Week 2 HW: DNA r/w/e


1. Benchling & In-silico Gel Art
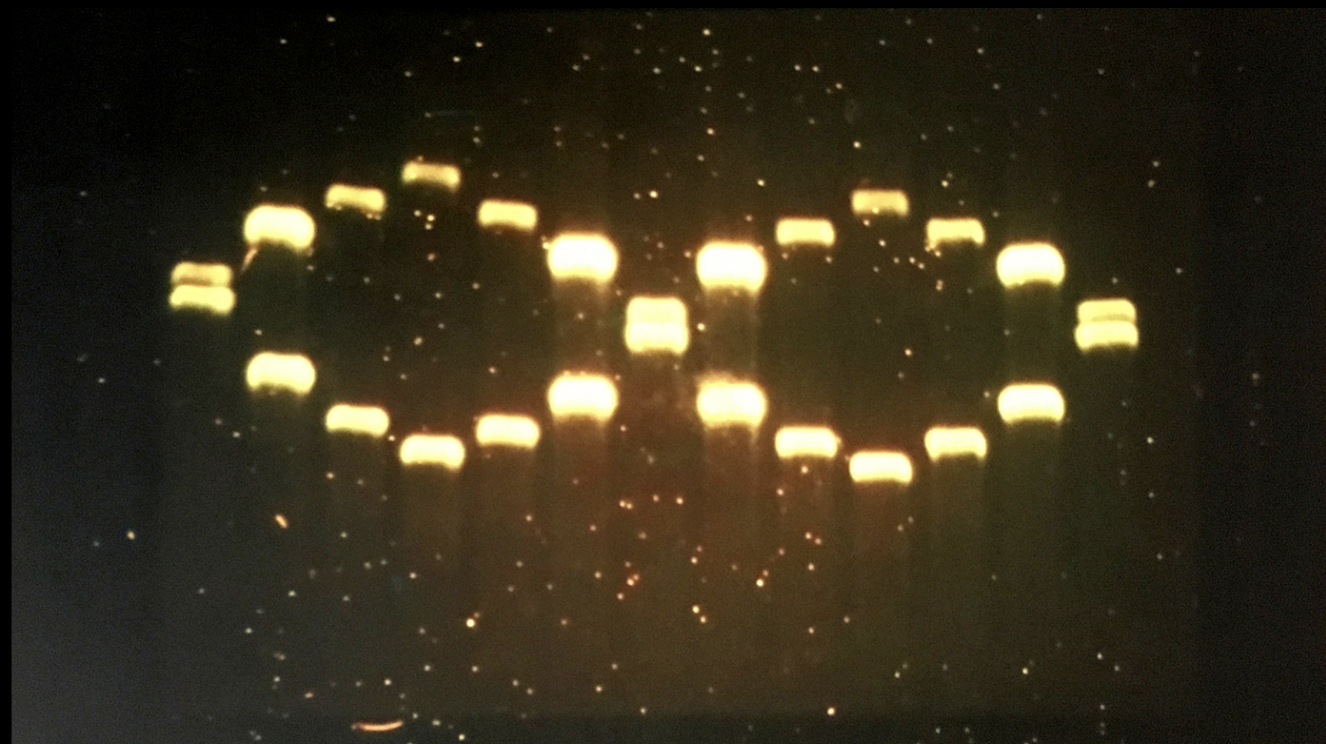
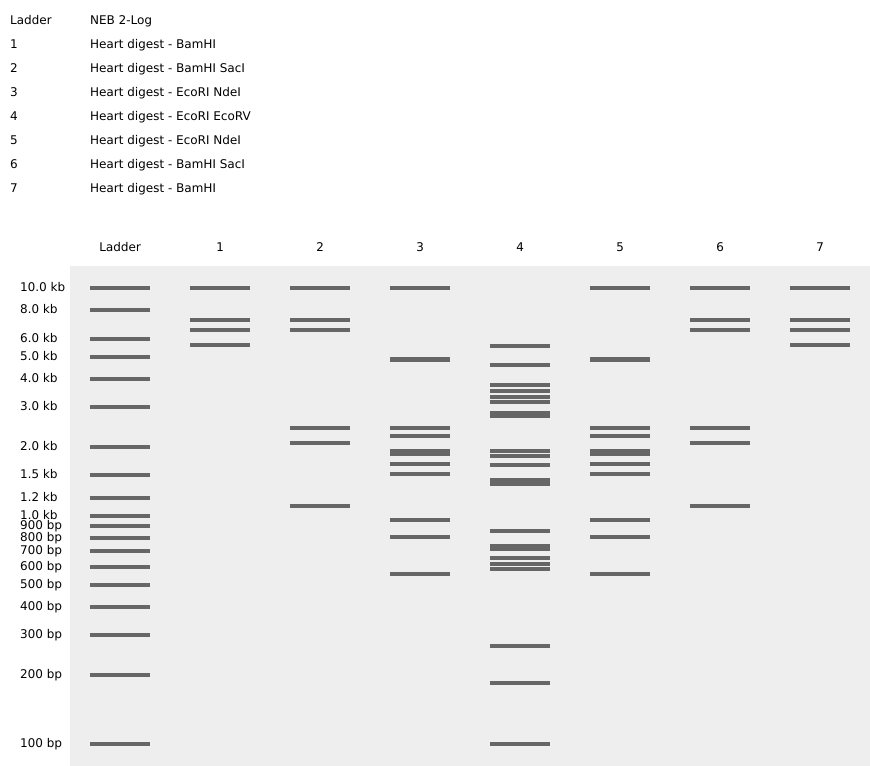
A. Electrophoresis simulation
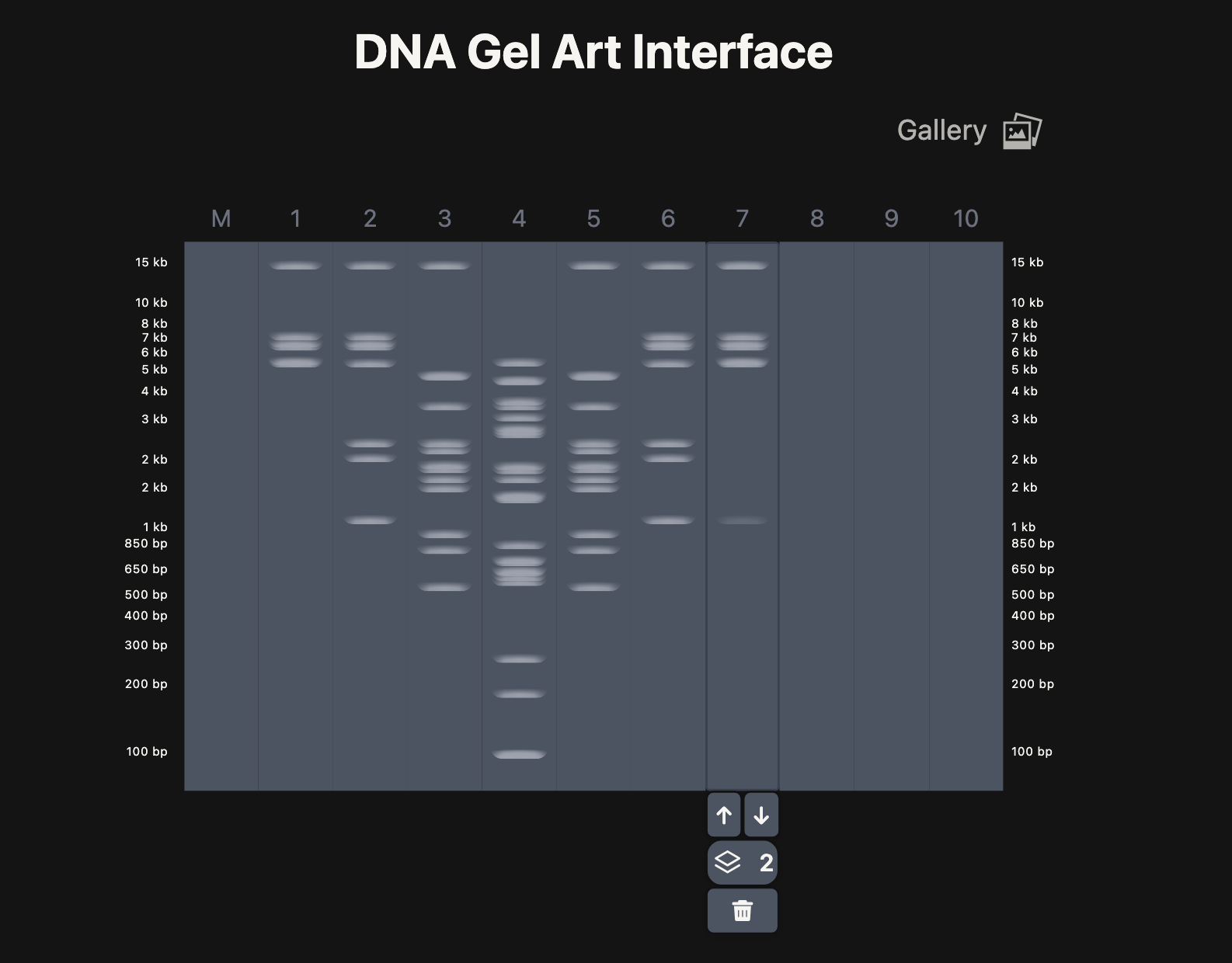

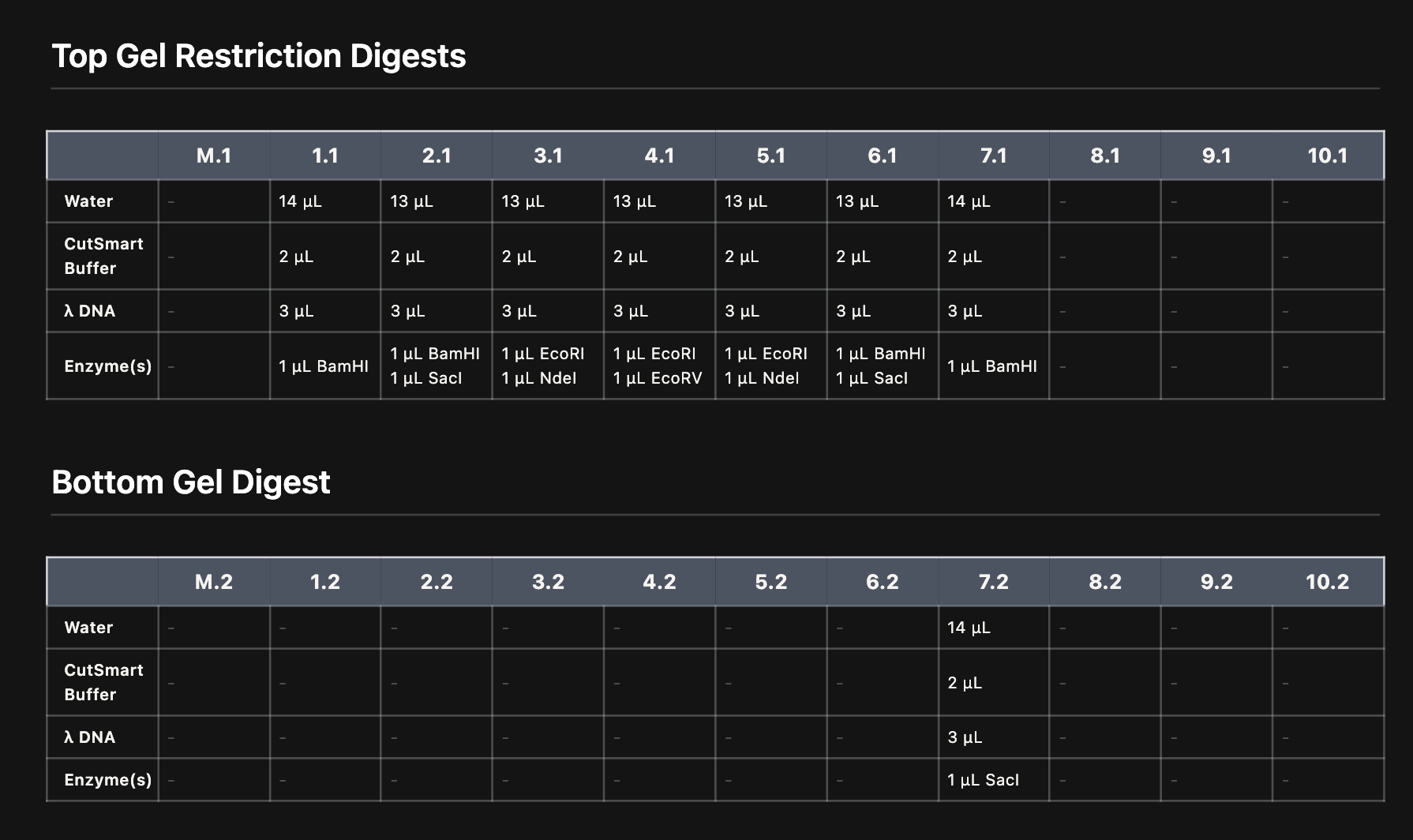

B. Benchling simulation
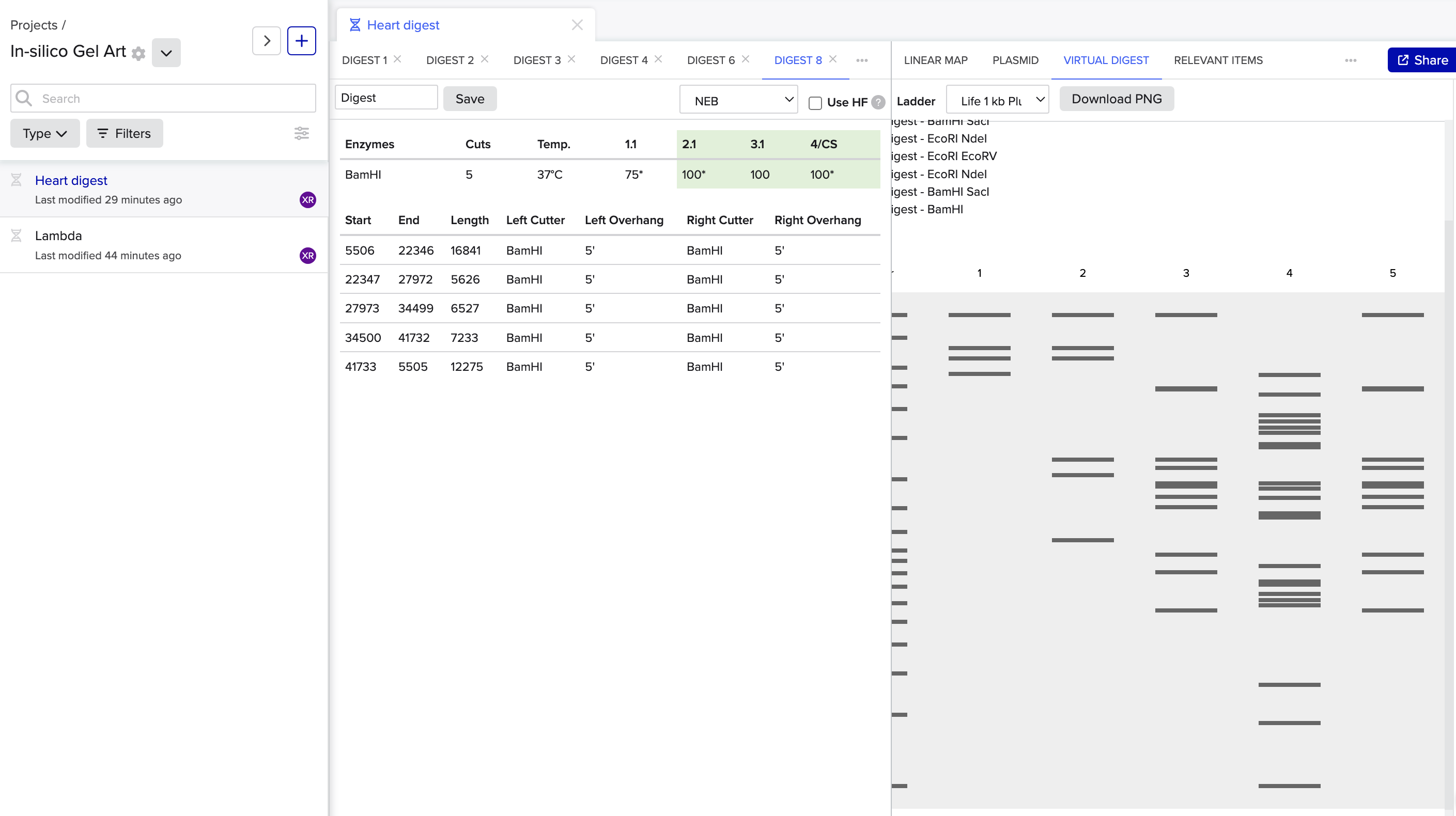

Final Digest: a heart <3

3. DNA design challenge
A. Protein chosen: CDG70667.1 Hydra vulgaris Proto-oncogene Wnt-3 [Hydra vulgaris]
MGTTRYKETLLCFLLIFMETQAQLWMALGTQTSAIESRPRSSINKNLCRALYLHHYQRTVCLNYTDLMLS VAEGIRLGIDECQVQFKHRKWNCTINEHGTSVFGPIITTASRESAFISGIISAGVAFSVTESCAEGKSVH CRCDNSVRGQTDEGWRWGGCNRPITYGIWFSQLFIDQVEKIVKKRKDPRKIMNLHNNKAGREVIKNLLQT ECKCHGTSGNCNLKTCWRSQPHFSEIGKILKEKYDSAHEMEFLYKVKANGERKIKDLIPKYKEYLPPSSL DFIYYEESPNYCVKNETLGIAGTKGRSCNITSSGVDGCELMCCQRGYNVNIVQKTHSCECKFVWCCKVSC NSCIKMTPEYTCK
I selected this protein due to my interest in tissue regeneration, intially my interest began from merismatic cells from plants. In vertebrates, Wtn regulates embryonic development, stem cell maintenance and fibrosis. Studying this protein’s impact in mammals might provide us knowledge about complete tissue regeneration without tumor development.
