week-06-hw-genetic-circuits-part-i
Homework — DUE BY START OF MAR 17 LECTURE
Assignment: DNA Assembly
Answer these questions about the protocol in this week’s lab:
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity master mix contains:
- Phusion DNA polymerase: An enzyme that synthesizes new DNA
- dNTPs (deoxynucleotide triphotes): The blocks that the polymerase incorporates to build the new DNA strand.
- Reaction buffer: Provides optimal pH and salt conditions for the polymerase.
- Stabilizers and additives: Help maintain enzyme activity during temperature cycling.
2. What are some factors that determine primer annealing temperature during PCR?
The annealing temperature (Tm) is determinaned mainly by:
1. Base composition: G-C pairs form three hydrogen bonds, a higher GC content increases Tm
2. