Week 2 HW: DNA Read, Write, & Edit
Part 1: Benchling & In-silico Gel Art
- Created a Benchling account and initialized a project for Week 2 focused on molecular cloning and restriction digest simulation.
- Import Lambda DNA from NCBI
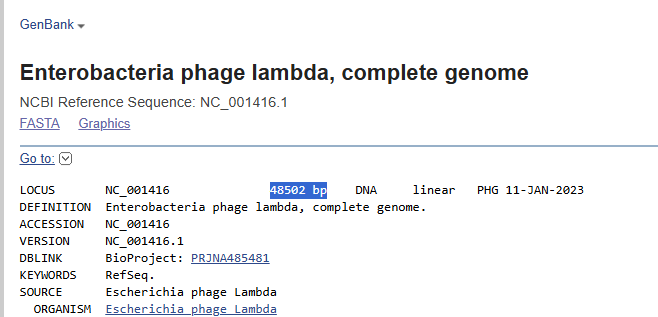
The NEB product specifies Lambda DNA derived from the cI857ind1 Sam7 strain. While this represents a specific laboratory variant, the full genomic backbone corresponds to the canonical 48,502 bp lambda phage genome (NC_001416).
For restriction digest simulations, the reference genome is appropriate, as restriction sites are conserved in the standard laboratory strain.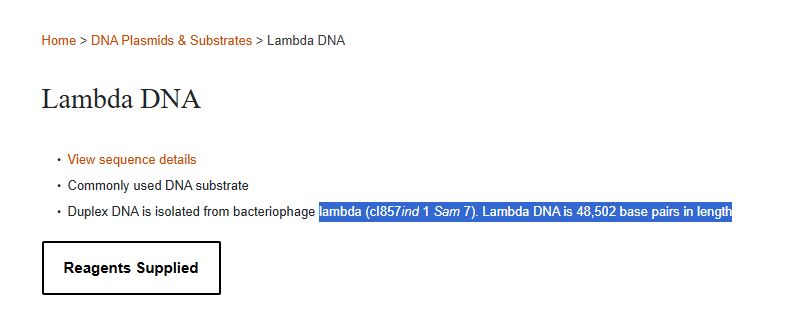
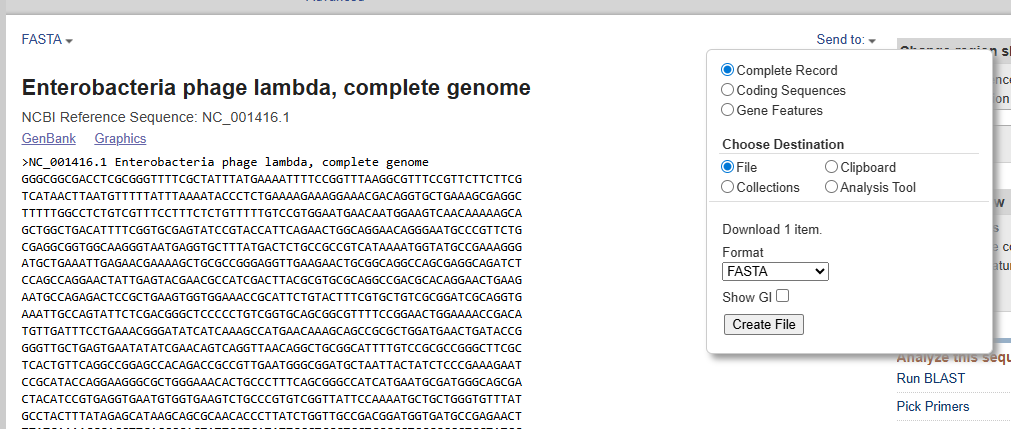
- Upload FASTA to Benchling
Enable translational progress in biomarker-integrated medical devices without imposing governance frameworks that unnecessarily slow legitimate research or clinical adoption.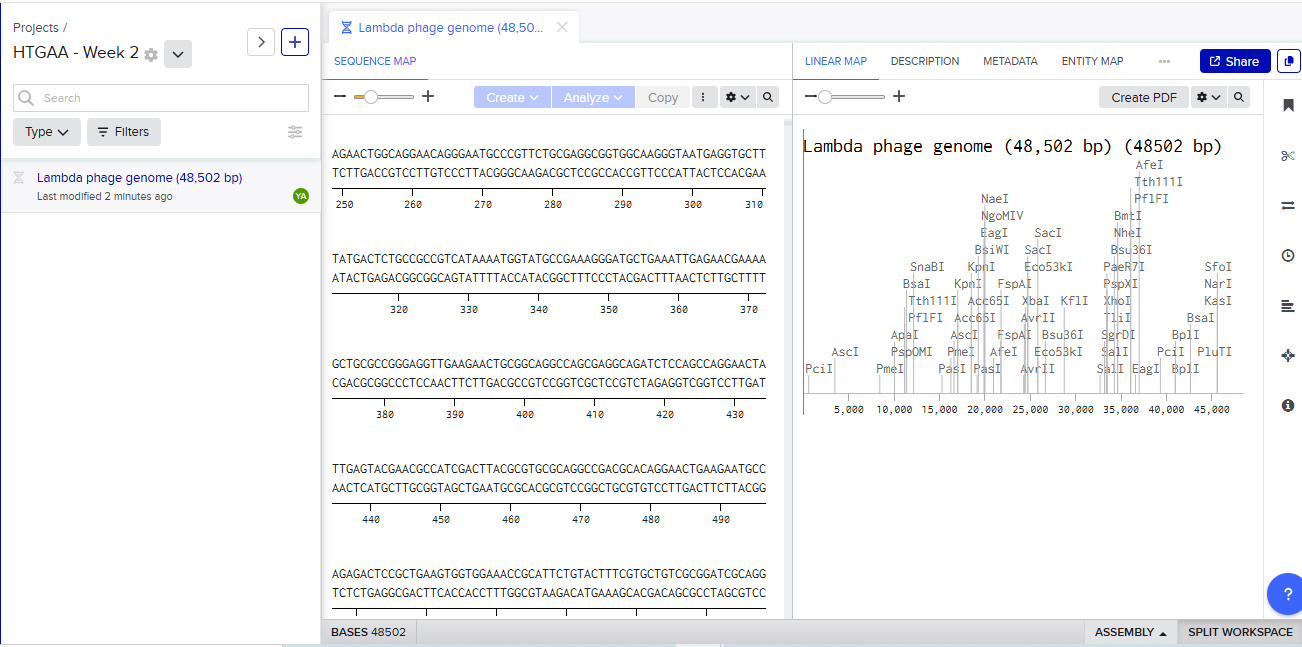
