finally, the PHERAstar could be used for reading fluorescence outputs.
Still developing this out from this branch.
Week 4 HW: Protein Design Part I
Part A: Conceptual Questions
We were allowed answer 9 out of 11 of the following questions:
- How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
I expect answers to vary since meat composition vary.
Water composition of meat is estimated to be about 65-80% meat.
Some estimates have proteins make up around 1/5 to less than 1/3 of overall muscle tissue.
We’ll work with 20% and 30% for upper and lower bounds for simplicity.
Assuming that 100 Daltons roughly equals 100 g/mo
Lower: 500 g × 0.2 = 100 g protein = 1 mol. 6.02 × 10²³ molecules
Upper: 500 g × 0.3 = 150 g protein = 1.5 mol. ~9 × 10²³ molecules
Quick Sources:
https://redtablemeats.com/fresh-meat/beef/what-percentage-of-beef-is-protein/
https://agriculture.institute/fresh-meat-technology/water-content-in-meat-importance/
For simplicity, let’s work with Taking the average, 500 grams of meat may yield
- Why do humans eat beef but do not become a cow, eat fish but do not become fish?
These products are broken down through digestion and recycled. Our genetic material largely informs the building process of new material.
- Why are there only 20 natural amino acids?
I’m not convinced that those are the only ones, in so far of what we have observed on Earth it is likely that the developmental “cost” as per our development over time was too high. 20 reflects optimization, no?
- Can you make other non-natural amino acids? Design some new amino acids.
Yes. These can be designed through new side chains, although stability and inclusion may be a problem.
- Where did amino acids come from before enzymes that make them, and before life started?
Abiotic processes. Some sources list thermal vents and delivery through meteorites.
Ehrenfreund, P., Bernstein, M.P., Dworkin, J.P., Sandford, S.A. and Allamandola, L.J., 2001. The photostability of amino acids in space. The Astrophysical Journal Letters, 550(1), pp.L95-L99.
Kobayashi, K., Mita, H., Kebukawa, Y., Nakagawa, K., Kaneko, T., Obayashi, Y., Sato, T., Yokoo, T., Minematsu, S., Fukuda, H. and Oguri, Y., 2021. Space exposure of amino acids and their precursors during the Tanpopo Mission. Astrobiology, 21(12), pp.1479-1493.
- If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
Left-handed
- Can you discover additional helices in proteins?
Yes
- Why are most molecular helices right-handed?
The stereochemistry of L-amino acids constrains backbone geometry in a way that right-handed helices are lower in energy. Right-handed chirality is favored and chirality from the bottom influences chirality at higher structures.
- Why do β-sheets tend to aggregate?
What is the driving force for β-sheet aggregation?
[Skipped]
- Why do many amyloid diseases form β-sheets?
β-Sheet structures are stable and can easy to produce from misfolding.
- Can you use amyloid β-sheets as materials? Design a β-sheet motif that forms a well-ordered structure.
[Skipped]
Part B: Protein Analysis and Visualization
The protein that I selected is U-box domain-containing protein 12.Possesses E3 ubiquitin-protein ligase in vitro. I was not too particular on its selection and wanted to start somewhere.
Source: https://www.uniprot.org/uniprotkb/Q5VRH9/entry
Oryza sativa subsp. japonica (Rice)
Amino Acid sequence:
sp|Q5VRH9|PUB12_ORYSJ U-box domain-containing protein 12 OS=Oryza sativa subsp. japonica OX=39947 GN=PUB12 PE=1 SV=1
MPKRVADEIAALPEPRGPLRRPCADLSRRVRLLAPLLDHLPASSSSSSSTPLADALGAAR
DLLRKTRDGSKIDQAMRGDAFLDEFAGVNRQIHLALDALPYNTFHMPQEVQEQVALVHSQ
FQRASTRTDPPDTQLSMDLAWALTDNPSDPALLTRISHKLQLHTMADMKNESIALHNMVI
STAGEPDGCVDQMSSLLKKLKDCVVTEDHANDALTTRSASIKHRSPIIPDEFRCPISLEL
MQDPVIVSSGQTYERSCIQKWLDSGHKTCPKTQQPLSHTSLTPNFVLKSLISQWCEANGI
ELPKNKQNSRDKKAAKSSDYDHAGLVSLMNRLRSGNQDEQRAAAGEIRLLAKRNVNNRIC
IAEAGAIPLLVNLLSSSDPRTQEHAVTALLNLSIHENNKASIVDSHAIPKIVEVLKTGSM
ETRENAAATLFSLSVVDENKVTIGAAGAIPPLINLLCDGSPRGKKDAATAIFNLCIYQGN
KVRAVKAGIVIHLMNFLVDPTGGMIDEALSLLSILAGNPEGKIVIARSEPIPPLVEVIKT
GSPRNRENAAAILWLLCSADTEQTLAAKAAGVEDALKELSETGTDRAKRKASSILELMHQ
ANEDSLKGNGH
The length of the protein is: 611 aminoacids.
The most common amino acid is: L, which appears 72 times.
This protein does not share 100% homology with any other although there are many with 50-90% homology, per UniProt. Via the Blast Tool, a lot of homologs were found.
Query= EMBOSS_001
Length=611
Score E
Sequences producing significant alignments: (Bits) Value
SP:Q5VRH9 PUB12_ORYSJ U-box domain-containing protein 12 OS=Oryza … 1199 0.0
TR:I1PYP8 I1PYP8_ORYGL U-box domain-containing protein 12 OS=Oryza… 1194 0.0
TR:A0A0D3GBR7 A0A0D3GBR7_9ORYZ RING-type E3 ubiquitin transferase … 1193 0.0
TR:A0A0E0PSC1 A0A0E0PSC1_ORYRU U-box domain-containing protein 12 … 1193 0.0
TR:A0A0E0HJJ2 A0A0E0HJJ2_ORYNI U-box domain-containing protein 12 … 1192 0.0
TR:A0A0E0A3S8 A0A0E0A3S8_9ORYZ U-box domain-containing protein 12 … 1182 0.0
SP:Q5VRH9-2 PUB12_ORYSJ Isoform 2 of U-box domain-containing prote… 1182 0.0
TR:A0A0D3GBR8 A0A0D3GBR8_9ORYZ RING-type E3 ubiquitin transferase … 1176 0.0
TR:A0A0E0PSC3 A0A0E0PSC3_ORYRU U-box domain-containing protein 12 … 1176 0.0
TR:A0A0E0HJJ3 A0A0E0HJJ3_ORYNI U-box domain-containing protein 12 … 1175 0.0
TR:A2Y893 A2Y893_ORYSI U-box domain-containing protein 12 OS=Oryza… 1167 0.0
TR:A0A0E0A3T5 A0A0E0A3T5_9ORYZ U-box domain-containing protein 12 … 1165 0.0
TR:A0A0E0DVD9 A0A0E0DVD9_9ORYZ U-box domain-containing protein 12 … 1165 0.0
TR:A0A0E0L6Q0 A0A0E0L6Q0_ORYPU U-box domain-containing protein 12 … 1152 0.0
TR:A0A6G1DGM2 A0A6G1DGM2_9ORYZ RING-type E3 ubiquitin transferase … 1128 0.0
TR:A0A0D9WKQ8 A0A0D9WKQ8_9ORYZ RING-type E3 ubiquitin transferase … 1102 0.0
TR:A0A0E0A3T4 A0A0E0A3T4_9ORYZ RING-type E3 ubiquitin transferase … 1009 0.0
TR:A0A5J9SK41 A0A5J9SK41_9POAL RING-type E3 ubiquitin transferase … 1005 0.0
TR:A0A2T7E463 A0A2T7E463_9POAL RING-type E3 ubiquitin transferase … 1000 0.0
TR:A0ABC9E206 A0ABC9E206_9POAL RING-type E3 ubiquitin transferase … 999 0.0
TR:A0A3L6RWE2 A0A3L6RWE2_PANMI RING-type E3 ubiquitin transferase … 999 0.0
TR:A0ABC9DBY8 A0ABC9DBY8_9POAL RING-type E3 ubiquitin transferase … 993 0.0
TR:A0A8T0T5H3 A0A8T0T5H3_PANVG RING-type E3 ubiquitin transferase … 993 0.0
TR:A0ABC9DW43 A0ABC9DW43_9POAL RING-type E3 ubiquitin transferase … 991 0.0
TR:A0A1E5WM86 A0A1E5WM86_9POAL RING-type E3 ubiquitin transferase … 991 0.0
TR:A0A811SD54 A0A811SD54_9POAL RING-type E3 ubiquitin transferase … 989 0.0
TR:A0A8T0TI26 A0A8T0TI26_PANVG RING-type E3 ubiquitin transferase … 988 0.0
TR:A0A811S773 A0A811S773_9POAL RING-type E3 ubiquitin transferase … 987 0.0
TR:A0A835KU84 A0A835KU84_9POAL RING-type E3 ubiquitin transferase … 986 0.0
TR:A0AAV5CS17 A0AAV5CS17_ELECO U-box domain-containing protein 12 … 986 0.0
TR:J3MAH2 J3MAH2_ORYBR U-box domain-containing protein 12 OS=Oryza… 985 0.0
TR:C5Z221 C5Z221_SORBI RING-type E3 ubiquitin transferase OS=Sorgh… 984 0.0
TR:A0A835FWE8 A0A835FWE8_9POAL RING-type E3 ubiquitin transferase … 983 0.0
TR:I1H245 I1H245_BRADI U-box domain-containing protein 12 OS=Brach… 983 0.0
TR:A0A3L6PH60 A0A3L6PH60_PANMI RING-type E3 ubiquitin transferase … 982 0.0
TR:A0A4U6UXZ9 A0A4U6UXZ9_SETVI U-box domain-containing protein 12 … 981 0.0
TR:B6SXY9 B6SXY9_MAIZE RING-type E3 ubiquitin transferase OS=Zea m… 964 0.0
TR:A0ACD5Y0V4 A0ACD5Y0V4_AVESA Uncharacterized protein OS=Avena sa… 957 0.0
TR:A0A3B6THH5 A0A3B6THH5_WHEAT RING-type E3 ubiquitin transferase … 956 0.0
TR:A0A8R7RDC8 A0A8R7RDC8_TRIUA RING-type E3 ubiquitin transferase … 955 0.0
TR:A0ACD6A429 A0ACD6A429_AVESA Uncharacterized protein OS=Avena sa… 950 0.0
TR:A0A8I6Y9M9 A0A8I6Y9M9_HORVV RING-type E3 ubiquitin transferase … 942 0.0
TR:A0A3B6I3Q2 A0A3B6I3Q2_WHEAT U-box domain-containing protein 12 … 917 0.0
TR:A0AAV5F2R6 A0AAV5F2R6_ELECO U-box domain-containing protein 12 … 914 0.0
TR:K3XW51 K3XW51_SETIT U-box domain-containing protein 12 OS=Setar… 905 0.0
TR:A0A0E0L6Q1 A0A0E0L6Q1_ORYPU U-box domain-containing protein 12 … 900 0.0
TR:A0A452XGN3 A0A452XGN3_AEGTS RING-type E3 ubiquitin transferase … 888 0.0
TR:A0A3B6R6B6 A0A3B6R6B6_WHEAT RING-type E3 ubiquitin transferase … 868 0.0
TR:A0A9R0Z145 A0A9R0Z145_TRITD RING-type E3 ubiquitin transferase … 866 0.0
TR:A0ACD5ZS62 A0ACD5ZS62_AVESA Uncharacterized protein OS=Avena sa… 843 0.0
TR:A0A452XGJ2 A0A452XGJ2_AEGTS RING-type E3 ubiquitin transferase … 836 0.0
TR:A0A0E0DVD6 A0A0E0DVD6_9ORYZ RING-type E3 ubiquitin transferase … 815 0.0
TR:A0A6I9S1S1 A0A6I9S1S1_ELAGV RING-type E3 ubiquitin transferase … 795 0.0
TR:A0A8B7C1E5 A0A8B7C1E5_PHODC RING-type E3 ubiquitin transferase … 786 0.0
TR:A0AAV8GY30 A0AAV8GY30_9POAL RING-type E3 ubiquitin transferase … 785 0.0
TR:A0AAV8EMN4 A0AAV8EMN4_9POAL RING-type E3 ubiquitin transferase … 783 0.0
TR:A0A8K0IKK4 A0A8K0IKK4_COCNU RING-type E3 ubiquitin transferase … 782 0.0
TR:A0AAV8F803 A0AAV8F803_9POAL RING-type E3 ubiquitin transferase … 781 0.0
TR:A0A8K0N659 A0A8K0N659_COCNU RING-type E3 ubiquitin transferase … 780 0.0
TR:A0A6I9SJB7 A0A6I9SJB7_ELAGV RING-type E3 ubiquitin transferase … 779 0.0
TR:A0A8B8Z9X0 A0A8B8Z9X0_PHODC U-box domain-containing protein 12 … 778 0.0
TR:A0A9Q0CB42 A0A9Q0CB42_9POAL RING-type E3 ubiquitin transferase … 777 0.0
TR:A0AAD5WDJ8 A0AAD5WDJ8_9POAL RING-type E3 ubiquitin transferase … 776 0.0
TR:A0A452XGQ0 A0A452XGQ0_AEGTS RING-type E3 ubiquitin transferase … 773 0.0
TR:A0A2I0BCA5 A0A2I0BCA5_9ASPA RING-type E3 ubiquitin transferase … 757 0.0
TR:A0A3B6TG51 A0A3B6TG51_WHEAT RING-type E3 ubiquitin transferase … 756 0.0
TR:A0A835RLE0 A0A835RLE0_VANPL RING-type E3 ubiquitin transferase … 755 0.0
TR:A0A9R0SRI3 A0A9R0SRI3_TRITD U-box domain-containing protein 12 … 754 0.0
TR:A0A6I9S1Y4 A0A6I9S1Y4_ELAGV RING-type E3 ubiquitin transferase … 753 0.0
TR:A0A833VES7 A0A833VES7_9POAL RING-type E3 ubiquitin transferase … 751 0.0
TR:A0A9R1BGT5 A0A9R1BGT5_TRITD RING-type E3 ubiquitin transferase … 751 0.0
TR:A0A9D5BZQ6 A0A9D5BZQ6_9LILI RING-type E3 ubiquitin transferase … 750 0.0
TR:A0ABD0UM27 A0ABD0UM27_DENTH RING-type E3 ubiquitin transferase … 746 0.0
TR:A0A8B8ZB12 A0A8B8ZB12_PHODC RING-type E3 ubiquitin transferase … 745 0.0
TR:A0A8T3AWN2 A0A8T3AWN2_DENNO RING-type E3 ubiquitin transferase … 744 0.0
TR:A0A2I0WEQ6 A0A2I0WEQ6_9ASPA RING-type E3 ubiquitin transferase … 743 0.0
TR:A0A6P5GVN1 A0A6P5GVN1_ANACO RING-type E3 ubiquitin transferase … 741 0.0
TR:A0A5P1FQ11 A0A5P1FQ11_ASPOF RING-type E3 ubiquitin transferase … 734 0.0
TR:A0AAP0G926 A0AAP0G926_9ASPA RING-type E3 ubiquitin transferase … 729 0.0
TR:A0AAP0WZL6 A0AAP0WZL6_LIQFO RING-type E3 ubiquitin transferase … 728 0.0
TR:A0A843WFY6 A0A843WFY6_COLES RING-type E3 ubiquitin transferase … 723 0.0
TR:A0ACB7WUI3 A0ACB7WUI3_DIOAL Zinc finger RING/FYVE/PHD-type prot… 723 0.0
TR:A0AB40BCZ8 A0AB40BCZ8_DIOCR RING-type E3 ubiquitin transferase … 723 0.0
TR:A0AAQ3K0H0 A0AAQ3K0H0_9LILI RING-type E3 ubiquitin transferase … 720 0.0
TR:F6HND8 F6HND8_VITVI RING-type E3 ubiquitin transferase OS=Vitis… 718 0.0
TR:A0ABY9D2S6 A0ABY9D2S6_VITVI RING-type E3 ubiquitin transferase … 718 0.0
TR:A0AAV7E209 A0AAV7E209_ARIFI RING-type E3 ubiquitin transferase … 717 0.0
TR:A0AA39DJS7 A0AA39DJS7_VITRO RING-type E3 ubiquitin transferase … 715 0.0
TR:A0ABS8SCS2 A0ABS8SCS2_DATST RING-type E3 ubiquitin transferase … 712 0.0
TR:A0AAV5LAD4 A0AAV5LAD4_9ROSI RING-type E3 ubiquitin transferase … 712 0.0
TR:V4TIZ6 V4TIZ6_CITCL RING-type E3 ubiquitin transferase OS=Citru… 710 0.0
TR:A0A804I6U7 A0A804I6U7_MUSAM RING-type E3 ubiquitin transferase … 710 0.0
TR:A0A200Q9A7 A0A200Q9A7_MACCD RING-type E3 ubiquitin transferase … 709 0.0
TR:A0A7J6W5Z9 A0A7J6W5Z9_THATH RING-type E3 ubiquitin transferase … 709 0.0
TR:A0ACB8L513 A0ACB8L513_CITSI U-box domain-containing protein 14 … 709 0.0
TR:A0A067E149 A0A067E149_CITSI RING-type E3 ubiquitin transferase … 709 0.0
TR:A0ACB9PUD9 A0ACB9PUD9_BAUVA Uncharacterized protein OS=Bauhinia… 707 0.0
TR:A0A9Q1MDB3 A0A9Q1MDB3_9SOLA RING-type E3 ubiquitin transferase … 707 0.0
TR:A0AAV8RJ11 A0AAV8RJ11_ENSVE RING-type E3 ubiquitin transferase … 707 0.0
TR:A0AAN8VNY8 A0AAN8VNY8_9MAGN RING-type E3 ubiquitin transferase … 706 0.0
TR:A0ABR2QF73 A0ABR2QF73_9ROSI RING-type E3 ubiquitin transferase … 706 0.0
TR:A0A061FPF5 A0A061FPF5_THECC RING-type E3 ubiquitin transferase … 706 0.0
TR:A0A6P5YZR2 A0A6P5YZR2_DURZI RING-type E3 ubiquitin transferase … 705 0.0
TR:A0A6A4QEE0 A0A6A4QEE0_LUPAL RING-type E3 ubiquitin transferase … 705 0.0
TR:A0A6I9TG78 A0A6I9TG78_SESIN RING-type E3 ubiquitin transferase … 705 0.0
TR:A0A835LLS2 A0A835LLS2_9MAGN RING-type E3 ubiquitin transferase … 705 0.0
TR:A0AAD4NZN1 A0AAD4NZN1_PERFH RING-type E3 ubiquitin transferase … 704 0.0
TR:A0ABR2S2R6 A0ABR2S2R6_9ROSI RING-type E3 ubiquitin transferase … 704 0.0
TR:A0AAE1RYG1 A0AAE1RYG1_9SOLA RING-type E3 ubiquitin transferase … 704 0.0
TR:A0A1J6IL89 A0A1J6IL89_NICAT RING-type E3 ubiquitin transferase … 704 0.0
TR:A0A9Q1M5W7 A0A9Q1M5W7_9SOLA RING-type E3 ubiquitin transferase … 704 0.0
TR:A0A6J1B7W8 A0A6J1B7W8_9ROSI RING-type E3 ubiquitin transferase … 704 0.0
TR:A0A9W7HPH7 A0A9W7HPH7_HIBTR RING-type E3 ubiquitin transferase … 703 0.0
TR:A0A9W7HNE6 A0A9W7HNE6_HIBTR RING-type E3 ubiquitin transferase … 703 0.0
TR:A0A2P5ANF8 A0A2P5ANF8_TREOI RING-type E3 ubiquitin transferase … 702 0.0
TR:A0A1S2XZS7 A0A1S2XZS7_CICAR RING-type E3 ubiquitin transferase … 702 0.0
TR:A0AAE1SAH3 A0AAE1SAH3_9SOLA RING-type E3 ubiquitin transferase … 702 0.0
TR:A0A1J7FNJ6 A0A1J7FNJ6_LUPAN RING-type E3 ubiquitin transferase … 702 0.0
TR:A0A2P5AJ14 A0A2P5AJ14_PARAD RING-type E3 ubiquitin transferase … 702 0.0
TR:A0A1U7VEA5 A0A1U7VEA5_NICSY RING-type E3 ubiquitin transferase … 702 0.0
TR:M1ABT1 M1ABT1_SOLTU RING-type E3 ubiquitin transferase OS=Solan… 702 0.0
TR:A0ABQ7W6L0 A0ABQ7W6L0_SOLTU RING-type E3 ubiquitin transferase … 702 0.0
TR:A0AAV9K3G9 A0AAV9K3G9_9SOLN RING-type E3 ubiquitin transferase … 702 0.0
TR:A0A1S4BCS6 A0A1S4BCS6_TOBAC RING-type E3 ubiquitin transferase … 702 0.0
TR:A0A6J5U2T1 A0A6J5U2T1_PRUAR RING-type E3 ubiquitin transferase … 702 0.0
TR:A0ACB8L4W7 A0ACB8L4W7_CITSI U-box domain-containing protein 14 … 702 0.0
TR:A0A2G2VCI1 A0A2G2VCI1_CAPBA RING-type E3 ubiquitin transferase … 702 0.0
TR:A0AAE1YU04 A0AAE1YU04_9LAMI RING-type E3 ubiquitin transferase … 701 0.0
TR:A0AAV1X6J7 A0AAV1X6J7_LUPLU RING-type E3 ubiquitin transferase … 701 0.0
TR:A0A1U8EX65 A0A1U8EX65_CAPAN RING-type E3 ubiquitin transferase … 701 0.0
TR:A0A5E4ECC4 A0A5E4ECC4_PRUDU RING-type E3 ubiquitin transferase … 701 0.0
TR:A0ABD1WYG0 A0ABD1WYG0_9LAMI RING-type E3 ubiquitin transferase … 701 0.0
TR:A0AAD7M5U3 A0AAD7M5U3_QUISA RING-type E3 ubiquitin transferase … 701 0.0
TR:A0AAD7Q5P3 A0AAD7Q5P3_QUISA RING-type E3 ubiquitin transferase … 700 0.0
TR:A0A6P8CYA9 A0A6P8CYA9_PUNGR RING-type E3 ubiquitin transferase … 700 0.0
TR:A0A1S3ZKQ3 A0A1S3ZKQ3_TOBAC RING-type E3 ubiquitin transferase … 700 0.0
TR:A0A2G5D164 A0A2G5D164_AQUCA RING-type E3 ubiquitin transferase … 700 0.0
TR:A0A6P5U3J9 A0A6P5U3J9_PRUAV RING-type E3 ubiquitin transferase … 700 0.0
TR:A0ABQ9M5E9 A0ABQ9M5E9_HEVBR RING-type E3 ubiquitin transferase … 699 0.0
TR:A0ACC0ZAK8 A0ACC0ZAK8_9ROSI Uncharacterized protein OS=Pistacia… 699 0.0
TR:A0ABM0P219 A0ABM0P219_PRUMU RING-type E3 ubiquitin transferase … 699 0.0
TR:A0A3Q7IQ73 A0A3Q7IQ73_SOLLC RING-type E3 ubiquitin transferase … 699 0.0
TR:A0AA39SE18 A0AA39SE18_ACESA RING-type E3 ubiquitin transferase … 699 0.0
TR:A0A5J5AW04 A0A5J5AW04_9ASTE RING-type E3 ubiquitin transferase … 699 0.0
TR:A0A314Y6E1 A0A314Y6E1_PRUYE RING-type E3 ubiquitin transferase … 699 0.0
TR:A0A251QMB1 A0A251QMB1_PRUPE RING-type E3 ubiquitin transferase … 699 0.0
TR:A0A7J7M6E3 A0A7J7M6E3_9MAGN RING-type E3 ubiquitin transferase … 699 0.0
TR:A0AAV1QYQ7 A0AAV1QYQ7_9ROSI RING-type E3 ubiquitin transferase … 699 0.0
TR:A0A835H9Y2 A0A835H9Y2_9MAGN RING-type E3 ubiquitin transferase … 699 0.0
TR:A0AAD4XDG0 A0AAD4XDG0_9MAGN RING-type E3 ubiquitin transferase … 698 0.0
TR:A0AAE1ILH4 A0AAE1ILH4_9FABA RING-type E3 ubiquitin transferase … 698 0.0
TR:A0A8J6CUL5 A0A8J6CUL5_9ROSI RING-type E3 ubiquitin transferase … 698 0.0
TR:A0ABD2QVW4 A0ABD2QVW4_9SOLN RING-type E3 ubiquitin transferase … 697 0.0
TR:A0A5N5FI06 A0A5N5FI06_9ROSA RING-type E3 ubiquitin transferase … 697 0.0
TR:A0A9Q0UCA7 A0A9Q0UCA7_SALPP RING-type E3 ubiquitin transferase … 697 0.0
TR:A0AAE0BAR6 A0AAE0BAR6_9ROSI RING-type E3 ubiquitin transferase … 697 0.0
TR:A0ABM1FMC2 A0ABM1FMC2_SOLPN RING-type E3 ubiquitin transferase … 697 0.0
TR:A0A6P6AKQ0 A0A6P6AKQ0_DURZI RING-type E3 ubiquitin transferase … 697 0.0
TR:A0AAP0LDP2 A0AAP0LDP2_9MAGN RING-type E3 ubiquitin transferase … 697 0.0
TR:A0AAP0QEX9 A0AAP0QEX9_9ROSI RING-type E3 ubiquitin transferase … 697 0.0
TR:A0A9J5WGK4 A0A9J5WGK4_SOLCO RING-type E3 ubiquitin transferase … 697 0.0
TR:A0ACB8L597 A0ACB8L597_CITSI U-box domain-containing protein 14 … 697 0.0
TR:A0A5C7GYA8 A0A5C7GYA8_9ROSI RING-type E3 ubiquitin transferase … 697 0.0
TR:A0A5D2XPV8 A0A5D2XPV8_GOSMU RING-type E3 ubiquitin transferase … 697 0.0
TR:A0A8T2Y9A9 A0A8T2Y9A9_POPDE RING-type E3 ubiquitin transferase … 697 0.0
TR:A0ABM2YU16 A0ABM2YU16_GOSHI RING-type E3 ubiquitin transferase … 697 0.0
TR:A0AAN8SZP9 A0AAN8SZP9_SOLBU RING-type E3 ubiquitin transferase … 697 0.0
TR:A0AAD5IAS4 A0AAD5IAS4_ACENE RING-type E3 ubiquitin transferase … 697 0.0
TR:A0A9Q0WKY8 A0A9Q0WKY8_9ROSI RING-type E3 ubiquitin transferase … 696 0.0
TR:A0A5D2NX47 A0A5D2NX47_GOSTO RING-type E3 ubiquitin transferase … 696 0.0
TR:A0ABD3TPL4 A0ABD3TPL4_9LAMI RING-type E3 ubiquitin transferase … 696 0.0
TR:A0A3S4PK96 A0A3S4PK96_9MAGN RING-type E3 ubiquitin transferase … 696 0.0
TR:A0AAD6PFY3 A0AAD6PFY3_9ROSI RING-type E3 ubiquitin transferase … 696 0.0
TR:A0A5J5B9Y0 A0A5J5B9Y0_9ASTE RING-type E3 ubiquitin transferase … 696 0.0
TR:A0ABC8UN55 A0ABC8UN55_9AQUA RING-type E3 ubiquitin transferase … 696 0.0
TR:B9HLH5 B9HLH5_POPTR RING-type E3 ubiquitin transferase OS=Popul… 696 0.0
TR:A0A5D2JAA3 A0A5D2JAA3_GOSTO RING-type E3 ubiquitin transferase … 696 0.0
TR:A0A8K0GRB9 A0A8K0GRB9_9ROSA RING-type E3 ubiquitin transferase … 695 0.0
TR:A0A5D2TDZ1 A0A5D2TDZ1_GOSMU RING-type E3 ubiquitin transferase … 695 0.0
TR:A0A5D2B520 A0A5D2B520_GOSDA RING-type E3 ubiquitin transferase … 695 0.0
TR:A0AAE1X2D8 A0AAE1X2D8_9LAMI RING-type E3 ubiquitin transferase … 695 0.0
TR:A0ABR2BSQ0 A0ABR2BSQ0_9ROSI RING-type E3 ubiquitin transferase … 695 0.0
TR:A0ABR2NSZ0 A0ABR2NSZ0_9ROSI RING-type E3 ubiquitin transferase … 695 0.0
TR:A0ABM3AXC1 A0ABM3AXC1_GOSHI RING-type E3 ubiquitin transferase … 695 0.0
TR:A0AAF0UY34 A0AAF0UY34_SOLVR RING-type E3 ubiquitin transferase … 695 0.0
TR:A0ABC8QLK3 A0ABC8QLK3_9AQUA RING-type E3 ubiquitin transferase … 695 0.0
TR:A0ABC8SRL1 A0ABC8SRL1_9AQUA RING-type E3 ubiquitin transferase … 695 0.0
TR:A0A9Q0SV54 A0A9Q0SV54_SALVM RING-type E3 ubiquitin transferase … 694 0.0
TR:A0ABU6RFH6 A0ABU6RFH6_9FABA RING-type E3 ubiquitin transferase … 694 0.0
TR:A0A8X7Z8K1 A0A8X7Z8K1_POPTO RING-type E3 ubiquitin transferase … 694 0.0
TR:A0ACC4BRY9 A0ACC4BRY9_POPAL Uncharacterized protein OS=Populus … 694 0.0
TR:A0AAD6MJN4 A0AAD6MJN4_9ROSI RING-type E3 ubiquitin transferase … 694 0.0
TR:A0ABD2TIP0 A0ABD2TIP0_9SOLN RING-type E3 ubiquitin transferase … 694 0.0
TR:A0ACC2KS43 A0ACC2KS43_PERAE Uncharacterized protein OS=Persea a… 694 0.0
TR:A0ABD3E7B2 A0ABD3E7B2_9LAMI RING-type E3 ubiquitin transferase … 694 0.0
TR:A0AAP0NZD1 A0AAP0NZD1_9MAGN RING-type E3 ubiquitin transferase … 694 0.0
TR:A0A834W7W7 A0A834W7W7_9FABA RING-type E3 ubiquitin transferase … 694 0.0
TR:A0A5J5PUV7 A0A5J5PUV7_GOSBA RING-type E3 ubiquitin transferase … 694 0.0
TR:A0A0B0NXG6 A0A0B0NXG6_GOSAR RING-type E3 ubiquitin transferase … 694 0.0
TR:A0ABR0NKE6 A0ABR0NKE6_GOSAR Uncharacterized protein OS=Gossypiu… 694 0.0
TR:A0A9D3UIB5 A0A9D3UIB5_9ROSI RING-type E3 ubiquitin transferase … 694 0.0
TR:A0A1R3H5H3 A0A1R3H5H3_COCAP RING-type E3 ubiquitin transferase … 694 0.0
TR:A0ACC2KS90 A0ACC2KS90_PERAE Uncharacterized protein OS=Persea a… 694 0.0
TR:A0AAJ6UKJ1 A0AAJ6UKJ1_POPEU RING-type E3 ubiquitin transferase … 694 0.0
TR:A0ABD1GH90 A0ABD1GH90_SALDI RING-type E3 ubiquitin transferase … 694 0.0
TR:A0A1R3JE18 A0A1R3JE18_9ROSI RING-type E3 ubiquitin transferase … 694 0.0
TR:A0AA88D9K5 A0AA88D9K5_FICCA RING-type E3 ubiquitin transferase … 694 0.0
TR:A0A7J9IC62 A0A7J9IC62_9ROSI RING-type E3 ubiquitin transferase … 693 0.0
TR:A0A7N2R9R8 A0A7N2R9R8_QUELO RING-type E3 ubiquitin transferase … 693 0.0
TR:A0A835JZV4 A0A835JZV4_9ROSI RING-type E3 ubiquitin transferase … 693 0.0
TR:A0A8J5FTW6 A0A8J5FTW6_ZINOF RING-type E3 ubiquitin transferase … 693 0.0
TR:A0A4Y7KVD5 A0A4Y7KVD5_PAPSO RING-type E3 ubiquitin transferase … 693 0.0
TR:A0A5D2F4H4 A0A5D2F4H4_GOSDA RING-type E3 ubiquitin transferase … 693 0.0
TR:A0A8J4VKI5 A0A8J4VKI5_9ROSI RING-type E3 ubiquitin transferase … 693 0.0
TR:A0AAN9F206 A0AAN9F206_CROPI RING-type E3 ubiquitin transferase … 693 0.0
TR:A0A5J5U6C5 A0A5J5U6C5_GOSBA RING-type E3 ubiquitin transferase … 693 0.0
TR:A0A6A2Y6Q4 A0A6A2Y6Q4_HIBSY RING-type E3 ubiquitin transferase … 693 0.0
TR:A0A5B6WJW5 A0A5B6WJW5_9ROSI RING-type E3 ubiquitin transferase … 692 0.0
TR:A0A498JGF4 A0A498JGF4_MALDO RING-type E3 ubiquitin transferase … 692 0.0
TR:A0A7N2MEE7 A0A7N2MEE7_QUELO RING-type E3 ubiquitin transferase … 692 0.0
TR:A0AAV6Y5W4 A0AAV6Y5W4_9LAMI RING-type E3 ubiquitin transferase … 692 0.0
TR:A0AAP0KTP2 A0AAP0KTP2_9MAGN RING-type E3 ubiquitin transferase … 692 0.0
TR:A0A7J6WEB3 A0A7J6WEB3_THATH RING-type E3 ubiquitin transferase … 692 0.0
TR:A0A2G5CD20 A0A2G5CD20_AQUCA RING-type E3 ubiquitin transferase … 692 0.0
TR:W9SKF5 W9SKF5_9ROSA RING-type E3 ubiquitin transferase OS=Morus… 692 0.0
TR:A0A072VQ10 A0A072VQ10_MEDTR U-box domain-containing protein 12 … 692 0.0
TR:A0A9W7I114 A0A9W7I114_HIBTR RING-type E3 ubiquitin transferase … 691 0.0
TR:A0A9W7M2C2 A0A9W7M2C2_HIBTR RING-type E3 ubiquitin transferase … 691 0.0
TR:A0A371F3Y5 A0A371F3Y5_MUCPR RING-type E3 ubiquitin transferase … 691 0.0
TR:A0A8J5FNQ2 A0A8J5FNQ2_ZINOF RING-type E3 ubiquitin transferase … 691 0.0
TR:A0A5N5FBR6 A0A5N5FBR6_9ROSA RING-type E3 ubiquitin transferase … 691 0.0
TR:A0ABC8QLK2 A0ABC8QLK2_9AQUA RING-type E3 ubiquitin transferase … 691 0.0
TR:A0ACB8L5G0 A0ACB8L5G0_CITSI U-box domain-containing protein 14 … 691 0.0
TR:A0ABR0CTF8 A0ABR0CTF8_9LAMI RING-type E3 ubiquitin transferase … 691 0.0
TR:A0A1U7ZJ54 A0A1U7ZJ54_NELNU RING-type E3 ubiquitin transferase … 691 0.0
TR:A0A540L773 A0A540L773_MALBA RING-type E3 ubiquitin transferase … 691 0.0
TR:A0ACB9ZMC7 A0ACB9ZMC7_CATRO Uncharacterized protein OS=Catharan… 691 0.0
TR:A0A1U7Z4U1 A0A1U7Z4U1_NELNU RING-type E3 ubiquitin transferase … 691 0.0
TR:A0A8B8MC51 A0A8B8MC51_ABRPR RING-type E3 ubiquitin transferase … 691 0.0
TR:A0A822YGE3 A0A822YGE3_NELNU RING-type E3 ubiquitin transferase … 691 0.0
TR:A0A7J7CI66 A0A7J7CI66_TRIWF RING-type E3 ubiquitin transferase … 690 0.0
TR:A0A8X8ZET3 A0A8X8ZET3_SALSN U-box domain-containing protein 12 … 690 0.0
TR:A0ACB7HJV7 A0ACB7HJV7_MANES Uncharacterized protein OS=Manihot … 690 0.0
TR:A0A2P6SJA1 A0A2P6SJA1_ROSCH RING-type E3 ubiquitin transferase … 690 0.0
TR:A0A2Z7BEP1 A0A2Z7BEP1_9LAMI RING-type E3 ubiquitin transferase … 689 0.0
TR:A0A8X8ZH43 A0A8X8ZH43_SALSN RING-type E3 ubiquitin transferase … 689 0.0
TR:A0A8S0SA15 A0A8S0SA15_OLEEU RING-type E3 ubiquitin transferase … 689 0.0
TR:A0AAW2CXI6 A0AAW2CXI6_9ROSI RING-type E3 ubiquitin transferase … 689 0.0
TR:A0ACC1YBU1 A0ACC1YBU1_MELAZ RING-type E3 ubiquitin transferase … 689 0.0
TR:A0AAV1C6C9 A0AAV1C6C9_OLDCO RING-type E3 ubiquitin transferase … 689 0.0
This belongs to the U-box E3 ubiquitin ligase family.
- [Structure page for protein via RCSB]
Name: U-box domain-containing protein 12
Organism: Oryza sativa Japonica Group
Experimental Method: X-ray
Resolution: 2.5 Å
Molecular Weight: 8.952 kDa
- The resolution is 2.5 Å indicating good quality.
- There are no other molecules in the solved structure
- The structure contains only the protein and no additional bound molecules.
- It belongs to the U-box structural family.
- I used 3D Protein Imager, located at https://3dproteinimaging.com/. I found it via https://bcrf.biochem.wisc.edu/2024/02/06/3d-protein-imager-a-pymol-qutemol-web-alternative/. https://doi.org/10.1093/bioinformatics/btaa009
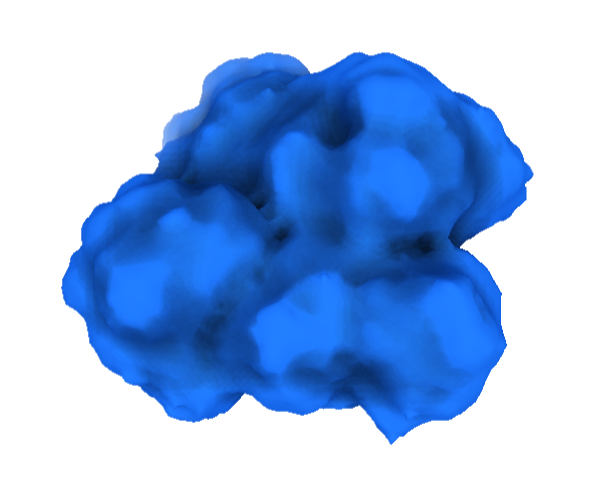
This can be accessed online, and free. They also provide an illustration service, which is pretty cool. It can be found below. This program did not have the “cartoon”, “ribbon”, and “ball and stick”, but it did have the “Real”, “Outline”, and “Goodsell” (cartoon-like) view modes where are shown from a “Sphere” mode of viewing. See image below.






To visualize it differently, I looked at it from a “tube” viewing mode. It has more helices than sheets. See image below.
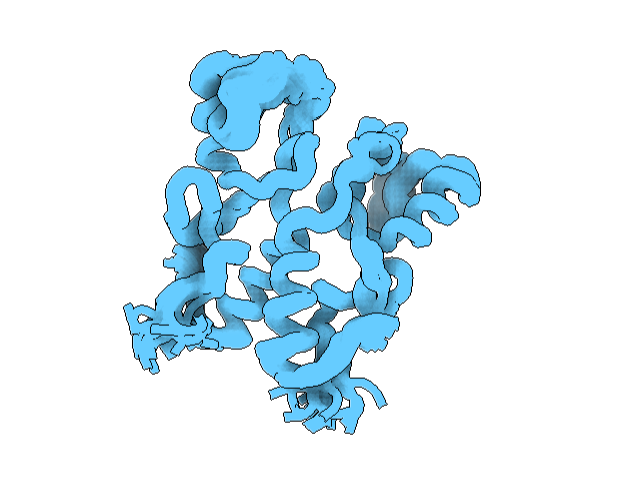

Hydrophobic residues are inside the protein core, and hydrophilic residues are on the surface. See image below.
Labeling by residue it has many charged and hydrophobic residues, with the latter being more focused inward. See image below.


Visualizing the surface, this structure appears compact, without deep binding cavities. See image below.


Part C
To be added with images
Part D
This part involved several of us students brainstorming for a lab according to the following instructions:
- Find a group of ~3–4 students
- Read through the Phage Reading material.
- Review the Bacteriophage Final Project Goals for engineering the L Protein:
- Choose one or two main goals from the list that you think you can address computationally (e.g., “We’ll try to stabilize the lysis protein,” or “We’ll attempt to disrupt its interaction with E. coli DnaJ.”).
- Write a 1-page proposal (bullet points or short paragraphs) describing:
- Each individually put your plan on your HTGAA website
Our initial proposal can be found below:


Week 5 HW: Protein Design Part II
This week’s homework was divided into 3 parts.
Part A involved SOD1 Binder Peptide Design.
That was broken into 3 parts:
Part 1: Part 1: Generate Binders with PepMLM
A) Begin by retrieving the human SOD1 sequence from UniProt (P00441) and introducing the A4V mutation.
Done
B) Using the PepMLM Colab linked from the HuggingFace PepMLM-650M model card:
Done
C) Generate four peptides of length 12 amino acids conditioned on the mutant SOD1 sequence.
AHWGVYTGVKKAKRX 15.649067
AWVPPYAVVYALRAX 18.835777
SRWPPYAARVEWAKA 19.332172
SRYDEVVGVKKLRKX 14.812749
D) To your generated list, add the known SOD1-binding peptide FLYRWLPSRRGG for comparison.
Done
Record the perplexity scores that indicate PepMLM’s confidence in the binders.
Recorded
Part 2: Evaluate Binders with AlphaFold3
A) Navigate to the AlphaFold Server: alphafoldserver.com
Done
B) For each peptide, submit the mutant SOD1 sequence followed by the peptide sequence as separate chains to model the protein-peptide complex.
Submitted:
Original: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
1st: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQAHWGVYTGVKKAKRA
2nd: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQAWVPPYAVVYALRAA
3rd: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQSRWPPYAARVEWAKA
4th: MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQSRYDEVVGVKKLRKA
5th (with mutant): MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQFLYRWLPSRRGG
All Xs were converted to As
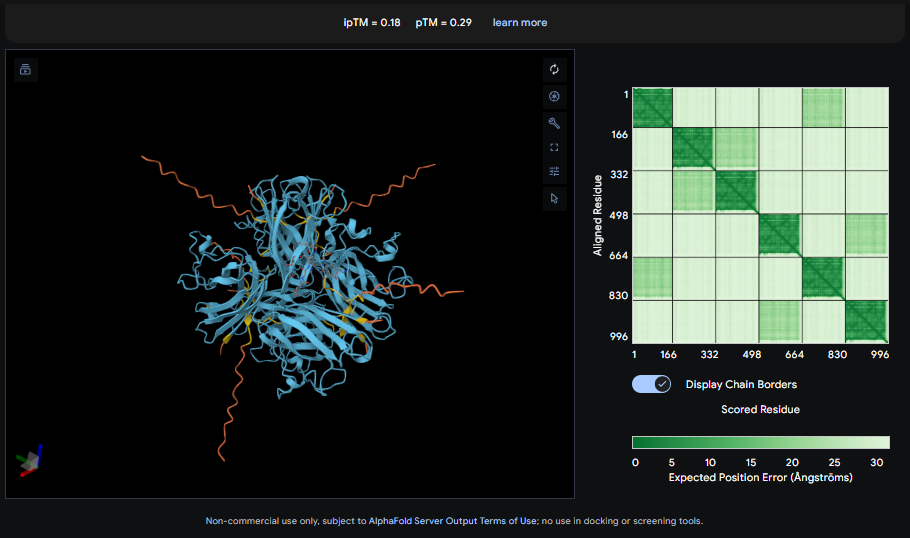
C/D) Record the ipTM score and briefly describe where the peptide appears to bind. Does it localize near the N-terminus where A4V sits? Does it engage the β-barrel region or approach the dimer interface? Does it appear surface-bound or partially buried? D) In a short paragraph, describe the ipTM values you observe and whether any PepMLM-generated peptide matches or exceeds the known binder.
The IpTM score is .18. This does not appear to indicate high confidence in binding. The lack of clear localiztion, strong β-barrel engagement, meaningful Dimer interfacing, combined with surface bounding of the peptides does not suggest strong binding interactions.

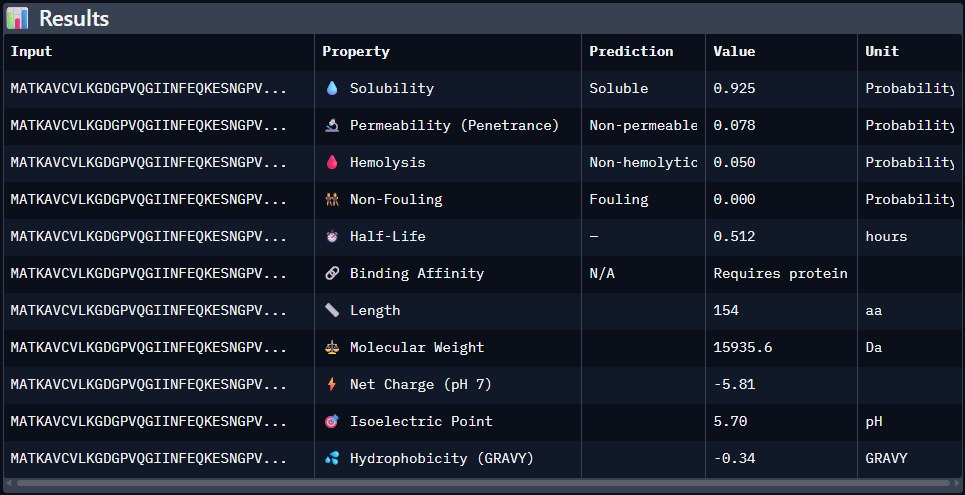
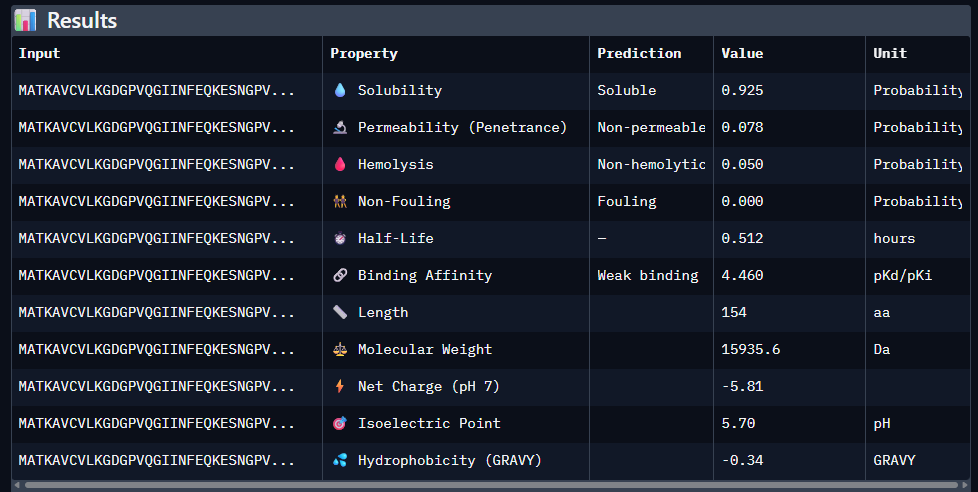
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
Paste the peptide sequence:
Paste the A4V mutant SOD1 sequence in the target field.
Check the boxes
Predicted binding affinity
Solubility
Hemolysis probability
Net charge (pH 7)
Molecular weight
Data for such were pasted as pictures below, in this order:
Original
1st Sequence: AHWGVYTGVKKAKRX
2nd Sequence: AWVPPYAVVYALRAX
3rd Sequence: SRWPPYAARVEWAKA
4th Sequence: SRYDEVVGVKKLRKX
5th Sequence: Mutant


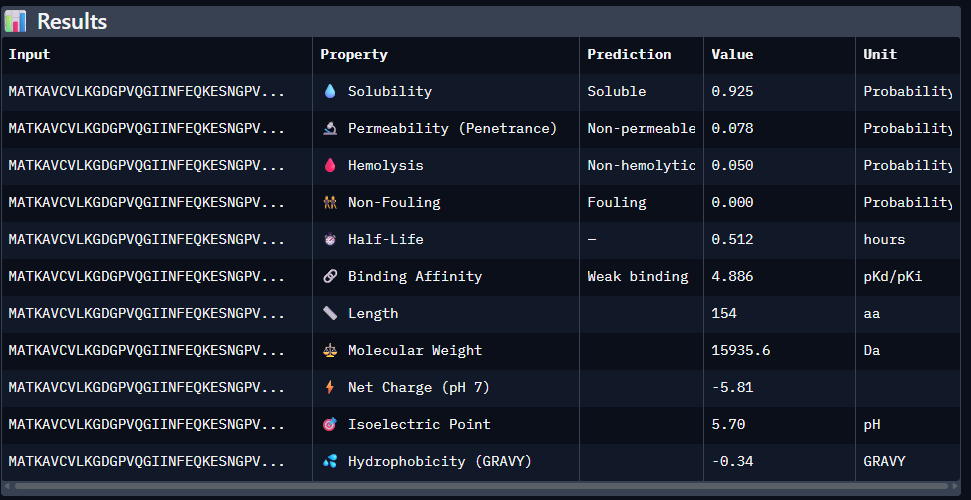

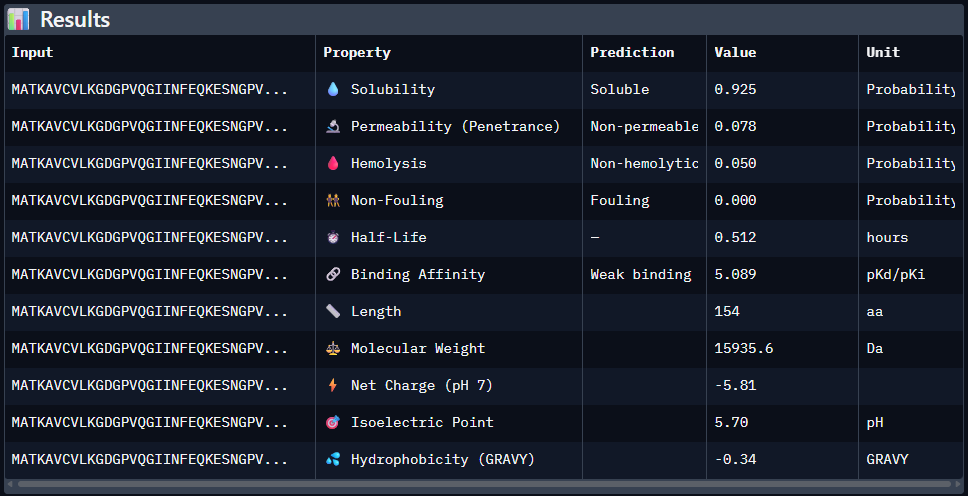

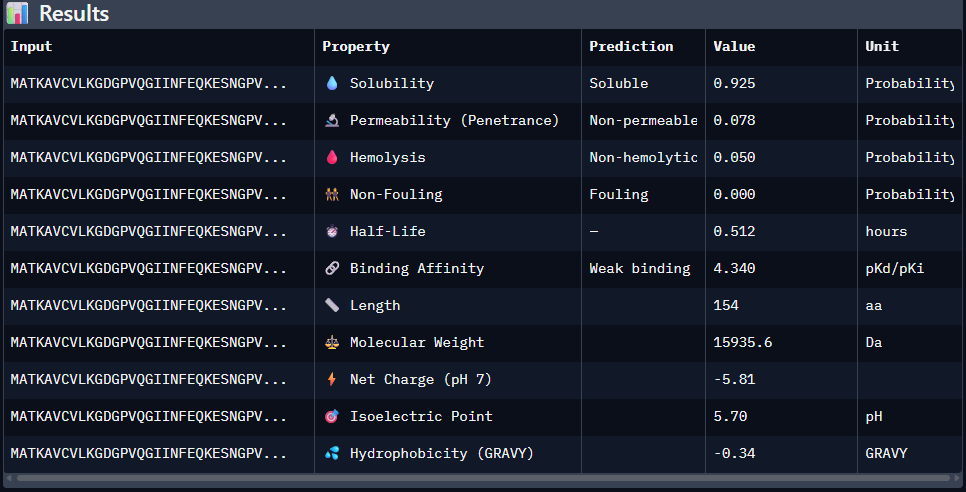

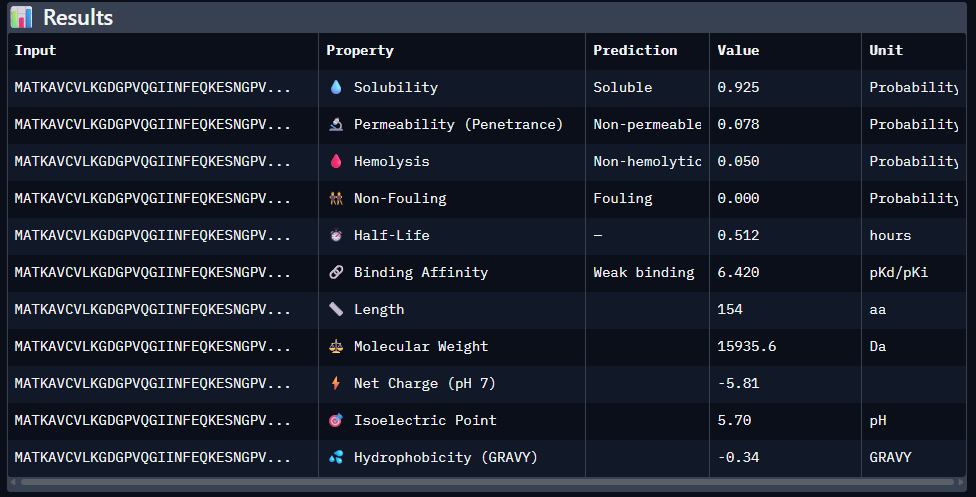

Part 4: Generate Optimized Peptides with moPPIt
Open the moPPit Colab linked from the HuggingFace moPPIt model card
Done
Make a copy and switch to a GPU runtime.
Done
In the notebook:
Paste your A4V mutant SOD1 sequence.
Choose specific residue indices on SOD1 that you want your peptide to bind (for example, residues near position 4, the dimer interface, or another surface patch).
Set peptide length to 12 amino acids.
Enable motif and affinity guidance (and solubility/hemolysis guidance if available). Generate peptides.
Done
After generation, briefly describe how these moPPit peptides differ from your PepMLM peptides. How would you evaluate these peptides before advancing them to clinical studies?
Erros were received and will be debugged.
Update:
This readout was given after fixing.
Binder Hemolysis Non-Fouling Solubility Half-Life Motif Specificity
TEVEEQEDRQHH 0.932494111 0.910815179 0.916666687 1.714286923 0.001987276 0.934523821
LAAGQALGITTA 0.918551639 0.573852718 0.416666627 6.734929085 0.006364056 0.886904776
ARKLTPEDQKQG 0.961578194 0.903518379 0.75 2.08972311 0.080991976 0.803571403
moPPit peptides appear better to develop. I would evaluate these peptides further computationally before advancing them to clinical studies. This would involve ranking by various qualities and further optimization. If successful, in vitro studies would follow.
Part B: BRD4 Drug Discovery Platform Tutorial (Gabriele)
This assignment was optional. For the sake of priorities, it will not be posted here.
Part C: Final Project: L-Protein Mutants
This was held between of 5 members, 3 of whom were able to provide their results jointly. We persued the Option 1: Mutagenesis.
To save space, given the large volume of images, a link to the inputs can be found in this google doc:
https://docs.google.com/document/d/1676c1tgFUlGaP-Bwp9_vDexbk3VsOJeuQeylNfvz76o/edit?usp=sharing.
Some of the results will be shown below:
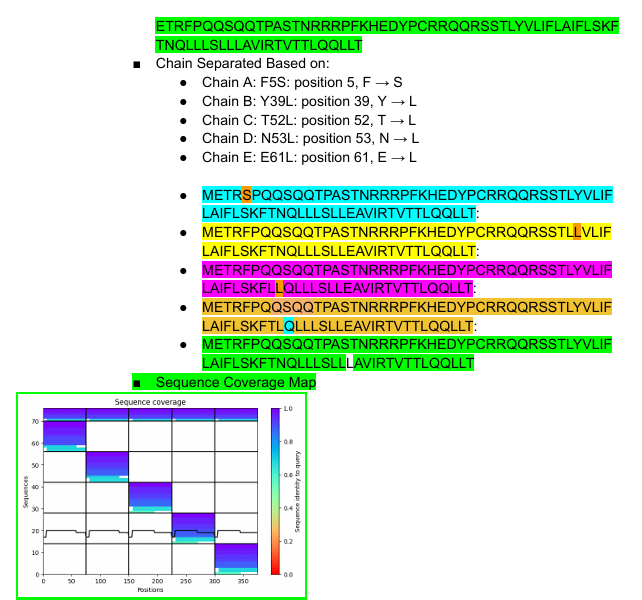


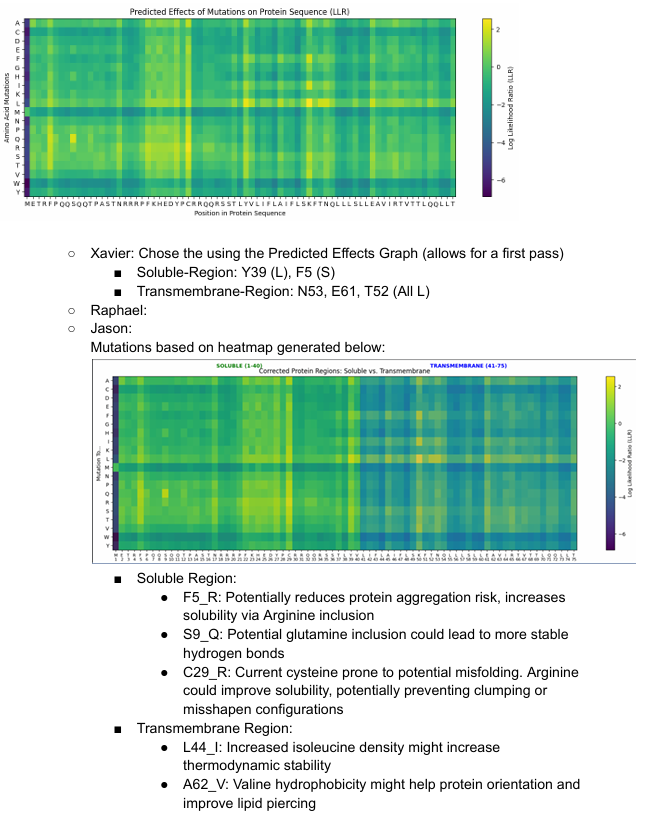



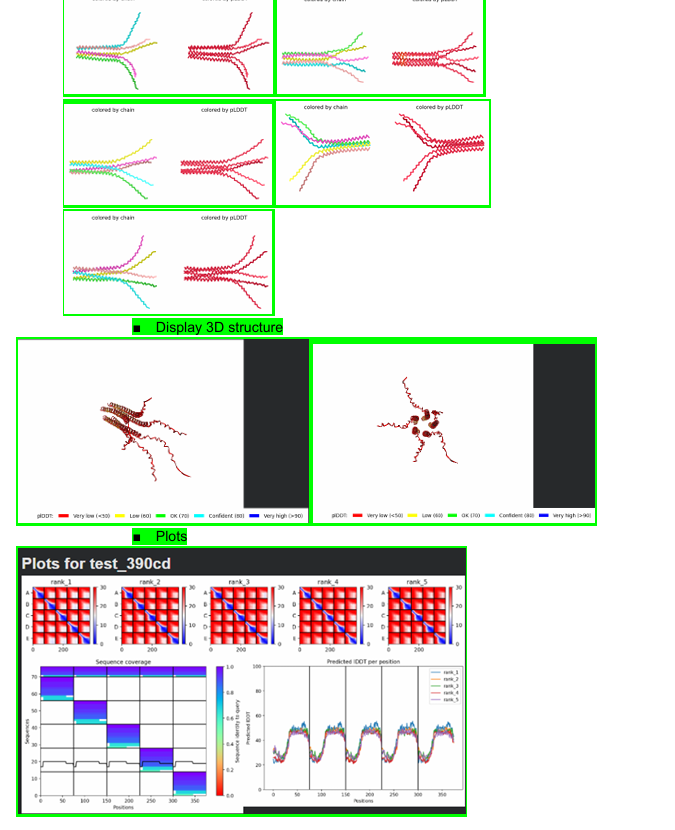

Week 6 HW: Genetic Circuits Part I: Assembly Technologies
Part 1: DNA Assembly
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Some of the components include the Phusion DNA Polymerease (key to PCR), dNTPs (these are used to synthesize new DNA), and buffering materials to stabilize the environmental conditions during the raction.
- What are some factors that determine primer annealing temperature during PCR?
Some of these include buffer conditions, primer melting temperature, primer length, and GC content.
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR uses primers, dNTPs, buffers and polymerase to amplify DNA.
Restriction enzyme digests are very precise in that they cut genomic material at precise locations, but they require restriction sites.
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
They need to be optimally designed for the system. This requires special attention primer design and their parameters.
- How does the plasmid DNA enter the E. coli cells during transformation?
After a shock is induced, plasmid DNA can enter E. Coli cells through pores that emerge in their membrane.
- Describe another assembly method in detail (such as Golden Gate Assembly)
6.1 Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Golden gate assembly is a method for efficient, seamless assembly that uses Type IIS restriction endonucleases to cleave DNA that exists outside of recognition sequences. Designs for inserts and cloning vectors place the Type IIS recognition sites distal to the cleavage site, and this allows efficient removal. NEB states 3 main advantages for GGA. One is that no scar sequence is introduced. Another is that the overhangs can allow for multiple simulaneous assemblies. Further, ligation and digestion can be done at the same time. A truncated example of a successful case is diagrammed below, with inspiration from a figure on NEB’s website.
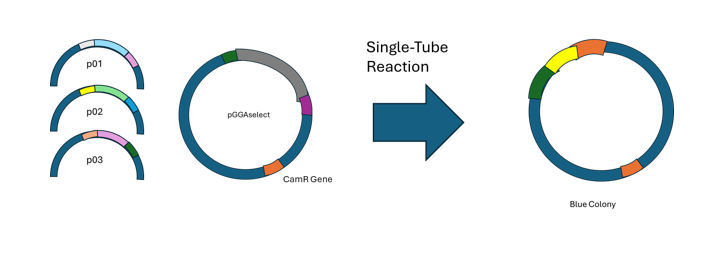

Source/Reference: https://www.neb.com/en/applications/cloning-and-synthetic-biology/dna-assembly-and-cloning/golden-gate-assembly
6.2 Model this assembly method with Benchling or Asimov Kernel!
Route chosen: Asimov Kernel
Owing to the brilliance of the Kernal from Asimov, redesigning the wheel was unnecessary.
I found the “pPD005 pcDNA Golden Gate” under the Addgene Plasmid Repository detailing a construct that was already created It is shown below in circular form.
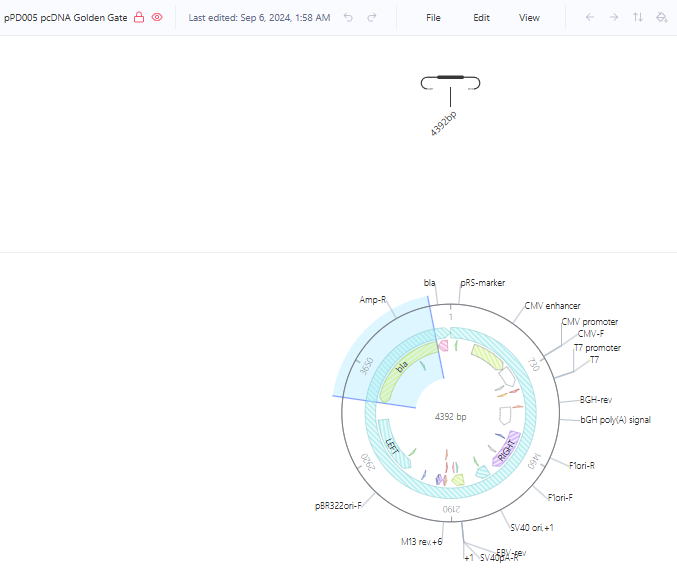

Part 2: Asimov Kernel
- Create a Repository for your work
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Create a blank Notebook entry to document the homework and save it to that Repository
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Explore the devices in the Bacterial Demos Repo to understand how the parts work together by running the Simulator on various examples, following the instructions for the simulator found in the “Info” panel (click the “i” icon on the right to open the Info panel)
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Create a blank Construct and save it to your Repository
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Recreate the Repressilator in that empty Construct by using parts from the Characterized Bacterial Parts repository
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Search the parts using the Search function in the right menu
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Drag and drop the parts into the Construct
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Confirm it works as expected by running the Simulator (“play” button) and compare your results with the Repressilator Construct found in the Bacterial Demos repository
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Document all of this work in your Notebook entry - you can copy the glyph image and the simulator graphs, and paste them into your Notebook
Done - Refer to W&M_Whale_Test_Rep_Notebook_Entry in Kernel
- Build three of your own Constructs using the parts in the Characterized Bacterials Parts Repo.
10.1. Explain in the Notebook Entry how you think each of the Constructs should function
10.2. Run the simulator and share your results in the Notebook Entry
10.3. If the results don’t match your expectations, speculate on why and see if you can adjust the simulator settings to get the expected outcome
10.1-10.3 These actions were all performed. I exported the images and markdown as below. However, they can be found under W&M_Whale_Test_Rep_Notebook_Entry
To explain constructs – these were recreations and thus were expected to act as the demos, prior to the run of their simulations. The success is in part a testament to effect design of the Kernal software.
What Follows is a markdown export of my Notebook Entry in Ansimov Kernal under W&M Whale Test Rep:
This entry starts with a general summary of the use of Asimov’s Kernel thus far. The user interface takes a little getting used to but is easy to learn.
I started by using the Bacterial Demos Repo as per the homework to understand how parts work together.
A simulation was ran which ran similarly to others other before it. I did not expect difference given that these were demos.
The Repressilator was the first, which was selected by starting from a blank construct using parts from the Characterized Bacterial Parts.
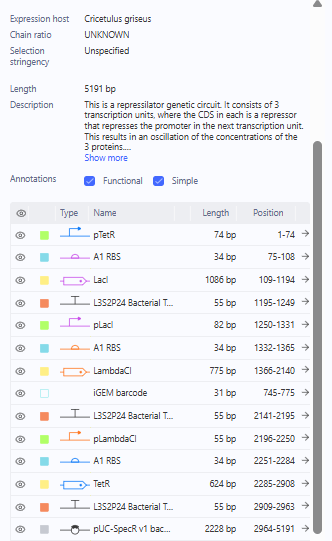

Characterized Bacterial Parts Examination for Repressilator


Represillator Recreation
This was then simulated with results expected from a Demo.
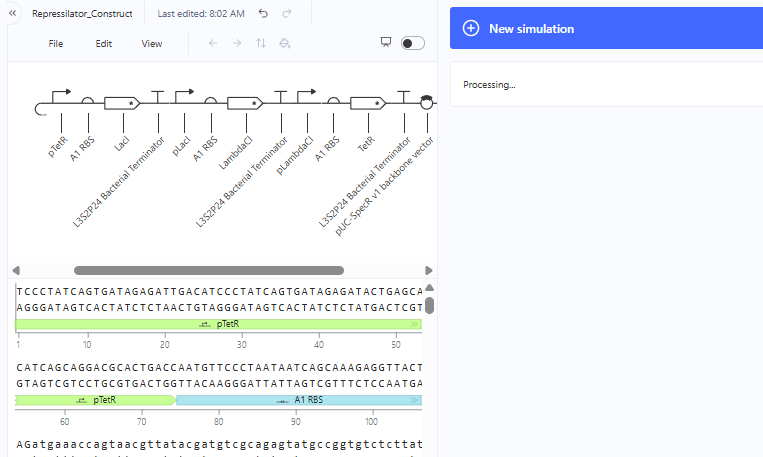

Represillator Recreation Simulation Start
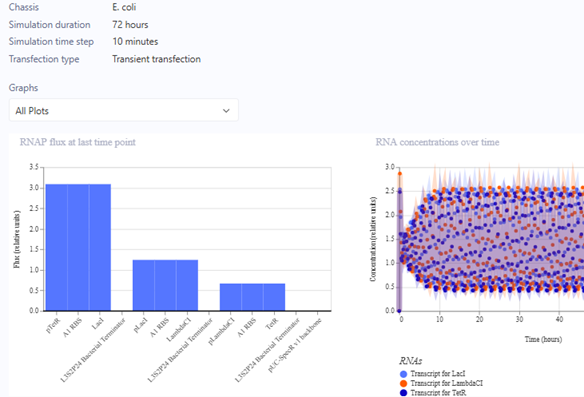

Represillator Recreation Preview
For the Homework, we were asked to create 3 more constructs. I chose recreations of the J23117 Promoter, Self-Replicating Promoter, and Circuit 3. Simulations accompanied them. I expected them to all run similar to the demos, which they did.
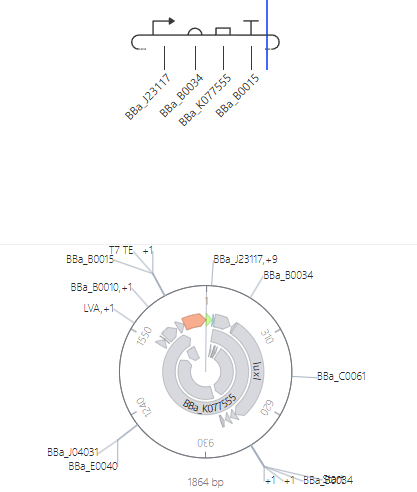

J23117 Promoter
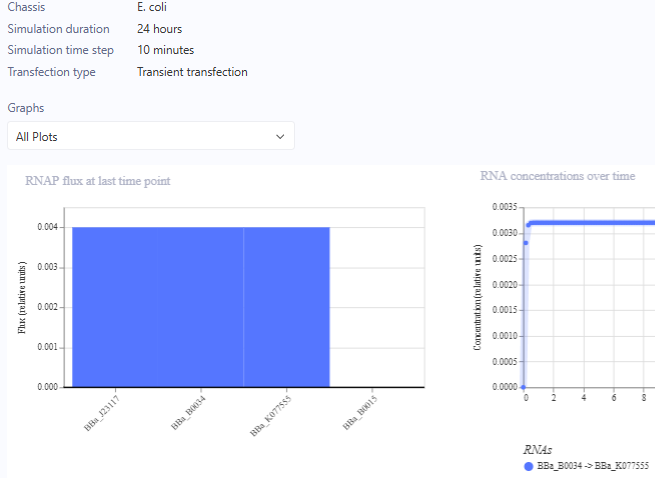

J23117 Promoter Simulation Preview
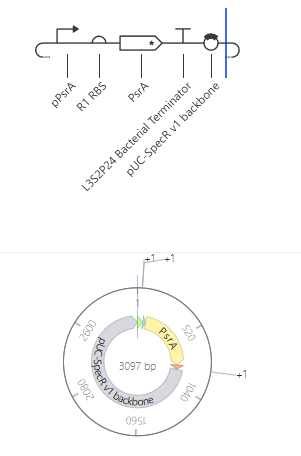

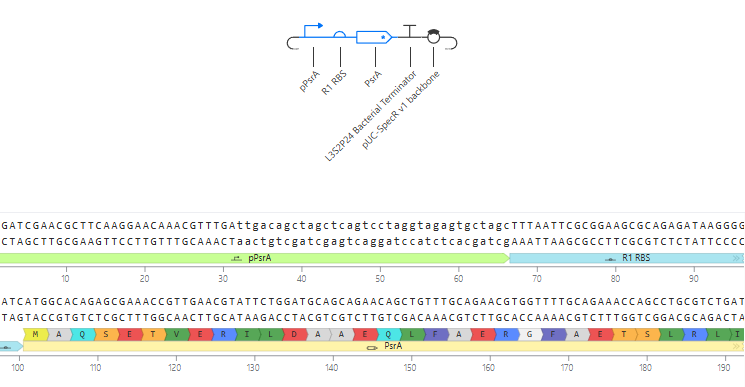
Self-Replicating Promoter
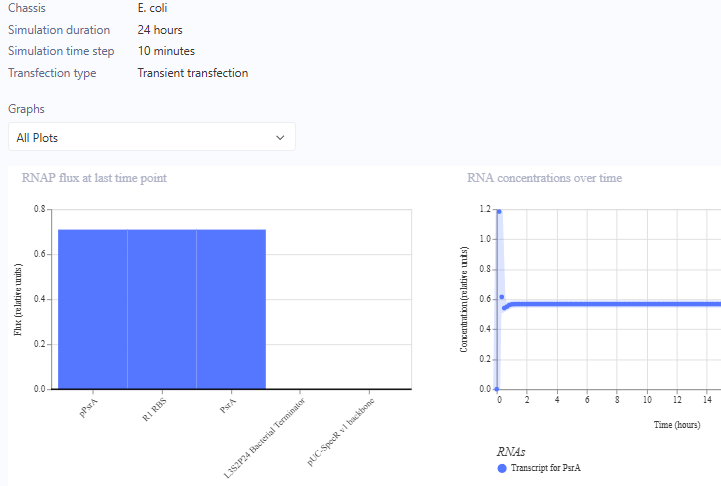

Self-Replicating Promoter Preview

Circuit 3
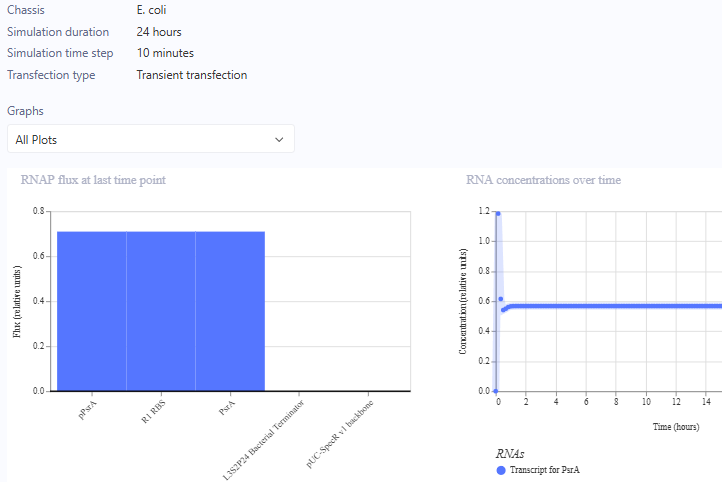

Circuit 3 Preview
Week 7 HW: Genetic Circuits Part II: Neuromorphic Circuits
Part 1:
What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
IANNs are great for analog computation, while also being scalable in deployment and useful for approximating functions.
Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
Disease diagnostics. Inputs would be expressions levels of biomarkers and signals deemed harmful. Outputs could be in the form of a gene with a strong signal or that is therapetic in nature. Strong limitations could come from unintended interactions with the output, as well as cross-talk generated. Noise within the biological system could affected outputs as well.
Part 2:
What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
I might genetically engineer fungi to eat waste products or grow as shielding or building material. In either case, the fungi can serve as a meaningful redesinging of the environment. Advantages of doing synthethic biology with fungi are at least three fold: A) you can perform unique protein modications and syntheses that you can’t yet easily do with bacteria, you can take advantage of more complex molecular regulatory systems within fungi, in some cases they may be more safely scalable, and you get easy “Last of Us” Jokes.
Part 3:
Objectives of this were as follows:
Review the Individual Final Project documentation guidelines.
Done
Submit this Google Form with your draft Aim 1, final project summary, HTGAA industry council selections, and shared folder for DNA designs.
In progress
Review Part 3: DNA Design Challenge of the week 2 homework. Design at least 1 insert sequence and place it into the Benchling/Kernel/Other folder you shared in the Google Form above. Document the backbone vector it will be synthesized in on your website.
This is TBA.
Week 9 HW: Week 9 — Cell-Free Systems
General homework questions
Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
To Be Added
Describe the main components of a cell-free expression system and explain the role of each component.
To Be Added
Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
To Be Added
Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
To Be Added
How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
To Be Added
Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
To Be Added
Homework question from Kate Adamala
Design an example of a useful synthetic minimal cell as follows:
Pick a function and describe it.
To Be Added
What would your synthetic cell do? What is the input and what is the output?
To Be Added
Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
To Be Added
Could this function be realized by genetically modified natural cell?
To Be Added
Describe the desired outcome of your synthetic cell operation.
To Be Added
Design all components that would need to be part of your synthetic cell.
To Be Added
What would be the membrane made of?
To Be Added
What would you encapsulate inside? Enzymes, small molecules.
To Be Added
Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
To Be Added
How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?)
Experimental details
To Be Added
List all lipids and genes. (bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.)
To Be Added
How will you measure the function of your system?
To Be Added
Homework question from Peter Nguyen
Freeze-dried cell-free systems can be incorporated into all kinds of materials as biological sensors or as inducible enzymes to modify the material itself or the surrounding environment. Choose one application field — Architecture, Textiles/Fashion, or Robotics — and propose an application using cell-free systems that are functionally integrated into the material. Answer each of these key questions for your proposal pitch:
To Be Added
Write a one-sentence summary pitch sentence describing your concept.
To Be Added
How will the idea work, in more detail? Write 3-4 sentences or more.
To Be Added
What societal challenge or market need will this address?
To Be Added
How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
To Be Added
Homework question from Ally Huang
Freeze-dried cell-free reactions have great potential in space, where resources are constrained. As described in my talk, the Genes in Space competition challenges students to consider how biotechnology, including cell-free reactions, can be used to solve biological problems encountered in space. While the competition is limited to only high school students, your assignment will be to develop your own mock Genes in Space proposal to practice thinking about biotech applications in space!
To Be Added
For this particular assignment, your proposal is required to incorporate the BioBits® cell-free protein expression system, but you may also use the other tools in the Genes in Space toolkit (the miniPCR® thermal cycler and the P51 Molecular Fluorescence Viewer). For more inspiration, check out https://www.genesinspace.org/ .
To Be Added
Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. (Maximum 100 words)
To Be Added
Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
To Be Added
Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
Clearly state your hypothesis or research goal and explain the reasoning behind it. (Maximum 150 words)
To Be Added
Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)
To Be Added
Part B:
Put your chosen final project slide in the appropriate slide deck following the instructions on slide 1:
MIT/Harvard/Wellesley ONE FINAL PROJECT IDEA
Committed Listener ONE FINAL PROJECT IDEA
Done
Submit this Final Project selection form if you have not already.
Done
Begin planning how you will write your final project documentation based on these guidelines
Initiated
Prepare your first DNA order and put it in the “Twist (MIT)” or “Twist (Nodes)” tab of the 2026 HTGAA Ordering: DNA, Reagents, Consumables spreadsheet, as appropriate.
In Progress
First Twist order deadline for MIT/Harvard/Wellesley students is Friday, April 3 at 11PM ET
First Twist order deadline for Committed Listeners is Friday, April 10 at 11PM ET. (Your Node Lead will place the Twist order, so please work with them to finalize your constructs and ordering decisions.)
In Progress
Week 10 HW: Week 10 — Advanced Imaging & Measurement Technology
Homework: Final Project
(Measurement Draft)
The main aspect of my project that I will measure is the functional activity of a mutated InaZ construct, specifically whether it increases ice nucleation efficiency relative to a control InaZ construct. I will perform this measurement using a controlled freezing assay in which replicate samples are cooled gradually and monitored for the onset of ice formation. The primary data collected will be the temperature at which freezing begins in each sample. Ice formation will be detected through optical observation of crystal formation and through temperature sensors that record the freezing point. In addition, I would verify the identity of the mutated construct using PCR, gel electrophoresis, and DNA sequencing. PCR and gel electrophoresis would be used to confirm the presence and approximate size of the inserted DNA, while DNA sequencing would confirm that the engineered inaZ mutation is correct. Together, these measurements allow me to confirm both that the construct was built properly and that it produces the intended increase in ice nucleation activity.
Restated:
Functional Assay: Ice Nucleation
Objective: Measure efficiency relative to a control construct.
Method: Controlled freezing assay with gradual cooling of replicate samples.
Data Points: Freezing onset temperature, recorded via optical observation and thermal sensors.
Genetic Verification
PCR & Gel Electrophoresis: Confirm the presence and approximate size of the inserted DNA.
DNA Sequencing: Verify the exact sequence of the engineered inaZ mutation.
Homework: Waters Part I — Molecular Weight
Work to be Added
Homework: Waters Part II — Secondary/Tertiary structure
Work to be Added
Homework: Waters Part III — Peptide Mapping - primary structure
Work to be Added
Homework: Waters Part IV — Oligomers
Work to be Added
Homework: Waters Part V — Did I make GFP?
Work to be Added
Week 11 HW: Bioproduction & Cloud Labs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
- Contribute at least one pixel to this global artwork experiment before the editing ends on Sunday 4/19 at 11:59 PM EST!
A personalized URL was sent to the email address associated with your Discourse account, and you can discuss the artwork on the Discourse!
If you did not have a chance to contribute, it’s okay, just make sure you become a TA this fall! 😉
Done: Added pixels.
- Make a note on your HTGAA webpages including:
Done: I added a total of 14 pixels by the end of the experiment: 12 colored pixels and 2 removal pixels.
what you contributed to the community bioart project (e.g., “I made part of the DNA on the bottom right plate”)
Done: I added a couple small details to subfeatures of some designs. One example was adding a temporary light to one of the spaceships.
what you liked about the project, and
Done: I liked the collaborative and semi-collaborative nature of the project. There was space for those who wanted to add with coordination and space for those who wanted to add individually from their own vision. Seeing what people came up with was great, as well.
what about this collaborative art experiment could be made better for next year.
Done: To improve it, one could widen the color section, plate area, and reduce the painting refresh-time.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
- Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
E. coli Lysate
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase)
Salts/Buffer
Potassium Glutamate
HEPES-KOH pH 7.5
Magnesium Glutamate
Potassium phosphate monobasic
Potassium phosphate dibasic
Energy / Nucleotide System
Ribose
Glucose
AMP
CMP
GMP
UMP
Guanine
Translation Mix (Amino Acids)
17 Amino Acid Mix
Tyrosine
Cysteine
Additives
Nicotinamide
Backfill
Nuclease Free Water
To Be Added.
- Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
To Be Added.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
- Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
sfGFP
mRFP1
mKO2
mTurquoise2
mScarlet_I
Electra2
The amino acid sequences are shown in the HTGAA Cell-Free Benchling folder.
To Be Added.
- Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
To Be Added.
- The second phase of this lab will be to define the precise reagent concentrations for your cell-free experiment. You will be assigned artwork wells with specific fluorescent proteins and receive an email with instructions this week (by April 24). You can begin composing master mix compositions here.
To Be Added.
Part D: Build-A-Cloud-Lab | (optional) Bonus Assignment
- Use this simulation tool to create an interesting looking cloud lab out of the Ginkgo Reconfigurable Automation Carts. This is just a minimal implementation so far, but I would love to see some fun designs!
To Be Added.
Week X HW: X
Week X HW: X
Week X HW: X
Subsections of Labs
Week 1 Lab: Pipetting
This was a lab review week. It covered standard lab practices. There’s nothing to note here aside from that review material from source and concepts below were reviewed.
Source: https://thecrashcourse.com/courses/unit-conversion-significant-figures-crash-course-chemistry-2/
Concepts: Dilution Problems, Chemistry, Molarity & Concentration Examples, Formula & Equations
![cover image]()
![cover image]()
Week 2 Lab: Lab DNA Gel Art
This lab covered gel art through restriction digests and gel electrophoresis
As a remote participant, I reviewed the material.
Week 3 Lab: Lab Automation
This lab, we were tasked with creating a design that could be generated by an OpenTrons Liquid Handling Robot.
As a remote participant, I prototyped a design using the GUI at opentrons-art.rcdonovan.com.
This resulted in a layered plus symbol shown below.
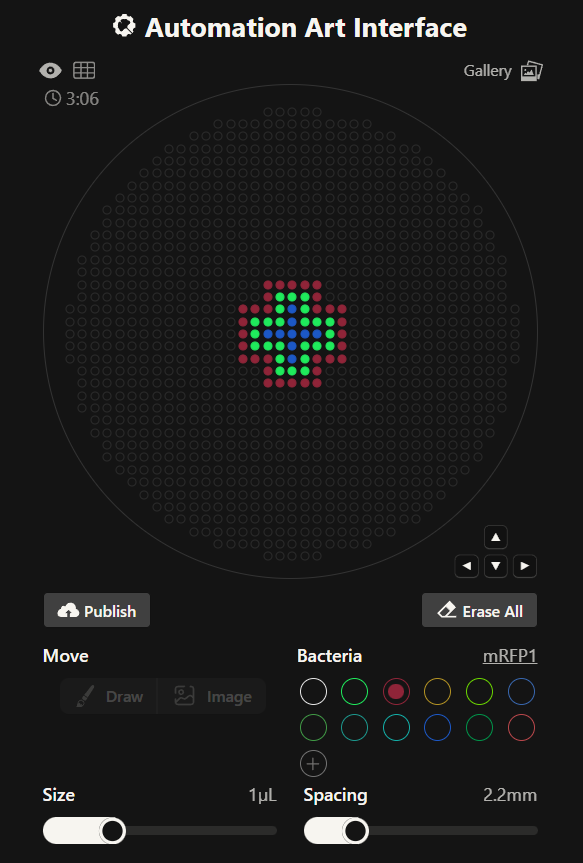

The coordinates for generating such can be found here, courtesy of RC Donovan’s tool:
sfgfp_points = [(-2.2, 6.6),(0, 6.6),(2.2, 6.6),(-2.2, 4.4),(2.2, 4.4),(-6.6, 2.2),(-4.4, 2.2),(-2.2, 2.2),(2.2, 2.2),(4.4, 2.2),(6.6, 2.2),(-6.6, 0),(6.6, 0),(-6.6, -2.2),(-4.4, -2.2),(-2.2, -2.2),(2.2, -2.2),(4.4, -2.2),(6.6, -2.2),(-2.2, -4.4),(2.2, -4.4),(-2.2, -6.6),(0, -6.6),(2.2, -6.6)]
electra2_points = [(0, 4.4),(0, 2.2),(-4.4, 0),(-2.2, 0),(0, 0),(2.2, 0),(4.4, 0),(0, -2.2),(0, -4.4)]
mrfp1_points = [(-4.4, 8.8),(-2.2, 8.8),(0, 8.8),(2.2, 8.8),(4.4, 8.8),(-4.4, 6.6),(4.4, 6.6),(-8.8, 4.4),(-6.6, 4.4),(-4.4, 4.4),(4.4, 4.4),(6.6, 4.4),(8.8, 4.4),(-8.8, 2.2),(8.8, 2.2),(-8.8, 0),(8.8, 0),(-8.8, -2.2),(8.8, -2.2),(-8.8, -4.4),(-6.6, -4.4),(-4.4, -4.4),(4.4, -4.4),(6.6, -4.4),(8.8, -4.4),(-4.4, -6.6),(4.4, -6.6),(-4.4, -8.8),(-2.2, -8.8),(0, -8.8),(2.2, -8.8),(4.4, -8.8)]
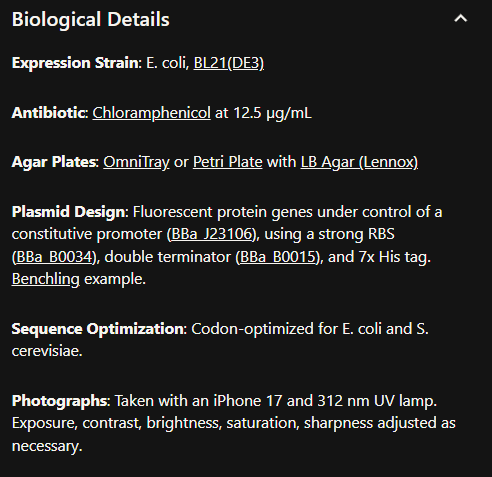

For my node, in order to work with their printer which had two colors, a modified version was created. Gemini within was tested and employed to deliver the following result. The following code which creates the following pattern was then emailed to the TA for printing.
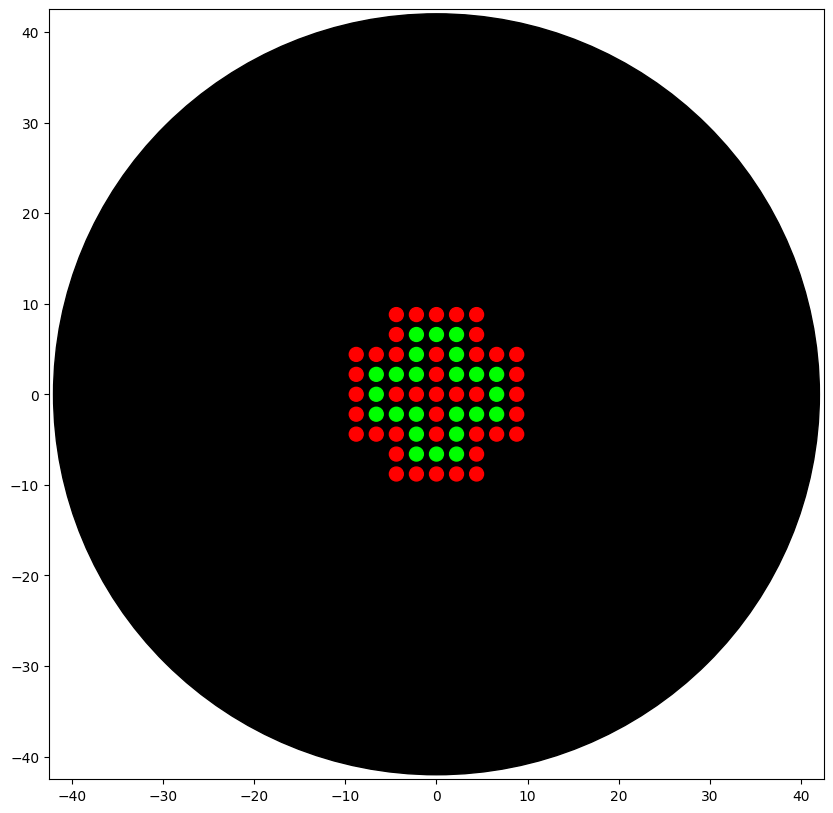

Week 4 Lab: Protein Design Part I
This lab was effectively part of Week4’s Homework (“This week’s Lab work is effectively part of this week’s Homework; see that assignment and document your work there.”).
As follows, the assigned work was:
- 1.Find a group of ~3–4 students
- 2.Read through the Phage Reading material listed under “Reading & Resources” below.
- 3.Review the Bacteriophage Final Project Goals for engineering the L Protein:
*Increased stability (easiest)
*Higher titers (medium)
*Higher toxicity of lysis protein (hard)
- 4.Brainstorm Session
- Choose one or two main goals from the list that you think you can address computationally (e.g., “We’ll try to stabilize the lysis protein,” or “We’ll attempt to disrupt its interaction with E. coli DnaJ.”).
- Write a 1-page proposal (bullet points or short paragraphs) describing:
Which tools/approaches from recitation you propose using (e.g., “Use Protein Language Models to do in silico mutagenesis, then AlphaFold-Multimer to check complexes.”).
- Why do you think those tools might help solve your chosen sub-problem?
- Name one or two potential pitfalls (e.g., “We lack enough training data on phage–bacteria interactions.”).
- Include a schematic of your pipeline.
- 5.Each individually put your plan on your HTGAA website
- Include your group’s short plan for engineering a bacteriophage
This can be found in Week 4’s HW Part D.
Week 5 Lab: Protein Design Part II
This week’s Lab work was effectively part of this week’s Homework. This is reflected in Part C of the week 5 homeowork.
Week 6 Lab — Genetic Circuits Part I: Assembly Technologies
This lab involved the Gibson Assembly and was in person.
As an online student, this lab did not apply for me.
Week 7 — Genetic Circuits Part II: Neuromorphic Circuits
This week’s lab had a dry and wet component. The focus was the building of our own IANN. According to the lab: “IANNs are also universal function approximators–given an adequate number of intracellular artificial neurons, you can use an IANN to achieve any input/output behavior you’d like.” https://2026a.htgaa.org/2026a/course-pages/weeks/week-07/lab/index.html
The Pre-lab involved us setting up and understanding Neuromorphic Wizard. We wrote instructions using this template (https://www.google.com/url?q=https://docs.google.com/spreadsheets/d/12S4Vv6e_am6U6dMgpijt1G9rtoRyfcdoKdIXvnkdGTo/edit?usp%3Dsharing&sa=D&source=editors&ust=1774224628668116&usg=AOvVaw2ayNzuuoVfm9mQYCP30sjK) and using these names (https://www.google.com/url?q=https://docs.google.com/spreadsheets/d/1cyEgmj08P40iUE5KOdvn_oaDhB7sOkQJwA7900rDqMc/edit?usp%3Dsharing&sa=D&source=editors&ust=1774224628668584&usg=AOvVaw3_lcXglYGq-h7wgkIkT-Tx).
Finally we entered our circuit into a google form.
Our Members were: Jason Ross, Nana Agyei, and Xavier Palmer. Jason served as the project submitter.
Pictures of aspects of the process can be found below.
Esssentially, we installed Neuromorphic Wizard, used the template, and generated outputs, before submitting our project.


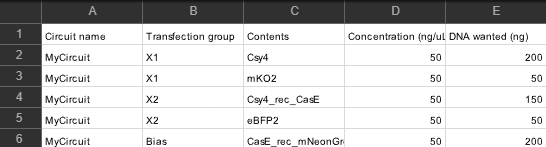

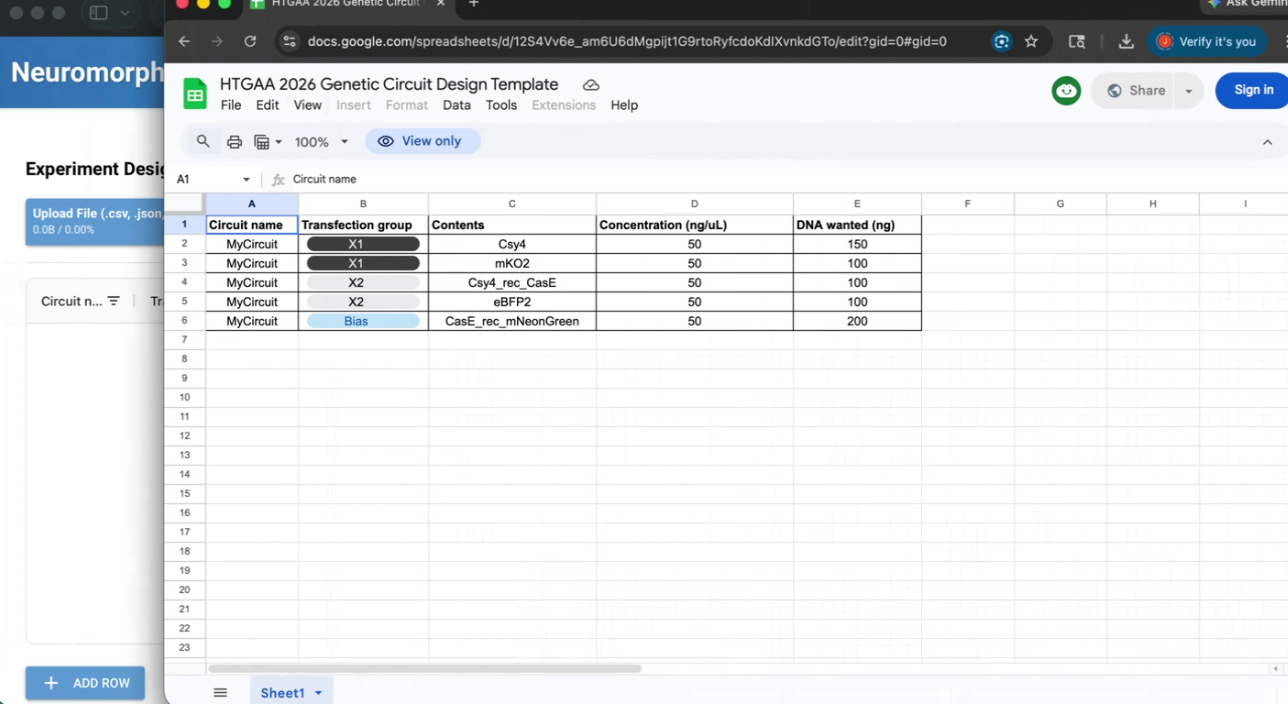

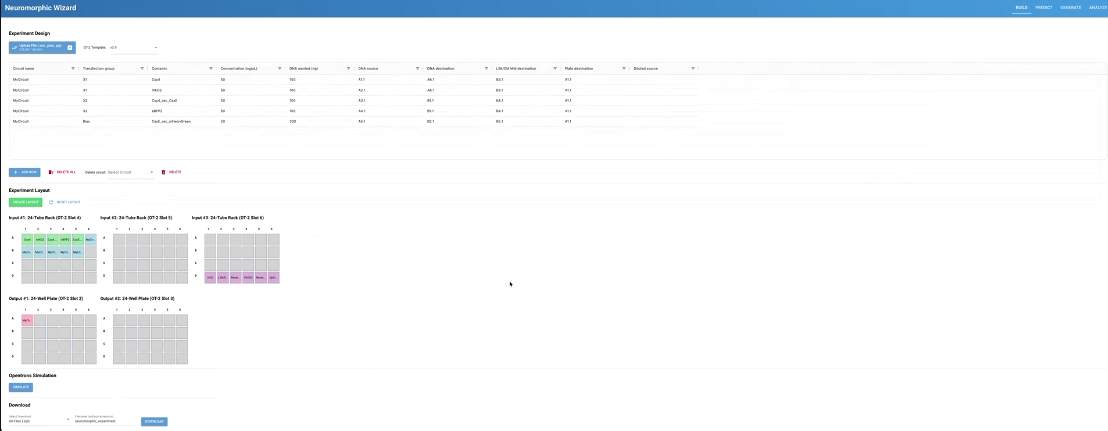

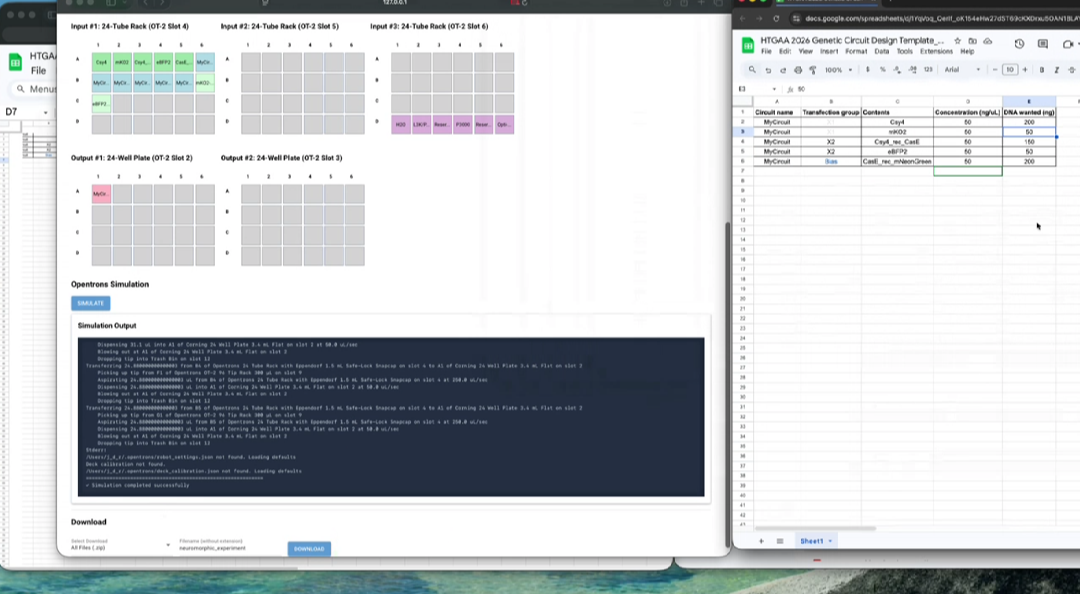


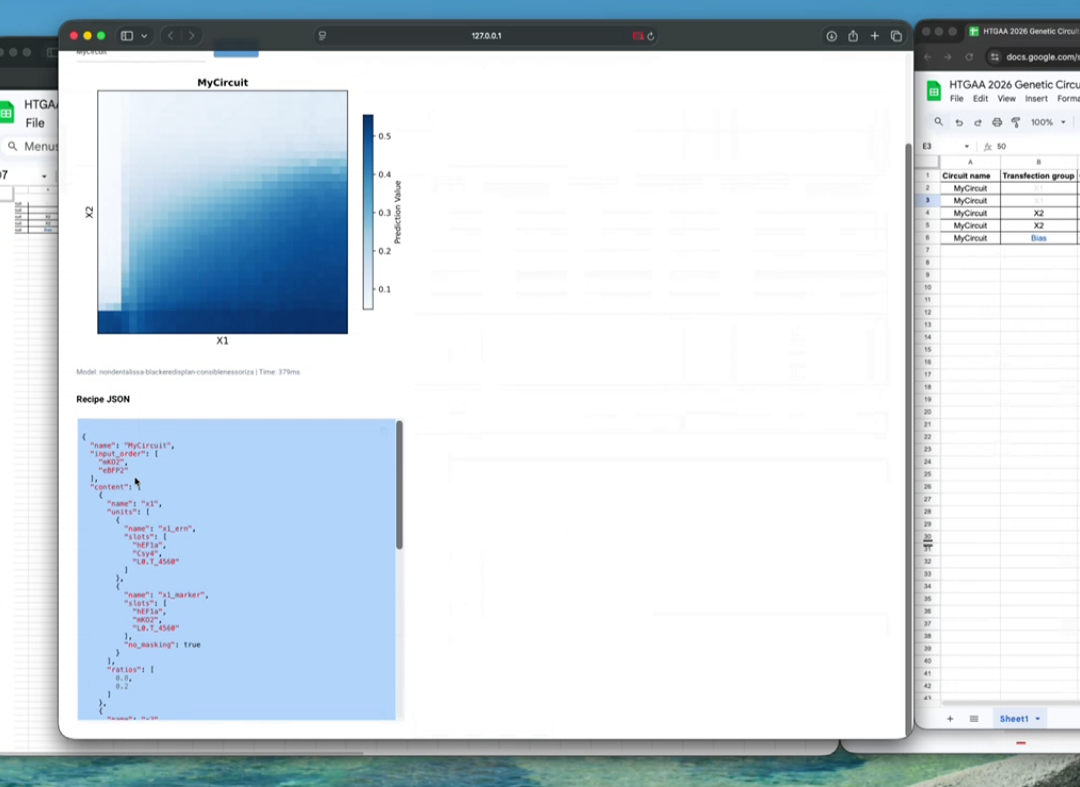

The code below the graph is:
{
“name”: “MyCircuit”,
“input_order”: [
“mKO2”,
“eBFP2”
],
“content”: [
{
“name”: “x1”,
“units”: [
{
“name”: “x1_ern”,
“slots”: [
“hEF1a”,
“Csy4”,
“L0.T_4560”
]
},
{
“name”: “x1_marker”,
“slots”: [
“hEF1a”,
“mKO2”,
“L0.T_4560”
],
“no_masking”: true
}
],
“ratios”: [
0.8,
0.2
]
},
{
“name”: “x2”,
“units”: [
{
“name”: “x2_output”,
“slots”: [
“hEF1a”,
“Csy4_rec”,
“CasE”,
“L0.T_4560”
]
},
{
“name”: “x2_marker”,
“slots”: [
“hEF1a”,
“eBFP2”,
“L0.T_4560”
],
“no_masking”: true
}
],
“ratios”: [
0.75,
0.25
]
},
{
“name”: “bias”,
“units”: [
{
“name”: “bias_output”,
“slots”: [
“hEF1a”,
“CasE_rec”,
“mNeonGreen”,
“L0.T_4560”
]
}
],
“ratios”: [
1.0
]
}
]
}