B. Reverse translation:
CTTTACATAGCAAATAGAAAGGGTCACCGATTAAAGAAAATAATAAAAGGATTCACACGTGCTAATCGTG CGCTTTAAAGATGACTGATTGTTTCTAACTTTATTTGAAATTTCACAAGAAAAGTTTTCACATTAATTTT TAAACACAATGCAAAAATAAACGCAAAGAATATGCAGTTTTAAAACATGCATTTTTAAAGAAAATGGGCA CGACGCGTTATAAAGAAACTTTGTTGTGTTTTTTATTAATTTTTATGGAAACTCAAGCACAACTCTGGAT GGCGCTTGGGACGCAAACGTCAGCAATTGAATCCAGGCCACGTTCATCAATCAATAAAAATTTATGTCGA GCGCTTTATCTTCATCACTACCAAAGAACGGTATGTTTAAATTACACTGATCTAATGTTAAGCGTTGCAG AAGGAATACGACTGGGAATTGACGAATGTCAAGTTCAATTTAAGCACCGTAAATGGAATTGTACGATAAA CGAACATGGAACATCCGTTTTTGGCCCAATTATTACAACAGCCAGCAGAGAAAGTGCATTTATTAGTGGA ATTATATCTGCGGGAGTTGCGTTTTCAGTGACTGAGTCATGTGCAGAAGGAAAATCTGTCCACTGTCGTT GCGATAATAGTGTACGAGGTCAAACGGACGAAGGTTGGCGCTGGGGAGGTTGTAACAGGCCAATCACATA TGGTATATGGTTTTCGCAGTTATTTATTGATCAAGTAGAAAAAATTGTAAAAAAAAGAAAAGATCCACGA AAAATAATGAATCTTCATAACAACAAGGCTGGACGAGAGGTAATAAAAAACCTTTTACAGACTGAATGTA AATGCCACGGAACATCAGGAAACTGCAACTTAAAAACATGCTGGCGTTCACAGCCCCACTTCAGTGAGAT TGGAAAAATACTTAAAGAAAAGTACGATTCAGCTCATGAAATGGAGTTTCTATACAAAGTTAAAGCTAAC GGTGAAAGAAAAATAAAAGACCTTATTCCAAAATATAAAGAATATCTTCCCCCCTCTTCACTGGACTTTA TTTACTATGAGGAATCTCCAAACTACTGCGTAAAAAACGAAACGTTGGGAATAGCAGGAACCAAAGGTCG TTCATGTAACATAACTTCTTCCGGAGTTGACGGTTGCGAACTTATGTGTTGCCAAAGAGGCTATAACGTT AATATTGTACAAAAAACACATTCTTGTGAATGTAAATTTGTATGGTGTTGCAAGGTTTCATGTAATAGCT GCATTAAAATGACGCCTGAATACACCTGTAAATAGTCTCAAAAAAATAAAATTTTAATTCAAATTGAAAA AAGTAATTTCATGATGTTAACACAATTTTAAAGTGTAAACTCTTTTTTTTTTAAGCGTATGTTTATTTAT TTCGGTTGTAAAAATAATTTCATAAAAACGTATATTTATTTAAGTATTGTATAAAATGCATGAATAC
C. Optimized sequence - Rattus norvegicus
CTTTACATAGCAAATAGAAAGGGTCACCGATTAAAGAAAATAATAAAAGGATTCACACGTGCTAATCGTG CGCTTTAAAGATGACTGATTGTTTCTAACTTTATTTGAAATTTCACAAGAAAAGTTTTCACATTAATTTT TAAACACAATGCAAAAATAAACGCAAAGAATATGCAGTTTTAAAACATGCATTTTTAAAGAAAATGGGCA CGACGCGTTATAAAGAAACTTTGTTGTGTTTTTTATTAATTTTTATGGAAACTCAAGCACAACTCTGGAT GGCGCTTGGGACGCAAACGTCAGCAATTGAATCCAGGCCACGTTCATCAATCAATAAAAATTTATGTCGA GCGCTTTATCTTCATCACTACCAAAGAACGGTATGTTTAAATTACACTGATCTAATGTTAAGCGTTGCAG AAGGAATACGACTGGGAATTGACGAATGTCAAGTTCAATTTAAGCACCGTAAATGGAATTGTACGATAAA CGAACATGGAACATCCGTTTTTGGCCCAATTATTACAACAGCCAGCAGAGAAAGTGCATTTATTAGTGGA ATTATATCTGCGGGAGTTGCGTTTTCAGTGACTGAGTCATGTGCAGAAGGAAAATCTGTCCACTGTCGTT GCGATAATAGTGTACGAGGTCAAACGGACGAAGGTTGGCGCTGGGGAGGTTGTAACAGGCCAATCACATA TGGTATATGGTTTTCGCAGTTATTTATTGATCAAGTAGAAAAAATTGTAAAAAAAAGAAAAGATCCACGA AAAATAATGAATCTTCATAACAACAAGGCTGGACGAGAGGTAATAAAAAACCTTTTACAGACTGAATGTA AATGCCACGGAACATCAGGAAACTGCAACTTAAAAACATGCTGGCGTTCACAGCCCCACTTCAGTGAGAT TGGAAAAATACTTAAAGAAAAGTACGATTCAGCTCATGAAATGGAGTTTCTATACAAAGTTAAAGCTAAC GGTGAAAGAAAAATAAAAGACCTTATTCCAAAATATAAAGAATATCTTCCCCCCTCTTCACTGGACTTTA TTTACTATGAGGAATCTCCAAACTACTGCGTAAAAAACGAAACGTTGGGAATAGCAGGAACCAAAGGTCG TTCATGTAACATAA
D & E.
Codon optimization is necessary because different organisms use specific synonymous codons based on tRNA availability and translational efficiency. Although multiple codons encode the same amino acid, using rare codons can reduce protein expression levels and slow translation. By redesigning the nucleotide sequence with common codons in the target model organism, translation efficiency and protein yield can be improved and at the same time preserving the original amino acid sequence of the protein.
I optimized the Hydra Wnt3 coding sequence for expression in Rattus norvegicus, a widely used mammalian model organism. Because Wnt3 is a secreted signaling protein that requires proper folding and post-translational processing, a mammalian system can provide the environment necessary to properly study the impact of this protein and its potential in biomedicine.
Now, when the optimized DNA sequence encoding Hydra Wnt3 is synthesized, it can be cloned into an expression vector containing regulatory elements such as a promoter, transcription start site, and polyadenylation signal. Since it is optimized for a mammal model, a cell-dependent system is best, the plasmid can be introduced into rat-derived cells through transfection. Once inside the cell, RNA polymerase should recognize the promoter and transcribe the inserted gene into messenger RNA (mRNA). The mRNA is then processed and exported to the cytoplasm, where ribosomes translate it into the Wnt3 protein. Because Wnt3 is a secreted signaling protein, it would enter the endoplasmic reticulum, undergo proper folding and post-translational modifications, and be transported through the Golgi apparatus before secretion.
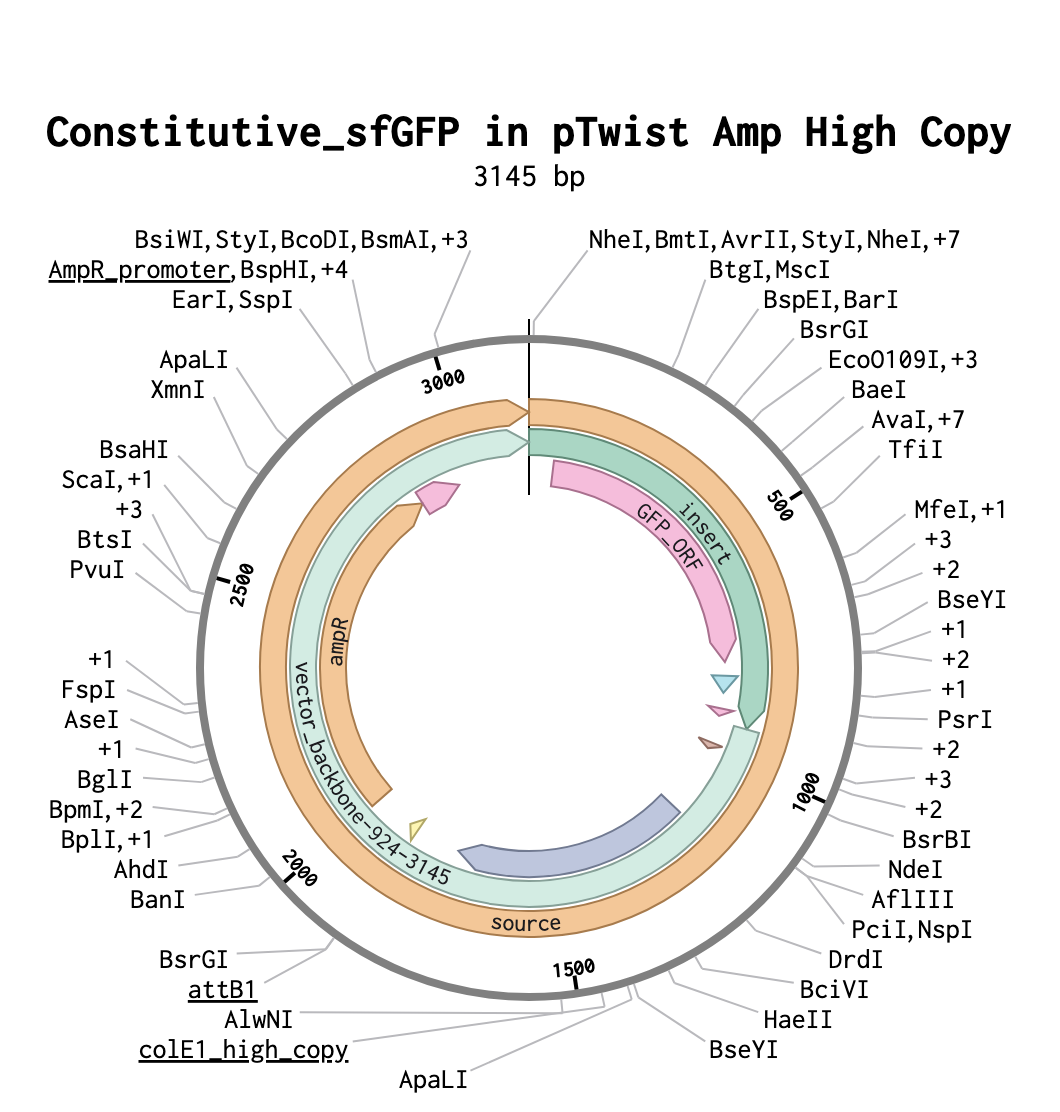
4. Prepare a Twist DNA Synthesis Order
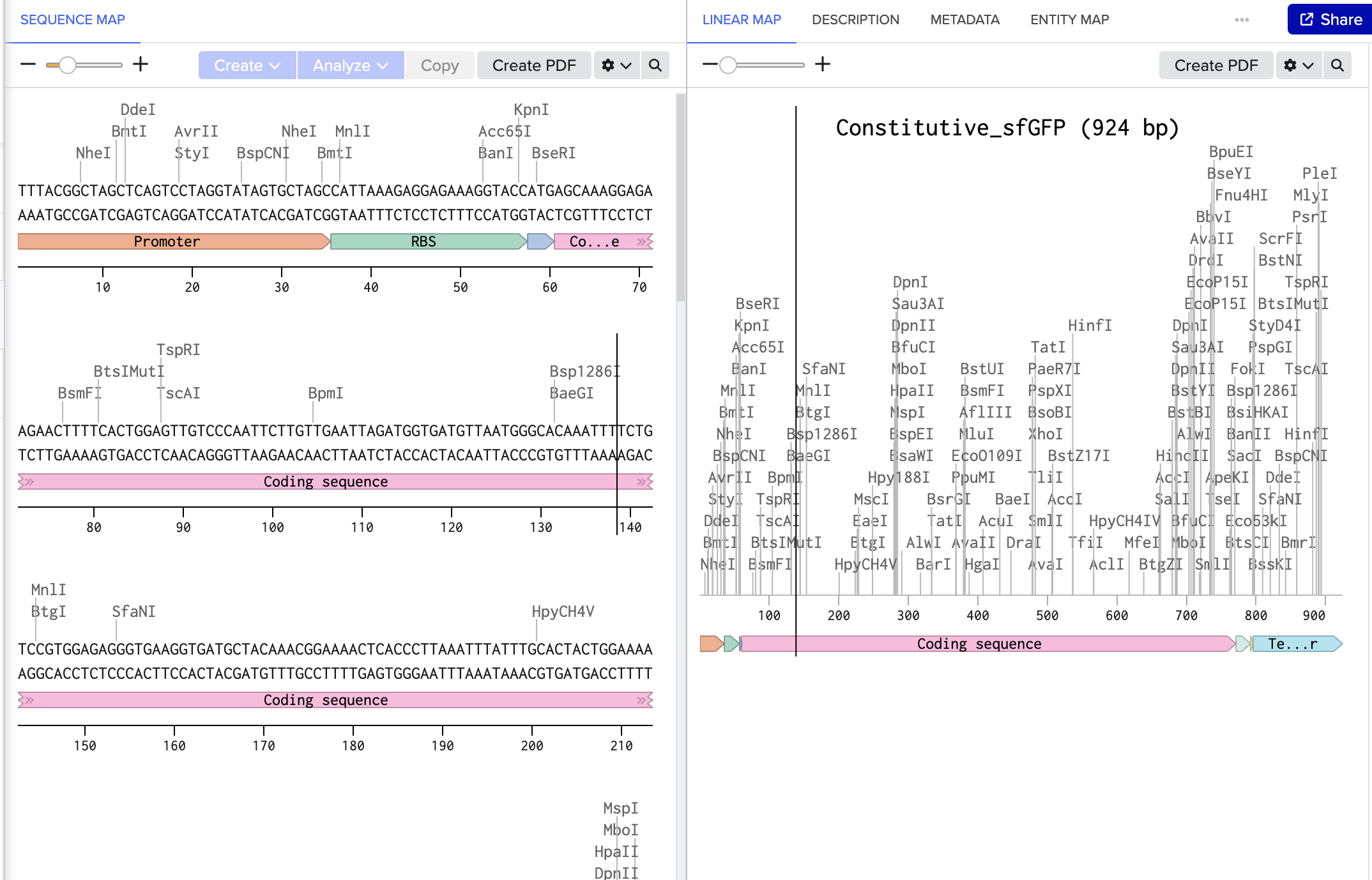

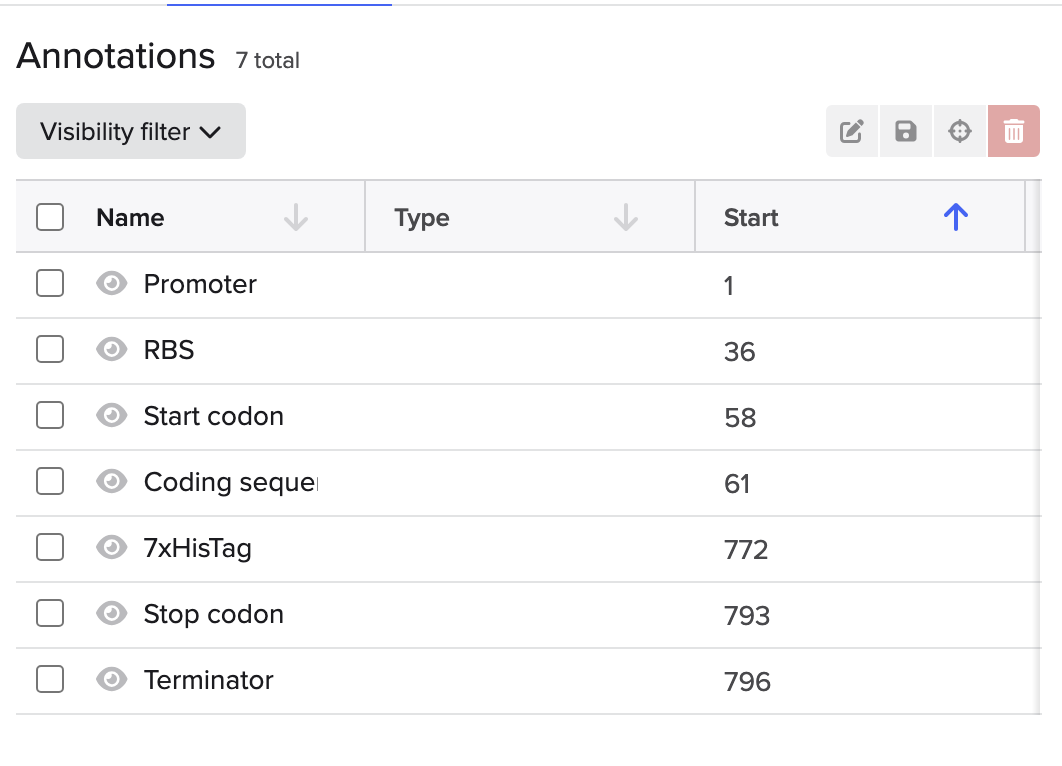

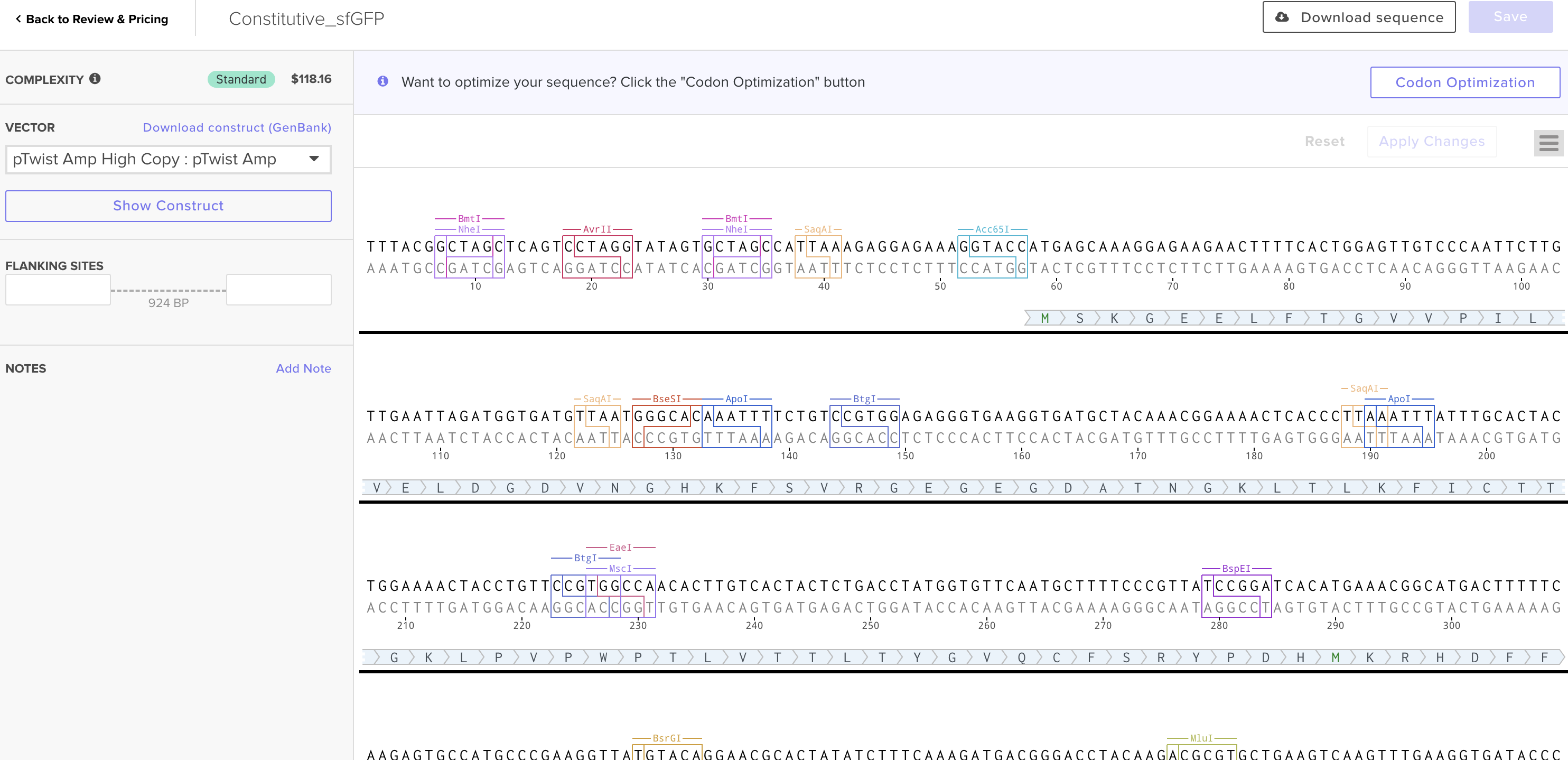

This is the plasmid result :)
5. DNA Read/Write/Edit
DNA read
I would sequence the genomic DNA of Cladosporium sphaerospermum to study it for astrobiology purposes as it is an organisms capable of resisting high levels of radiation. In space or on planetary surfaces like Mars, organisms face chronic ionizing radiation, oxidative stress, desiccation, and nutrient limitation. A fungus that tolerates high radiation is a strong candidate model for understanding which biological features make life resilient in extreme environments.
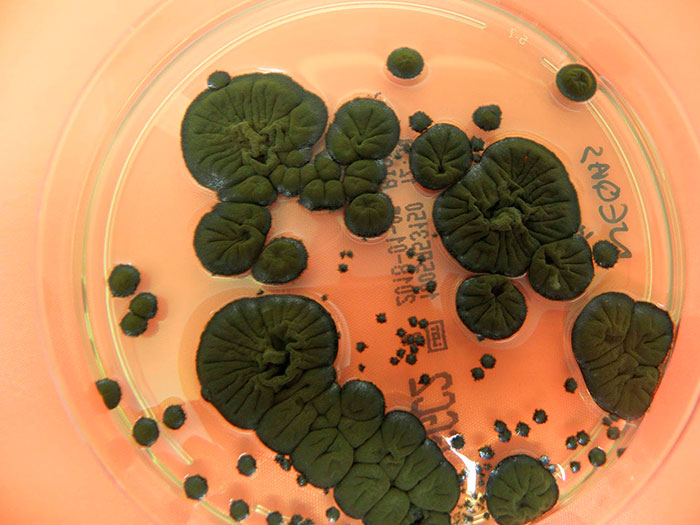


I would sequence it with Oxford Nanopore sequencing (ONT) plus Illumina sequencing-by-synthesis (SBS) as a hybrid strategy. This DNA would let me identify and study genetic systems that can potentially support survival in space-like conditions, such as DNA repair pathways, antioxidant and stress-response networks, and melanin biosynthesis (melanin is known to protect against radiation). I think I would be able to study concepts about long-term survival in other planets, planetary protection, and potential biotech for space habitats like devices or crops.
i. Second-generation sequencing (Illumina SBS)(since it can sequence millions of fragments in parallel using amplification and optical detection during synthesis). ii. Third-generation sequencing (Oxford Nanopore) because it reads single DNA molecules in real time without requiring amplification and can generate very long reads.
This method of sequencing is due that fungal genomes include repeats, variation, and complex regions where long reads are an important part of the genome stability. ONT gives long reads that would improve contiguity and resolve repeats, and Illumina would provide high accuracy to correct residual errors.
–Input: high-quality genomic DNA extracted from fungal cells.
Grow fungal culture and harvest biomass.
Lyse cells and extract genomic DNA.
Purify DNA and assess quality (purity + fragment length).
For Illumina library prep: fragment DNA to short sizes (~200–500 bp), end-repair/A-tailing, adapter ligation, PCR amplification, QC/quantification.
For Nanopore library prep: keep DNA long (minimal fragmentation), ligate Nanopore adapters, load onto flow cell.
–Essential steps of chosen sequencing technology and how does it decode bases:
A. Second-generation sequencing (Illumina SBS)
DNA fragments bind to a flow cell and are clonally amplified into clusters.
During sequencing cycles, fluorescently labeled nucleotides are added one base at a time.
After each cycle, imaging captures the fluorescence signal from each cluster.
The instrument/software assigns the base (A/T/C/G) based on the detected signal, removes the terminator, and repeats for the next base. Base calling is based on fluorescence signals during DNA synthesis.
B. Oxford Nanopore (Nanopore sequencing)
A single DNA molecule passes through a protein nanopore embedded in a membrane.
As different k-mers move through the pore, they cause characteristic disruptions in ionic current.
This method would measure the current changes over time. Base calling would be done by software that converts the electrical signal trace into the DNA sequence.
–Output of sequencing:
The output would consist of sequencing reads (FASTQ files) containing DNA sequences and quality scores. These reads would be assembled into a complete genome (FASTA file) and annotated to identify genes and pathways involved in radiation resistance.


DNA write
I would synthesize a melanin biosynthesis gene from Cladosporium sphaerospermum (such as a polyketide synthase involved in melanin production), it would be codon optimized for expression in a microbial host like Saccharomyces cerevisiae since it is a widely studied model.

Since melanin has been associated with radiation tolerance in fungi found in high-radiation environments. Engineering microbes to produce melanin could help us study biological radiation protection and explore potential applications for space habitats or planetary environments like Mars.
The construct would include: a promoter, ribosome binding site, the codon-optimized melanin gene and a terminator sequence
I would use chip-based oligonucleotide synthesis followed by enzymatic gene assembly, such as the silicon-based platform described in class (Twist Bioscience). This would allow: parallel production of multiple constructs and it would be cost-effective.
–Essential steps:
Design and codon-optimize the gene sequence.
Chemically synthesize short oligonucleotides on a chip.
Amplify the oligo pool.
Enzymatically assemble the full-length gene.
Clone into a plasmid vector.
Sequence-verify the final construct.
–Limitations:
Error rate: Chemical synthesis can introduce mutations, requiring sequence verification.
Length limitations: Very long or repetitive sequences are more difficult to synthesize.
Complexity: High GC content or secondary structure can reduce synthesis efficiency.
DNA Edit

I would edit the genome of Saccharomyces cerevisiae to enhance its tolerance to ionizing radiation and oxidative stress. Since fungi such as Cladosporium sphaerospermum are known to survive in high-radiation environments, I would attempt to transfer or optimize specific protective mechanisms in yeast. (Introducing melanin biosynthesis genes (polyketide synthase)). This would allow us to experimentally test how specific genetic changes contribute to radiation resilience which as discussed previously has implications for astrobiology, planetary protection, and biotechnology applications in space habitats. Apart from astrobiological applications, this could also be applied to conservation biology or restoration efforts, such as modifying species for increased climate resilience.
For yeast genome editing I would use CRISPR–Cas9 technology.
–Essential steps
Design a guide RNA (gRNA) complementary to the target genomic sequence of the yeast
Cas9 nuclease is directed by the gRNA to the target locus.
Cas9 introduces a double-strand break.
The cell repairs the break via: Homology-directed repair (HDR), with a donor template containing homologous sequences.
–Inputs: Designed guide RNA sequence, Cas9 protein, donor DNA template with homology arms, yeast cells, selectable marker, PCR primers for verification
–Design
Identify safe genomic integration site
Design guide RNA and donor DNA construct with flanking homology arms.
Clone into plasmid or assemble as linear repair template.
Transform yeast and select edited clones.
Validate edits via PCR and sequencing.
–Limitations
Variable HDR efficiency
Potential genomic instability if inserting large constructs
Fitness cost of engineered traits