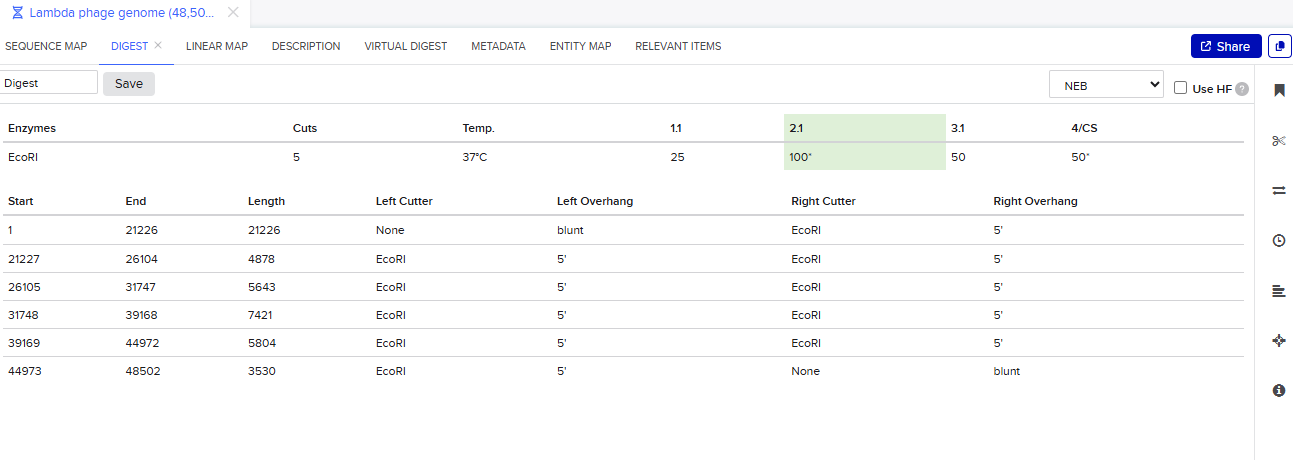
- Simulate Restriction Enzyme Digestion with the following Enzymes
Enable translational progress in biomarker-integrated medical devices without imposing governance frameworks that unnecessarily slow legitimate research or clinical adoption.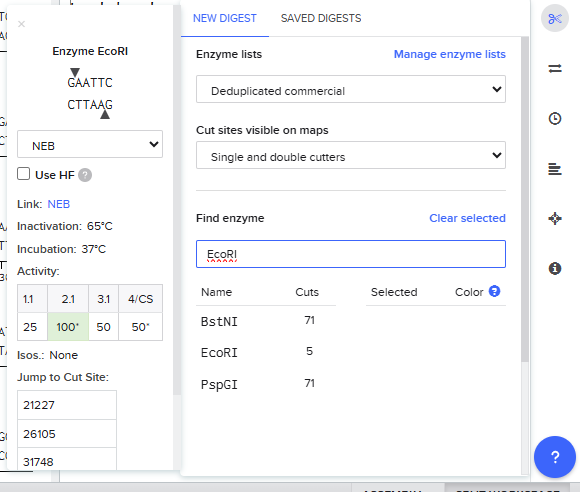
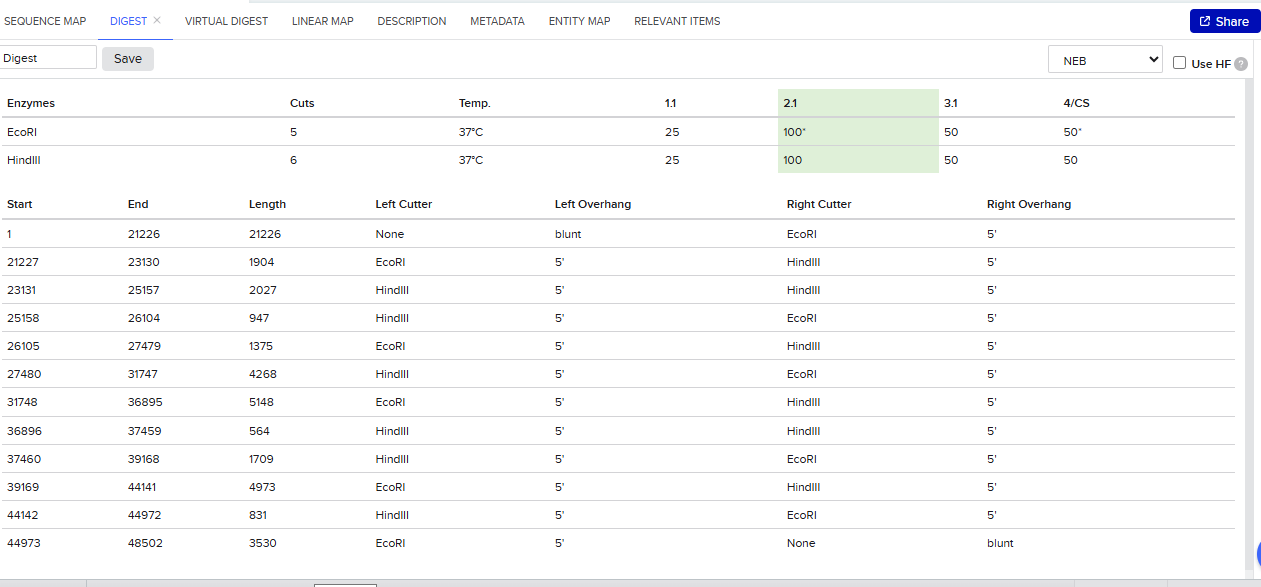
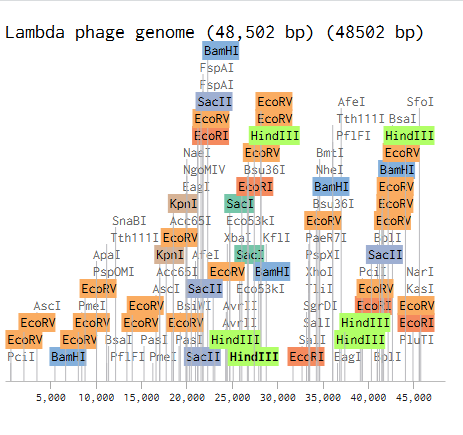
- Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks.
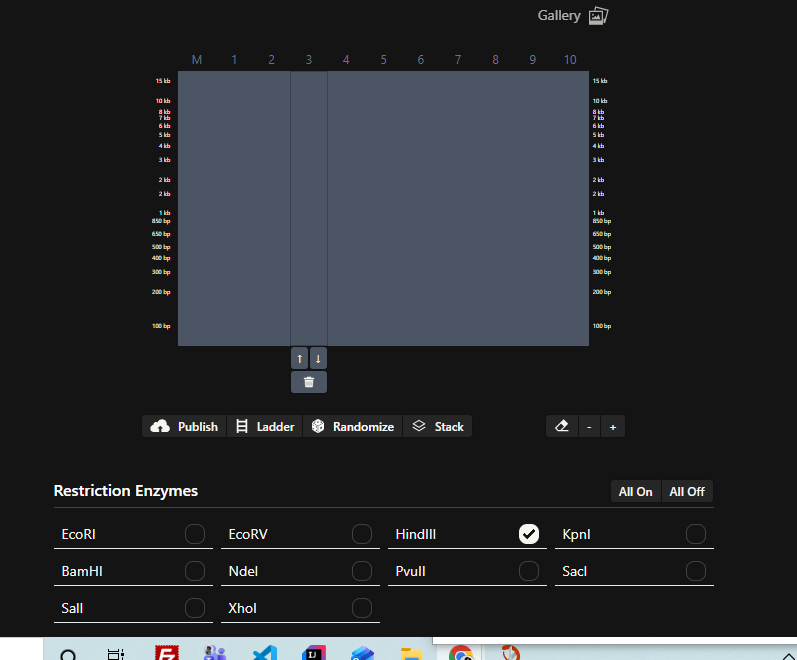
Part 3: DNA Design Challenge
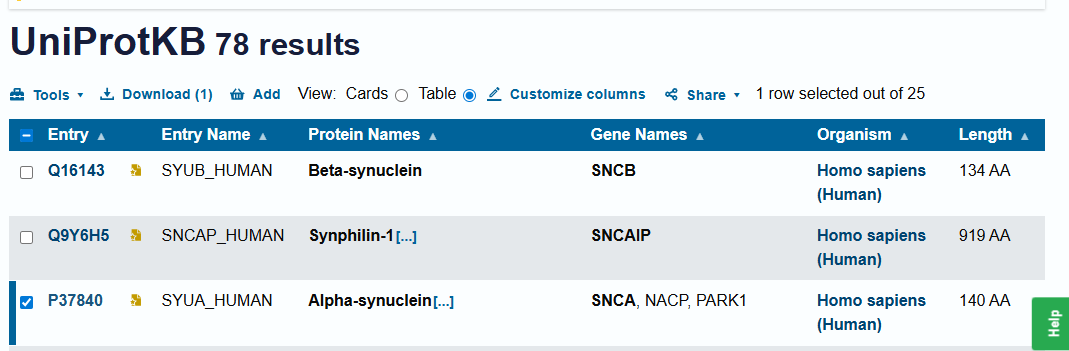
- 3.1 Choose Your Protein: Alpha-Synuclein (SNCA)
I chose human alpha-synuclein (SNCA) because of its central role in neurodegenerative diseases, particularly a group of disorders known as synucleinopathies.Synucleinopathies are a group of chronic and progressive neurodegenerative diseases characterized by the abnormal accumulation of misfolded alpha-synuclein protein. This misfolded protein aggregates into insoluble inclusions known as Lewy bodies, which are found in neurons and glial cells. The major synucleinopathies include Parkinson’s disease, Dementia with Lewy bodies, and Multiple system atrophy. These disorders typically appear in late adulthood and progressively worsen over time.
Alpha-synuclein is a neuronal protein encoded by the SNCA gene in humans. Under normal physiological conditions, it plays a role in synaptic vesicle regulation and neurotransmitter release. However, in neurodegenerative conditions, the protein undergoes misfolding and aggregation, forming toxic oligomers and fibrils that disrupt cellular homeostasis and ultimately lead to neuronal death.
Because protein misfolding and aggregation are central mechanisms in many neurodegenerative diseases, studying alpha-synuclein at the sequence level provides important insight into disease pathology and potential therapeutic strategies.
`>sp|P37840|SYUA_HUMAN Alpha-synuclein OS=Homo sapiens OX=9606 GN=SNCA PE=1 SV=1MDVFMKGLSKAKEGVVAAAEKTKQGVAEAAGKTKEGVLYVGSKTKEGVVHGVATVAEKTKEQVTNVGGAVVTGVTAVAQKTVEGAGSIAAATGFVKKDQLGKNEEGAPQEGILEDMPVDPDNEAYEMPSEEGYQDYEPEA`<br>Sequence taken from UNIPROT
- 3.2 Reverse Translation: Protein (amino acid sequence) → DNA (nucleotide sequence)
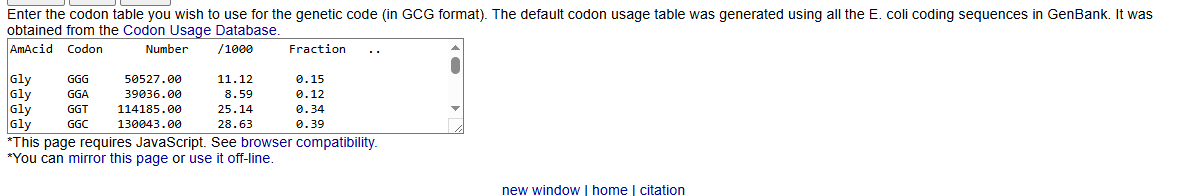
Because of the degeneracy of the genetic code, multiple nucleotide sequences can encode the same amino acid sequence. To obtain a DNA sequence corresponding to human alpha-synuclein, I used the Sequence Manipulation Suite Reverse Translate tool. Using the Escherichia coli codon usage table, the program generated a 420 base pair non-degenerate DNA sequence representing the most likely coding sequence for bacterial expression.The resulting optimized nucleotide sequence is:
atggatgtgtttatgaaaggcctgagcaaagcgaaagaaggcgtggtggcggcggcggaa aaaaccaaacagggcgtggcggaagcggcgggcaaaaccaaagaaggcgtgctgtatgtg ggcagcaaaaccaaagaaggcgtggtgcatggcgtggcgaccgtggcggaaaaaaccaaa gaacaggtgaccaacgtgggcggcgcggtggtgaccggcgtgaccgcggtggcgcagaaa accgtggaaggcgcgggcagcattgcggcggcgaccggctttgtgaaaaaagatcagctg ggcaaaaacgaagaaggcgcgccgcaggaaggcattctggaagatatgccggtggatccg gataacgaagcgtatgaaatgccgagcgaagaaggctatcaggattatgaaccggaagcg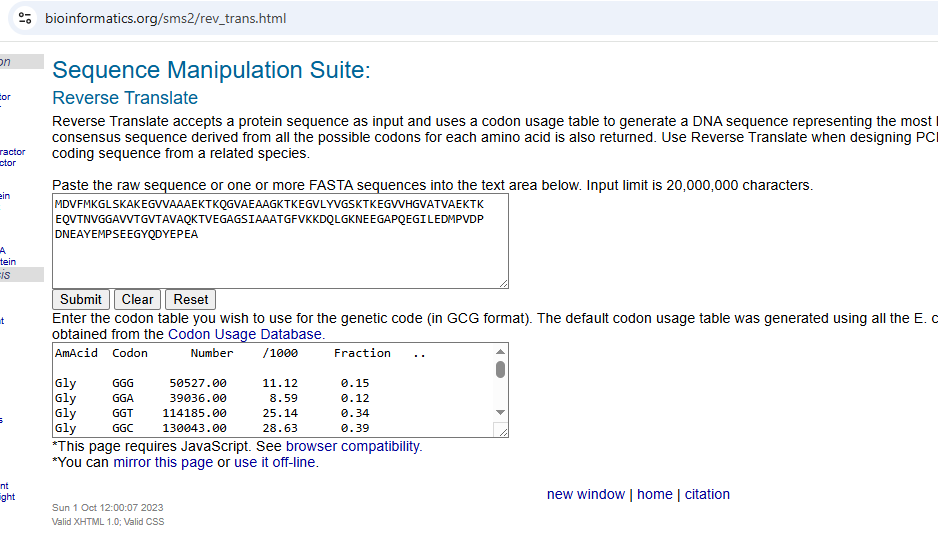
- 3.3 Codon Optimization
Although the genetic code is universal, different organisms exhibit codon usage bias, meaning that certain codons are used more frequently than others. Using rare codons in a host organism can reduce translation efficiency, slow protein synthesis, and negatively impact protein folding.To improve expression efficiency, I optimized the alpha-synuclein coding sequence for Escherichia coli, a commonly used bacterial expression system in molecular biology. Using the Reverse Translate tool with an E. coli codon usage table, the most frequently used codons were selected for each amino acid. This increases the likelihood of efficient transcription and translation when the gene is expressed in bacterial cells.
- 3.4 How Can This DNA Be Used to Produce Alpha-Synuclein?
The optimized alpha-synuclein DNA sequence can be used to produce protein through recombinant expression in a bacterial system. Below is a step-by-step explanation of how the DNA sequence is transcribed and translated into protein.Step 1: Gene Synthesis and Cloning
The codon-optimized alpha-synuclein DNA sequence is first chemically synthesized and inserted into a plasmid vector. The plasmid contains a strong promoter (such as T7), a ribosome binding site, an antibiotic resistance gene for selection, and a transcription terminator. This plasmid serves as a vehicle to introduce and express the gene inside bacterial cells.Step 2: Transformation into Bacterial Cells
The recombinant plasmid is introduced into competent Escherichia coli cells through heat shock or electroporation. The bacteria are then plated on antibiotic-containing media so that only cells that successfully incorporate the plasmid survive.Step 3: Transcription (DNA → mRNA)
Inside the bacterial cell, RNA polymerase binds to the promoter and transcribes the alpha-synuclein DNA sequence into messenger RNA (mRNA). During transcription, thymine (T) in DNA is replaced by uracil (U) in RNA.Step 4: Translation (mRNA → Protein)
The mRNA binds to ribosomes, which read the sequence in sets of three nucleotides (codons). Each codon specifies one amino acid. Transfer RNAs (tRNAs) deliver the corresponding amino acids, which are linked together to form the alpha-synuclein protein. Because the gene was codon-optimized for E. coli, translation efficiency is increased.Step 5: Protein Folding and Purification
After translation, the protein folds into its functional structure. The bacterial cells can then be lysed, and the protein purified using methods such as affinity chromatography, ion exchange chromatography, or size exclusion chromatography.
In summary, the optimized DNA sequence is transcribed into RNA and translated into alpha-synuclein protein, enabling large-scale production for research on neurodegenerative diseases such as Parkinson’s disease. - 3.5 How Does It Work in Nature?
In biological systems, genetic information flows according to the Central Dogma: DNA → RNA → Protein. A single gene can give rise to multiple protein products through mechanisms such as alternative splicing, alternative promoters, and post-transcriptional modifications.The human SNCA gene primarily encodes the 140-amino-acid alpha-synuclein protein. However, alternative splicing can generate shorter isoforms (e.g., 126 or 112 amino acids), demonstrating how one gene can produce multiple protein variants.
Below is the alignment of the designed DNA sequence with its corresponding RNA and translated protein:This alignment confirms that the optimized DNA sequence correctly encodes alpha-synuclein without frame shifts or premature stop codons.DNA
ATG GAT GTG TTT ATG AAA GGC CTG AGC↓ TranscriptionmRNA
AUG GAU GUG UUU AUG AAA GGC CUG AGC↓ TranslationProtein
Met – Asp – Val – Phe – Met – Lys – Gly – Leu – Ser
Part 4: Prepare a Twist DNA Synthesis Order
/* Insert Pictures */Part 5: DNA Read/Write/Edit - First draft
- 5.1 DNA Read
(i) What DNA would you want to sequence and why?
For this project, I would focus on genes commonly mutated in adenocarcinoma endometrial, such as TP53, PTEN, PIK3CA. Sequencing these genes allows identification of mutations that drive tumor progression and informs targeted therapy strategies.Example source: NCBI Gene Database for TP53: https://www.ncbi.nlm.nih.gov/gene/7157
(ii) Sequencing technologies to use and why:
- I would use next-generation sequencing (NGS, second-generation) due to its high throughput and accuracy for detecting somatic mutations.
- Input preparation: DNA extraction from tissue, fragmentation, adapter ligation, PCR amplification.
- Sequencing steps: Cluster generation on flow cell, cyclic reversible termination (base incorporation), imaging, base calling.
- Output: FASTQ files with read sequences, aligned to human reference genome to identify mutations. - 5.2 DNA Write
(i) What DNA would you want to synthesize and why?
To simulate or study therapeutic interventions, I could synthesize DNA fragments of TP53, PTEN, or PIK3CA with specific mutations. These could be used in cell line models to test gene expression, protein production, or biosensor detection in a controlled environment.Example applications:
- Genetic circuits for biosensors embedded in the DIU to detect tumor markers.
- DNA origami structures for controlled drug release or as scaffolds for sensor integration.(ii) Technology to perform DNA synthesis and why:
- I would use oligonucleotide synthesis and assembly methods (Twist Bioscience, IDT), which are precise and scalable.
- Essential steps: Phosphoramidite synthesis, purification, assembly into larger constructs.
- Limitations: Sequence length constraints (usually <1–2 kb per construct), cost for large-scale synthesis, and potential errors in repetitive sequences. - 5.3 DNA Edit
(i) What DNA would you want to edit and why?
I would target mutated TP53 or PTEN alleles in endometrial cells. Editing these genes could theoretically restore normal tumor suppressor function, providing a model for localized gene therapy integrated with DIU-mediated monitoring.
For Example: The Company Colossal Biosciences demonstrates genome engineering in animals; similarly, precise editing tools can model human therapeutic strategies.(ii) Technology to perform DNA edits and why:
- I would use CRISPR/Cas9 or base editors for targeted nucleotide correction.
- Essential steps: Design guide RNA complementary to the mutation, deliver Cas9 and guide RNA into cells, allow DNA repair mechanisms to introduce the correct sequence.
- Input preparation: DNA template or plasmid with Cas9 and guide RNA, target cells (e.g., endometrial cell line), transfection reagents.
- Limitations: Off-target effects, variable editing efficiency, delivery challenges in vivo.
📚 References
- NCBI. (n.d.). Lambda phage genome, complete sequence. (link)
- Benchling. (n.d.). Lambda phage genome project – Week 2. (link)
- UniProt. (n.d.). Alpha-synuclein (SNCA) – Homo sapiens. (link)
- Bioinformatics.org. (n.d.). Sequence Manipulation Suite – Reverse Translate. (link)
- NCBI Gene. (n.d.). TP53 tumor protein p53 [Homo sapiens]. (link)
- R.C. Donovan. (n.d.). Gel art. (link)
- HTGAA 2026a. (2026). Week 2 Lab: Benchling & In-silico Gel Art. Course materials. (link)
- OpenAI. (2026). ChatGPT (GPT-5.2) [Large language model]. Used for drafting support and conceptual clarification. (link)
- This topic is related to my thesis project: Diseño Computacional de un DIU Antitumoral con Liberación Controlada y Monitoreo para Terapia Localizada en Adenocarcinoma Endometrial.