Ximena Roman Grajeda — HTGAA Spring 2026




Week 1 HW: Principles and Practices
First, describe a biological engineering application or tool you want to develop and why. This could be inspired by an idea for your HTGAA class project and/or something for which you are already doing in your research, or something you are just curious about: I would be interested in developing an approach to study and properly map interactions and pathways between the gut mycobiome and host gut cells. Specifically to examine how fungal components contribute to chronic inflammation and downstream cellular stress responses. This approach would contribute to the characterization of direct and indirect mycobiome inflammation pathways that are linked to disease (cancer, IBD) origin and progression.
For this homework I used gemini AI within colab to understand how the process worked as well as to achieve the design I had done in the Donovan simulation website. Once I was using colab, the design changed a lot since I could instruct my design better, but I also realized that making a planet desing was much more challenging than I initially thought. While doing this code I ran into some errors due cross contamination, which gemini helped me in localizing where the code did not indicate the robot to drop the tip, it was very helpful.

I would be interested in developing an approach to study and properly map interactions and pathways between the gut mycobiome and host gut cells. Specifically to examine how fungal components contribute to chronic inflammation and downstream cellular stress responses. This approach would contribute to the characterization of direct and indirect mycobiome inflammation pathways that are linked to disease (cancer, IBD) origin and progression.
A. Governance role 1 (Safety and security-Responsible research): Prevent misuse or unsafe utilization of mycobiome research. a1. Sub-goal 1: prevent wrongful application of findings outside the context they were designed for: to make sure that findings are not used as direct causal proof or direct clinical input. Responsible research will also involve proper methodologies and expertise input. a2. Sub-goal 2: ensure clear communication of model limitations: this to ensure that understanding of what exactly the model’s goals are and what it insight will be able to provide about the disease or other possible research areas
B. Governance role 2 (Equity): Ensuring equitable and representative research in mycobiome research models. b1. Sub-goal 1: Encourage inclusion of diverse mycobiome profiles in research design: so that models can reflect more accurately the variability of the mycobiome across populations. b2 Sub-goal 2: Ensure that model explicitly acknowledge variability of the mycobiome: to prevent results from being presented as universally representative of all human gut.
Actions:
A. Standardized reporting guidelines for mycobiome-based models
a. Purpose: Commonly, human gut microbiome is studied and it includes bacteria, viruses and fungi. Since fungi are biologically distinct than viruses and bacteria, reporting on methodology of research models should be set on their own category to better understand the behaviour of this fungi in in models and their roles in the gut. This distinction is important to assess the exact roles of bacteria, viruses and fungi in the human gut and to be able to pinpoint what fungal related pathways can be better charatcerized in relation to chronic inflammation. This would enforce that all biological systems in human microbiome research are represented equitably.
b. Design: Academic Journals and scientific institutions would be reporting checklists for mycobiome studies. Authors and contributors will need to include sections addressing fungal specific assumptions and limitations, which reviewers would asses during peer review process.
c.Assumptions: This action would assume that journals will make sure these guidelines are followed, that researchers successfully dispose limitations of the models, clearer reporting would improve interpretation for research replication.
d. Risk of failure and success: -Risk of failure: Reporting requirements may become superficial or inconsistent if they are not properly enforced by journals and institutions and mycobiome focused research might become diluted in microbiome studies. -Risk of success: This might be beneficial for small laboratories that want to focus their research on the mycobiome as a category inside the microbiome instead of the whole biological system that the microbiome is and provide access to reproducible research. It would also open possibilities for collaboration with labs focusing on bacteria or viruses to form equitable involvement of biological systems.
B. Required combined fungal biology and human biology expertise
a. Purpose: If there is mycobiome research proposed to be completed with the goal of reaching biomedical applications, the research cannot only be done by microbiologists due that the human gut involves knowledge of human biology. This action proposes that research addressinf fungal-host interactions in disease contexts is informed by expertise in both fungal biology, and human bioology, particularly when immune signalign and cellular stress are involved.
b. Design: Research teams, institutions and funding bodies would require mycobiome projects claiming relevance to human health would be exoected to include collaborators or consultants with fungal biology and human biology expertise or justify how this expertise is incorportated into their study design. Journals could further assess and confirm that appropiate expertise is oncluded when evaluating study scope. In case of missing expertise in study design scientific boards and universities could direct them to experts.
c. Assumptions: This action would assume that there is relevant expetise is included in research team before beggining experimentation and that revirewers and intitutions reasonably evalute whether this expertise is sufficient. It would also assume that experts involved are capable of carrying out similar research or have done it before.
d. Risk of failure and success: -Risk of failure: Expertise requirements might just be applied briefly or superficially for co authoring purposes, smaller labs may face barriers to collaboration, reviewers may inconsistently assess what counts as adequate expertise. -Risk of success: Increases collaboration requirements could slow research advances or interdisciplinary work may become performative. In the other hand if collaboration proves to be well organized there would be sucessful exchange of quality scientific knowledge and be and example of inclusive collaborative research and correct disctinction of all biological systems participating.
C. Limits on clinical language in mycobiome research
a. Purpose: Many studies that involve immune signaling or potential diseas related processes coould be framed using clinical or therapeutic langiage even when findings are model based and have not yet been appplied in clinical research. This can lead to misunderstandings of findings applications and contribute to misunformation when results are communicated to the public. This action would restric thr use of clinical language unless studies meet defined standards that help to distinguish exploratory research from apploed or clinical claims.
b. Design: Insitutions, science boards and journals would implement guidelines that discurage the use of terms implying treatment, prevention,or clinical efficacy unless it is supported. They would also require explicit labelling of studies as exploratory or non-clinical.
c. Assumptions: This action would assume that language can influence interpetation by both scientific and public audiences. That institutions, jorunals and scientific communities will enfornce the language guideliunes and that researches will be able to communicate their significance without utilizng clinical language.
d. Risk of failure and success: -Risk fo failure: Guidelines might not be consistently enforced. Researches might use other wording impliying clinical relevance. Public misunderstanding will still exists regardless of langiage guidelines. -Risk of success: overly strict language might disocurage translational discussion (although translational potential is not to be confused with claiming clinical relevance). Important findings might be perceived as less impactful due framing and many studies might be dismissed if not read aproppiately.
| Actions: | Action 1 | Action 2 | Action 3 |
|---|---|---|---|
| Enhance accurate science communication | |||
| •By preventing overlooking data | 1 | 1 | 1 |
| •By enhancing understanding | 1 | 1 | 1 |
| of specific biological systems | |||
| Foster Lab Safety | |||
| • By preventing incident | 2 | 1 | 3 |
| • By helping respond | 2 | 1 | 3 |
| Promote scientific collaboration | |||
| • By preventing incidents | 3 | 1 | 3 |
| • By helping respond | 3 | 1 | 2 |
| Other considerations | |||
| • Minimizing costs and burdens to stakeholders | 3 | 3 | 2 |
| • Feasibility? | 1 | 2 | 2 |
| • Not impede research | 2 | 2 | 2 |
| • Promote constructive applications | 1 | 1 | 1 |
Based on the scoring I would prioritize Action 2 (Requiring combines fungal biology and human biology expertise) supported by Action 3 (placing limits on clinical language). Action 2 scored highly in policy goals related to scientific accuracy, lab safety, and collaboration, which suggetss that expertise is the most effective way to reduce misinterpretation, methodological errors and to avoid the dismissal of important data. Action 3 complements this since it adresses downstream risks of misinformation and misapplication, especially when research findings are communicated beyong the scientific community.
Action 1 (standarized reporting guidelines) scored high on improving communication, it scored lower in terms of cost, burden, and feasibility. This tells us that reporting standards might cause delays in research and may be most effective as a secondary long term action rather than a priority.
By prioritizing action 2, I noticed a trade off between impriving scientific accuracy and the potential burden on smaller laboratories, which may find interdisciplinary collaboration more difficult to enfornce. Action 3 assumes that limiting clinical language will meaningfully reduce public misunderstanding as well as misundersatnding in broader academic communties, though it will remain uncertain how research findings are interpreted once they leave academic contexts.
These recommendations assume thay appropiate expertise is avaliable and that journals and relevant institutions will be able to evaluate appropiately both disciplinary competence and laguage use. I do not think this actions will eliminate the issue but I would expoect them to reduce risks without stalling or impeding research progress.
—-> Questions from professor Jacobson:
DNA polymerase error is 1 per 10⁶ bases added. The human genome is around 3.1 billion base pairs. So if polymerase made one error per 10⁶ bases without corrections there would be thousands of nucleotide misplacements and life would not be viable.
Biology deals with this discrepancy by different systems that reduce errors.
a. DNA polymerases have 3’ to 5’ exonuclease activity which allows them to detect nucleotide misplacements, remove them and place a correct match. b. To lessen the misplacements even more there exists a mechanism called Post replication mismatch repair (MMR) that takes place after replication specifically through the MutS Repair system. In this system specialized proteins, MutS detects mismatches, to allow Pol III and ligase to synthesize correct matches.
The number of possible ways to code for a human protein is very large, this is due of the degeneracy of the genetic code that allows different codon triplets to code for the same amino acid. The third possiton of the codon is flexible and is able to bind with other bases and even though the codon triplets are different: GCC, GCU, GCA, GCG all code for alanine.
The reasons that even if all of these codons code for the same amino acid but don’t produce a functional protein:
a. mRNA secondary structure: Different codon arrangements change the mRNA sequence and alter how the strand folds, this can cause the strands to fold and form strucutres near the ribosome binding site or nearby the start codons. This would prevent the ribosome from binding and unwinding the mRNA for translation.
b. Different cells have different tRNAs: if there are many strange combinations of codon triplets and a cell does not have enough tRNA to code for them, the result will be an incomplete peptide or a unstable petide that will be prone to degradation.
c. GC content: if there is a very high or low content of GC in a sequence, it can affect folding, coding efficiency, ribosome activity or mRNA stability.It is also known that high GC content can cause the creation of ridig structures affecting their function.
—-> Questions from Dr. LeProust:
The most commonly used method is Phosphoramidite DNA Synhesis cycle.
Phosphoramidite synthesis has no proofreading and a non zero error rate (it is small but non-zero, they can be deletions, side reactions or incomplete reactions) meaning that long oligos could accumulate several errors and final molecule would contain them since there is no mechanism to fix them, at that point purification would be lenghty and impractical.
Attempting to make a 2000bp would accumulate errors exponentially since there is no proofreading mechanism, so we would obtain faulty sequences. Assembly would make several oligos of smaller size, they would accumulate less errors which can then be corrected and assembled via enzymatic processes.
—-> Question from George Church
The 10 essential amino acids are Histidine, Isoleucine, Leucine, Lysine, Methionine, Phenylalanine, Threonine, Tryptophan, Valine and Arginine. Animals including humans cannot synthesize Lysine molecules, they must obtain it from other sources through their diet. Plants and some microbes are able to synthesize lysine, so if consumed animals are able to supply their lysine.
This understanding challenges the concept of “lysisne contingency” that was seen in the movies of Jurassic Park, where Henry Wu performed a genetic alteration in the dinosaur genome supposedly knocking down the ability to produce the amino acid lysine. This was a plan made for it that anything went wrong, the dinosaurs were not able to escape and survive long to disrupt ecosystems. It was assumed they would die due the lack of lysine.
Since we know that lysine is an essential amino acid, meaning that animals cannot synthesize it. It is nonsensical that they would say that they performed this genetic alteration, animals (including dinosaurs I imagine) were already unable to produce lysine, so no genetic alteration could have been made.
In any case, if we took that this dinosaurs in that world were supposed to be able to produce lysine but they were genetically altered to not be able to synthesize it, hervibores would be living well, carnivores fed through cattle would be living well, as I suppose they did not want their carnivores eating anything else but the food provided (also not hervibores). But if they escape, carnivores will eat hervibores, so they all survive either way.
The lysine contigency idea does not work. But I believe the author of jurassic park, Michael Chrichton wrote it this way as a symbol of corporate overconfidence and musinderstanding of biological systems. It is is a recurring theme in Michael Chrichton’s work to critique political roles and intitutions, including one of my favorite novels “The Andromeda Strain”.




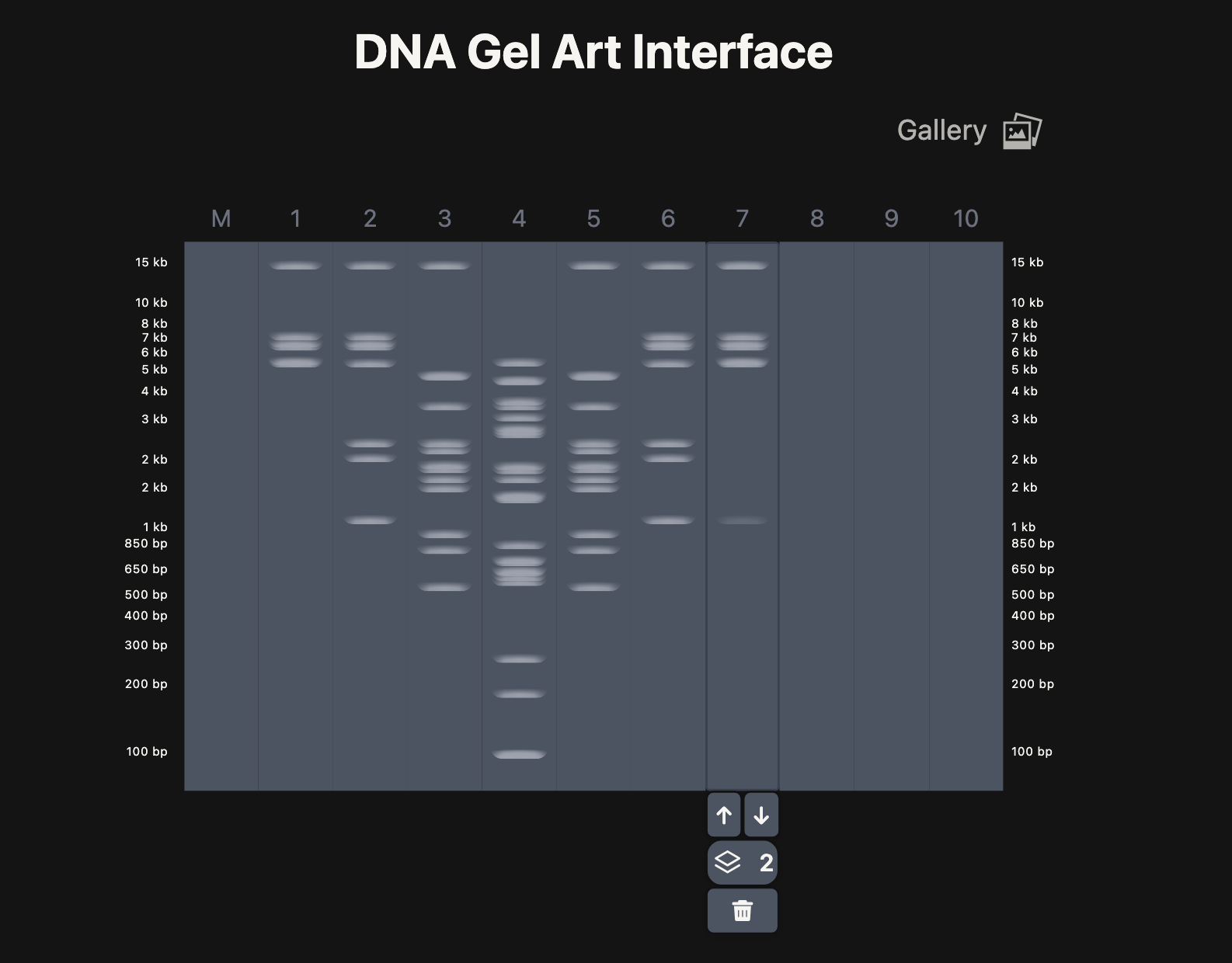
A. Electrophoresis simulation

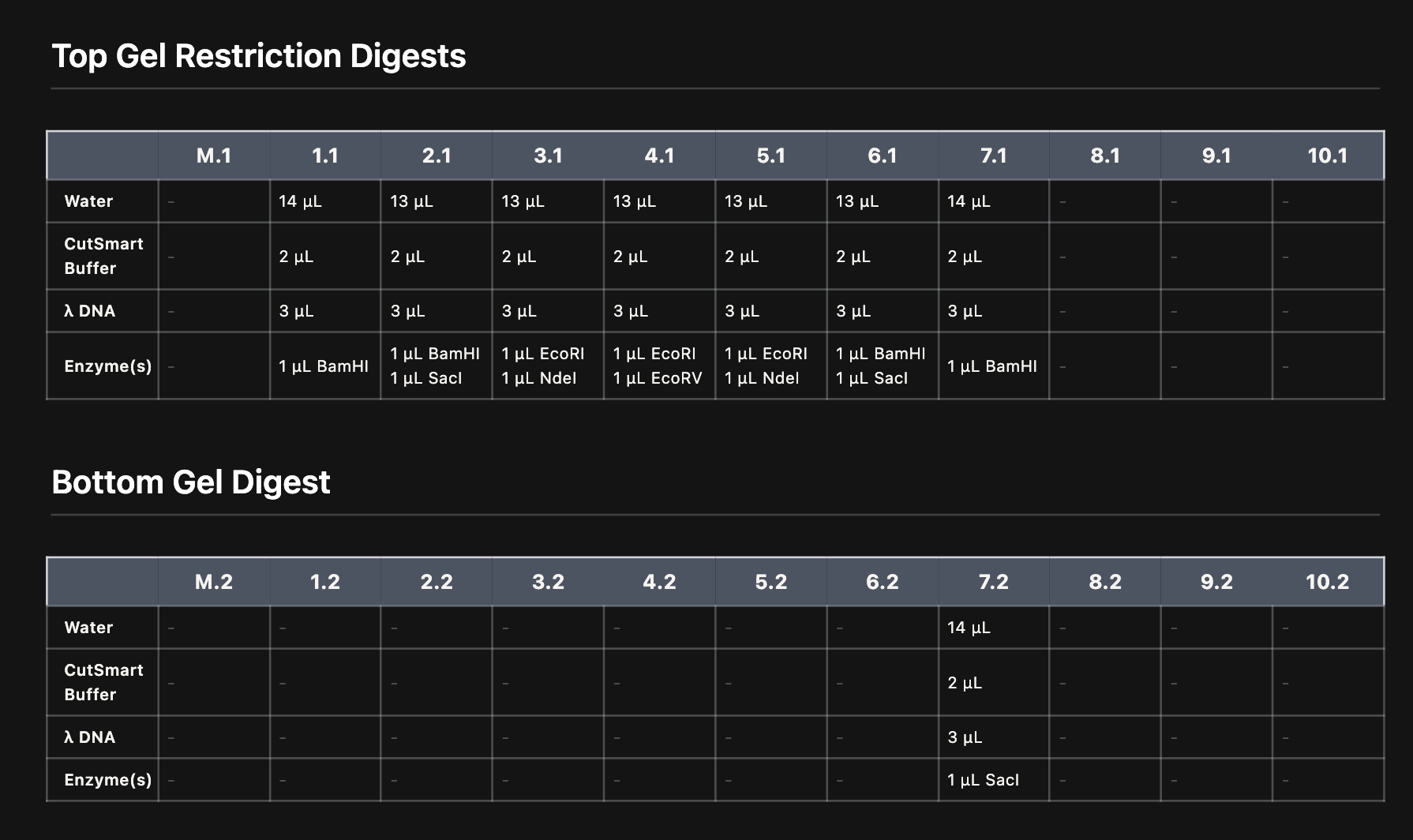

B. Benchling simulation
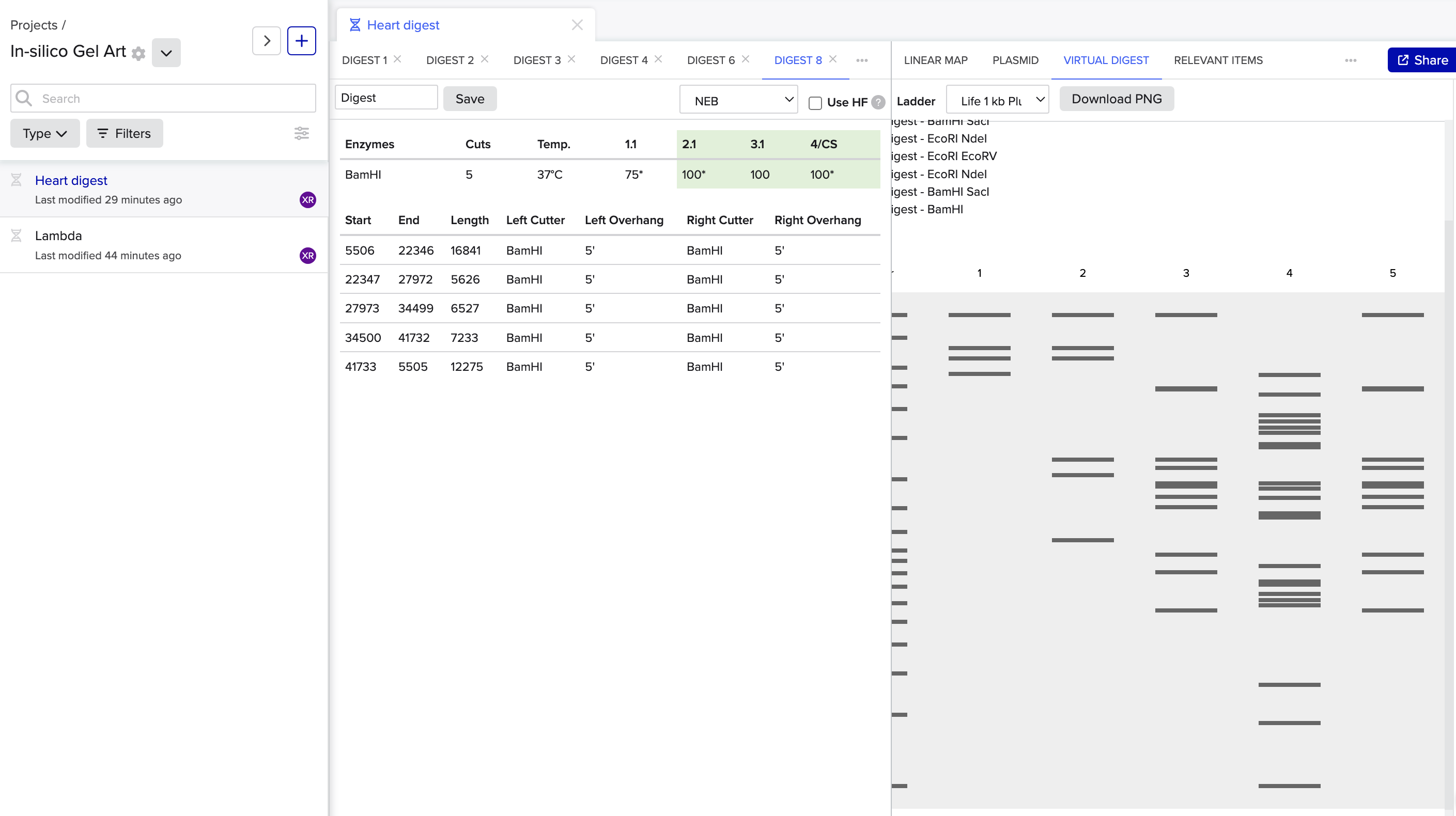

Final Digest: a heart <3
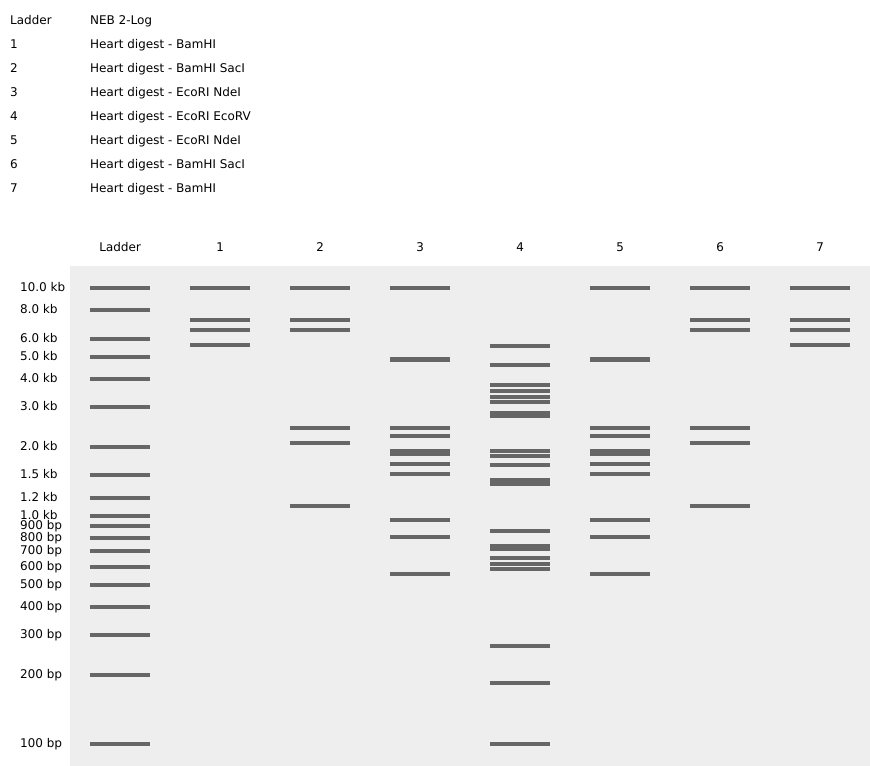

A. Protein chosen: CDG70667.1 Hydra vulgaris Proto-oncogene Wnt-3 [Hydra vulgaris]
MGTTRYKETLLCFLLIFMETQAQLWMALGTQTSAIESRPRSSINKNLCRALYLHHYQRTVCLNYTDLMLS VAEGIRLGIDECQVQFKHRKWNCTINEHGTSVFGPIITTASRESAFISGIISAGVAFSVTESCAEGKSVH CRCDNSVRGQTDEGWRWGGCNRPITYGIWFSQLFIDQVEKIVKKRKDPRKIMNLHNNKAGREVIKNLLQT ECKCHGTSGNCNLKTCWRSQPHFSEIGKILKEKYDSAHEMEFLYKVKANGERKIKDLIPKYKEYLPPSSL DFIYYEESPNYCVKNETLGIAGTKGRSCNITSSGVDGCELMCCQRGYNVNIVQKTHSCECKFVWCCKVSC NSCIKMTPEYTCK
I selected this protein due to my interest in tissue regeneration, intially my interest began from merismatic cells from plants. In vertebrates, Wtn regulates embryonic development, stem cell maintenance and fibrosis. Studying this protein’s impact in mammals might provide us knowledge about complete tissue regeneration without tumor development.
B. Reverse translation:
CTTTACATAGCAAATAGAAAGGGTCACCGATTAAAGAAAATAATAAAAGGATTCACACGTGCTAATCGTG CGCTTTAAAGATGACTGATTGTTTCTAACTTTATTTGAAATTTCACAAGAAAAGTTTTCACATTAATTTT TAAACACAATGCAAAAATAAACGCAAAGAATATGCAGTTTTAAAACATGCATTTTTAAAGAAAATGGGCA CGACGCGTTATAAAGAAACTTTGTTGTGTTTTTTATTAATTTTTATGGAAACTCAAGCACAACTCTGGAT GGCGCTTGGGACGCAAACGTCAGCAATTGAATCCAGGCCACGTTCATCAATCAATAAAAATTTATGTCGA GCGCTTTATCTTCATCACTACCAAAGAACGGTATGTTTAAATTACACTGATCTAATGTTAAGCGTTGCAG AAGGAATACGACTGGGAATTGACGAATGTCAAGTTCAATTTAAGCACCGTAAATGGAATTGTACGATAAA CGAACATGGAACATCCGTTTTTGGCCCAATTATTACAACAGCCAGCAGAGAAAGTGCATTTATTAGTGGA ATTATATCTGCGGGAGTTGCGTTTTCAGTGACTGAGTCATGTGCAGAAGGAAAATCTGTCCACTGTCGTT GCGATAATAGTGTACGAGGTCAAACGGACGAAGGTTGGCGCTGGGGAGGTTGTAACAGGCCAATCACATA TGGTATATGGTTTTCGCAGTTATTTATTGATCAAGTAGAAAAAATTGTAAAAAAAAGAAAAGATCCACGA AAAATAATGAATCTTCATAACAACAAGGCTGGACGAGAGGTAATAAAAAACCTTTTACAGACTGAATGTA AATGCCACGGAACATCAGGAAACTGCAACTTAAAAACATGCTGGCGTTCACAGCCCCACTTCAGTGAGAT TGGAAAAATACTTAAAGAAAAGTACGATTCAGCTCATGAAATGGAGTTTCTATACAAAGTTAAAGCTAAC GGTGAAAGAAAAATAAAAGACCTTATTCCAAAATATAAAGAATATCTTCCCCCCTCTTCACTGGACTTTA TTTACTATGAGGAATCTCCAAACTACTGCGTAAAAAACGAAACGTTGGGAATAGCAGGAACCAAAGGTCG TTCATGTAACATAACTTCTTCCGGAGTTGACGGTTGCGAACTTATGTGTTGCCAAAGAGGCTATAACGTT AATATTGTACAAAAAACACATTCTTGTGAATGTAAATTTGTATGGTGTTGCAAGGTTTCATGTAATAGCT GCATTAAAATGACGCCTGAATACACCTGTAAATAGTCTCAAAAAAATAAAATTTTAATTCAAATTGAAAA AAGTAATTTCATGATGTTAACACAATTTTAAAGTGTAAACTCTTTTTTTTTTAAGCGTATGTTTATTTAT TTCGGTTGTAAAAATAATTTCATAAAAACGTATATTTATTTAAGTATTGTATAAAATGCATGAATAC
C. Optimized sequence - Rattus norvegicus
CTTTACATAGCAAATAGAAAGGGTCACCGATTAAAGAAAATAATAAAAGGATTCACACGTGCTAATCGTG CGCTTTAAAGATGACTGATTGTTTCTAACTTTATTTGAAATTTCACAAGAAAAGTTTTCACATTAATTTT TAAACACAATGCAAAAATAAACGCAAAGAATATGCAGTTTTAAAACATGCATTTTTAAAGAAAATGGGCA CGACGCGTTATAAAGAAACTTTGTTGTGTTTTTTATTAATTTTTATGGAAACTCAAGCACAACTCTGGAT GGCGCTTGGGACGCAAACGTCAGCAATTGAATCCAGGCCACGTTCATCAATCAATAAAAATTTATGTCGA GCGCTTTATCTTCATCACTACCAAAGAACGGTATGTTTAAATTACACTGATCTAATGTTAAGCGTTGCAG AAGGAATACGACTGGGAATTGACGAATGTCAAGTTCAATTTAAGCACCGTAAATGGAATTGTACGATAAA CGAACATGGAACATCCGTTTTTGGCCCAATTATTACAACAGCCAGCAGAGAAAGTGCATTTATTAGTGGA ATTATATCTGCGGGAGTTGCGTTTTCAGTGACTGAGTCATGTGCAGAAGGAAAATCTGTCCACTGTCGTT GCGATAATAGTGTACGAGGTCAAACGGACGAAGGTTGGCGCTGGGGAGGTTGTAACAGGCCAATCACATA TGGTATATGGTTTTCGCAGTTATTTATTGATCAAGTAGAAAAAATTGTAAAAAAAAGAAAAGATCCACGA AAAATAATGAATCTTCATAACAACAAGGCTGGACGAGAGGTAATAAAAAACCTTTTACAGACTGAATGTA AATGCCACGGAACATCAGGAAACTGCAACTTAAAAACATGCTGGCGTTCACAGCCCCACTTCAGTGAGAT TGGAAAAATACTTAAAGAAAAGTACGATTCAGCTCATGAAATGGAGTTTCTATACAAAGTTAAAGCTAAC GGTGAAAGAAAAATAAAAGACCTTATTCCAAAATATAAAGAATATCTTCCCCCCTCTTCACTGGACTTTA TTTACTATGAGGAATCTCCAAACTACTGCGTAAAAAACGAAACGTTGGGAATAGCAGGAACCAAAGGTCG TTCATGTAACATAA
D & E.
Codon optimization is necessary because different organisms use specific synonymous codons based on tRNA availability and translational efficiency. Although multiple codons encode the same amino acid, using rare codons can reduce protein expression levels and slow translation. By redesigning the nucleotide sequence with common codons in the target model organism, translation efficiency and protein yield can be improved and at the same time preserving the original amino acid sequence of the protein.
I optimized the Hydra Wnt3 coding sequence for expression in Rattus norvegicus, a widely used mammalian model organism. Because Wnt3 is a secreted signaling protein that requires proper folding and post-translational processing, a mammalian system can provide the environment necessary to properly study the impact of this protein and its potential in biomedicine.
Now, when the optimized DNA sequence encoding Hydra Wnt3 is synthesized, it can be cloned into an expression vector containing regulatory elements such as a promoter, transcription start site, and polyadenylation signal. Since it is optimized for a mammal model, a cell-dependent system is best, the plasmid can be introduced into rat-derived cells through transfection. Once inside the cell, RNA polymerase should recognize the promoter and transcribe the inserted gene into messenger RNA (mRNA). The mRNA is then processed and exported to the cytoplasm, where ribosomes translate it into the Wnt3 protein. Because Wnt3 is a secreted signaling protein, it would enter the endoplasmic reticulum, undergo proper folding and post-translational modifications, and be transported through the Golgi apparatus before secretion.
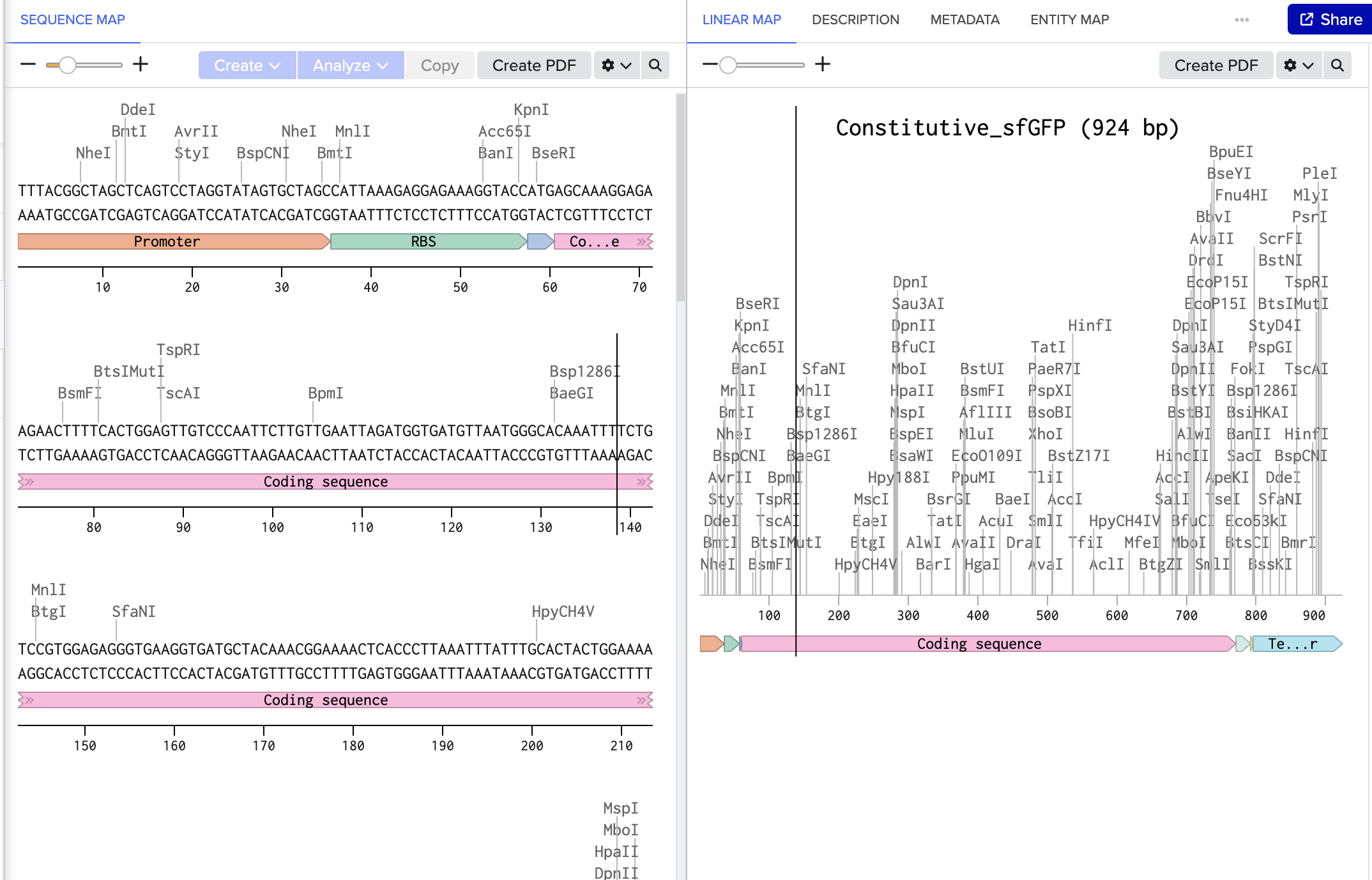

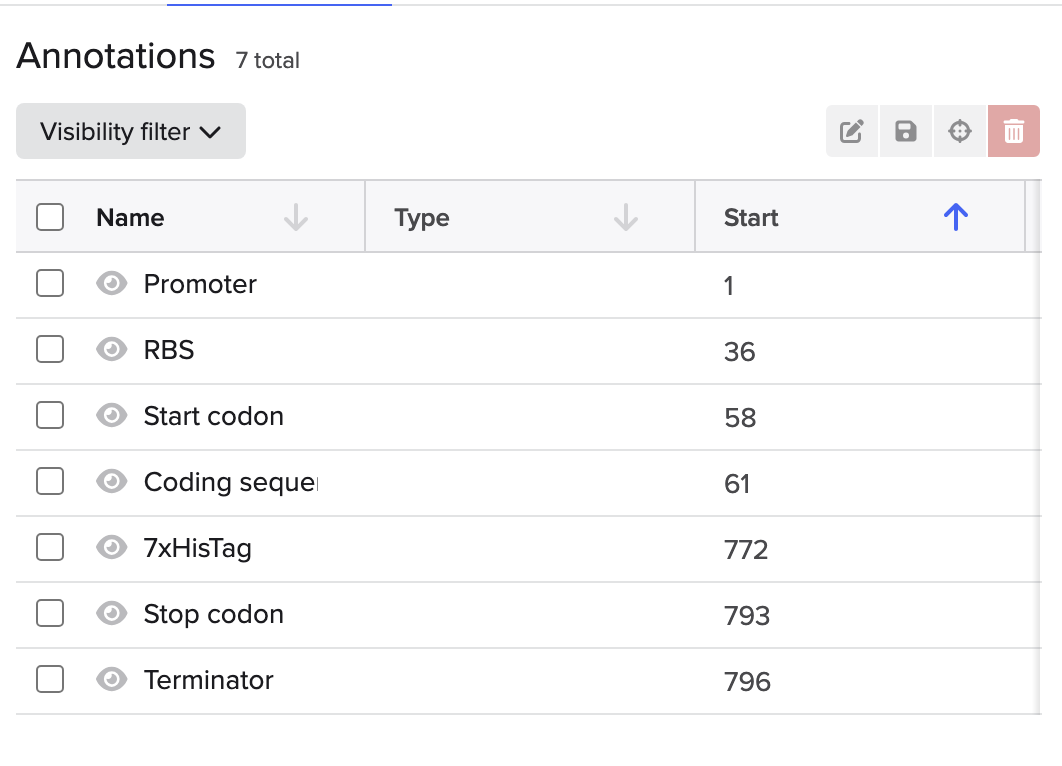

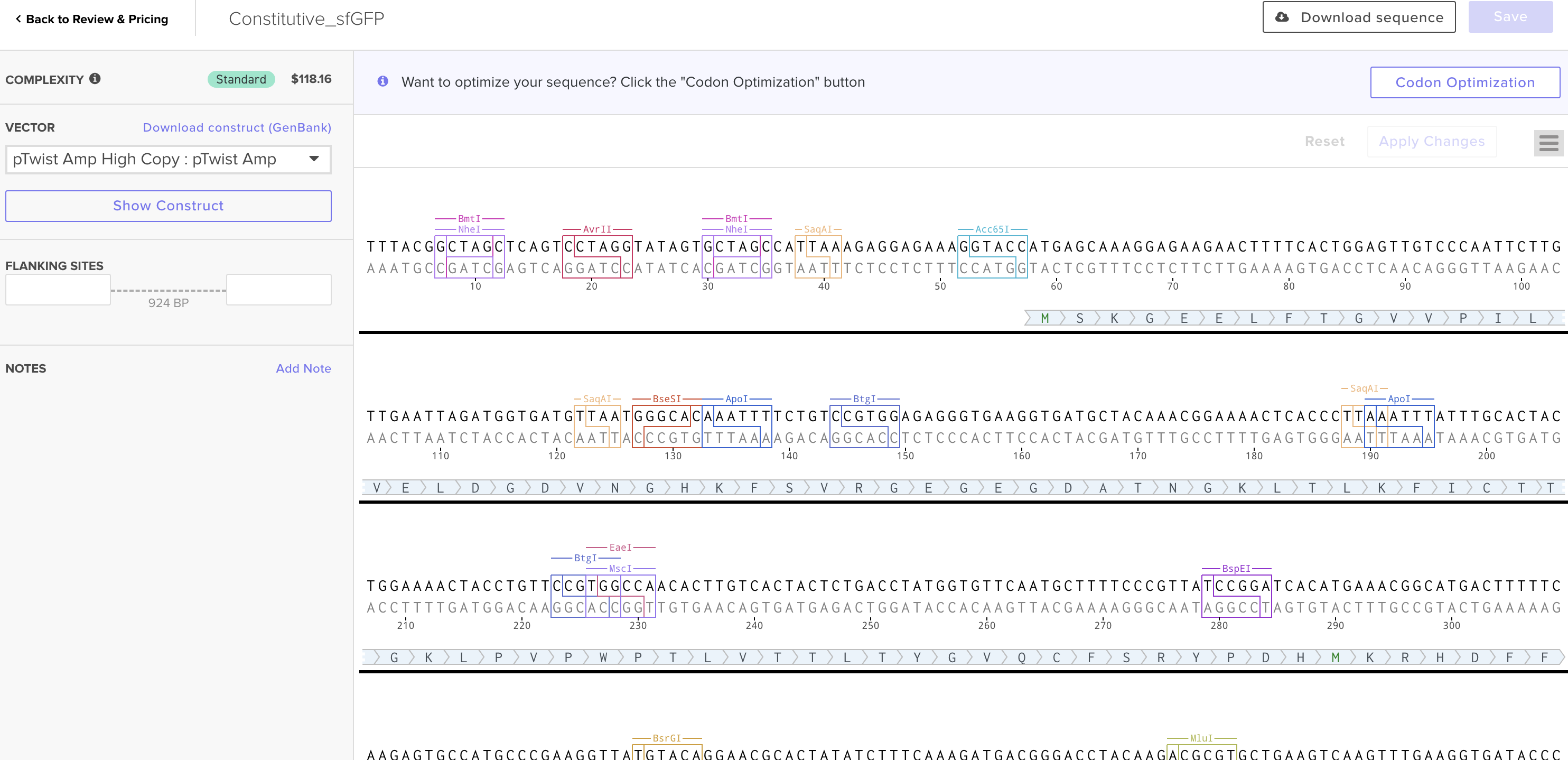

This is the plasmid result :)
DNA read
I would sequence the genomic DNA of Cladosporium sphaerospermum to study it for astrobiology purposes as it is an organisms capable of resisting high levels of radiation. In space or on planetary surfaces like Mars, organisms face chronic ionizing radiation, oxidative stress, desiccation, and nutrient limitation. A fungus that tolerates high radiation is a strong candidate model for understanding which biological features make life resilient in extreme environments.
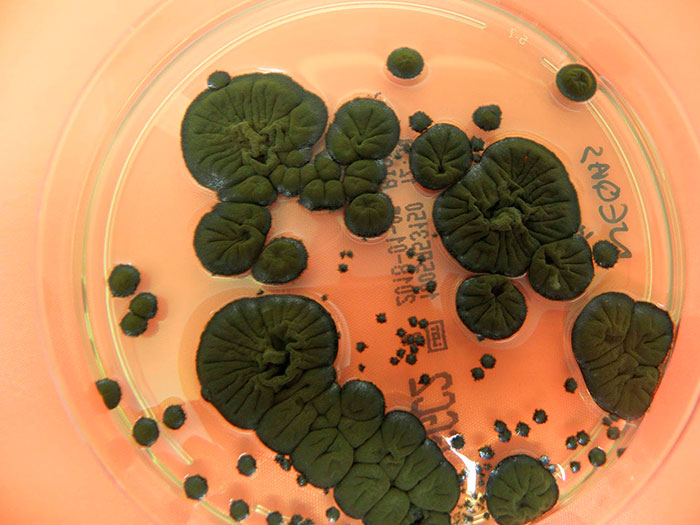


I would sequence it with Oxford Nanopore sequencing (ONT) plus Illumina sequencing-by-synthesis (SBS) as a hybrid strategy. This DNA would let me identify and study genetic systems that can potentially support survival in space-like conditions, such as DNA repair pathways, antioxidant and stress-response networks, and melanin biosynthesis (melanin is known to protect against radiation). I think I would be able to study concepts about long-term survival in other planets, planetary protection, and potential biotech for space habitats like devices or crops.
i. Second-generation sequencing (Illumina SBS)(since it can sequence millions of fragments in parallel using amplification and optical detection during synthesis). ii. Third-generation sequencing (Oxford Nanopore) because it reads single DNA molecules in real time without requiring amplification and can generate very long reads.
This method of sequencing is due that fungal genomes include repeats, variation, and complex regions where long reads are an important part of the genome stability. ONT gives long reads that would improve contiguity and resolve repeats, and Illumina would provide high accuracy to correct residual errors.
–Input: high-quality genomic DNA extracted from fungal cells.
Grow fungal culture and harvest biomass.
Lyse cells and extract genomic DNA.
Purify DNA and assess quality (purity + fragment length).
For Illumina library prep: fragment DNA to short sizes (~200–500 bp), end-repair/A-tailing, adapter ligation, PCR amplification, QC/quantification.
For Nanopore library prep: keep DNA long (minimal fragmentation), ligate Nanopore adapters, load onto flow cell.
–Essential steps of chosen sequencing technology and how does it decode bases:
A. Second-generation sequencing (Illumina SBS)
DNA fragments bind to a flow cell and are clonally amplified into clusters.
During sequencing cycles, fluorescently labeled nucleotides are added one base at a time.
After each cycle, imaging captures the fluorescence signal from each cluster.
The instrument/software assigns the base (A/T/C/G) based on the detected signal, removes the terminator, and repeats for the next base. Base calling is based on fluorescence signals during DNA synthesis.
B. Oxford Nanopore (Nanopore sequencing)
A single DNA molecule passes through a protein nanopore embedded in a membrane.
As different k-mers move through the pore, they cause characteristic disruptions in ionic current.
This method would measure the current changes over time. Base calling would be done by software that converts the electrical signal trace into the DNA sequence.
–Output of sequencing:
The output would consist of sequencing reads (FASTQ files) containing DNA sequences and quality scores. These reads would be assembled into a complete genome (FASTA file) and annotated to identify genes and pathways involved in radiation resistance.


DNA write
I would synthesize a melanin biosynthesis gene from Cladosporium sphaerospermum (such as a polyketide synthase involved in melanin production), it would be codon optimized for expression in a microbial host like Saccharomyces cerevisiae since it is a widely studied model.

Since melanin has been associated with radiation tolerance in fungi found in high-radiation environments. Engineering microbes to produce melanin could help us study biological radiation protection and explore potential applications for space habitats or planetary environments like Mars.
The construct would include: a promoter, ribosome binding site, the codon-optimized melanin gene and a terminator sequence
I would use chip-based oligonucleotide synthesis followed by enzymatic gene assembly, such as the silicon-based platform described in class (Twist Bioscience). This would allow: parallel production of multiple constructs and it would be cost-effective.
–Essential steps:
Design and codon-optimize the gene sequence.
Chemically synthesize short oligonucleotides on a chip.
Amplify the oligo pool.
Enzymatically assemble the full-length gene.
Clone into a plasmid vector.
Sequence-verify the final construct.
–Limitations:
Error rate: Chemical synthesis can introduce mutations, requiring sequence verification.
Length limitations: Very long or repetitive sequences are more difficult to synthesize.
Complexity: High GC content or secondary structure can reduce synthesis efficiency.
DNA Edit

I would edit the genome of Saccharomyces cerevisiae to enhance its tolerance to ionizing radiation and oxidative stress. Since fungi such as Cladosporium sphaerospermum are known to survive in high-radiation environments, I would attempt to transfer or optimize specific protective mechanisms in yeast. (Introducing melanin biosynthesis genes (polyketide synthase)). This would allow us to experimentally test how specific genetic changes contribute to radiation resilience which as discussed previously has implications for astrobiology, planetary protection, and biotechnology applications in space habitats. Apart from astrobiological applications, this could also be applied to conservation biology or restoration efforts, such as modifying species for increased climate resilience.
For yeast genome editing I would use CRISPR–Cas9 technology.
–Essential steps
Design a guide RNA (gRNA) complementary to the target genomic sequence of the yeast
Cas9 nuclease is directed by the gRNA to the target locus.
Cas9 introduces a double-strand break.
The cell repairs the break via: Homology-directed repair (HDR), with a donor template containing homologous sequences.
–Inputs: Designed guide RNA sequence, Cas9 protein, donor DNA template with homology arms, yeast cells, selectable marker, PCR primers for verification
–Design
Identify safe genomic integration site
Design guide RNA and donor DNA construct with flanking homology arms.
Clone into plasmid or assemble as linear repair template.
Transform yeast and select edited clones.
Validate edits via PCR and sequencing.
–Limitations
Variable HDR efficiency
Potential genomic instability if inserting large constructs
Fitness cost of engineered traits

For this homework I used gemini AI within colab to understand how the process worked as well as to achieve the design I had done in the Donovan simulation website. Once I was using colab, the design changed a lot since I could instruct my design better, but I also realized that making a planet desing was much more challenging than I initially thought. While doing this code I ran into some errors due cross contamination, which gemini helped me in localizing where the code did not indicate the robot to drop the tip, it was very helpful.
Idea at the simulation:
Python Script for Opentrons Artwork
Post lab questions
John A Bryant, Mason Kellinger, Cameron Longmire, Ryan Miller, R Clay Wright, AssemblyTron: flexible automation of DNA assembly with Opentrons OT-2 lab robots, Synthetic Biology, Volume 8, Issue 1, 2023, ysac032, https://doi.org/10.1093/synbio/ysac032
This paper introduces AssemblyTron, a software system that helps automate DNA assembly using Opentrons OT-2 to improve the speed, accuracy, and accessibility of synthetic biology workflows. In synthetic biology, building DNA constructs is usually done by hand, which can be slow and prone to mistakes. This paper states that AssemblyTron solves this by taking DNA designs created on a computer and turning them into step-by-step instructions that a robot can follow to perform tasks like PCR, fragment mixing, and plasmid assembly. The authors demonstrate that this system can successfully assemble multi-fragment plasmids, perform site-directed mutagenesis, and achieve transformation efficiencies comparable to manual methods while reducing training time, human error, and costs. (I think that the application of opentrons in research will allow biotechnology and personalized medicine to advance exponentially)
Phase 1: Final Project ideas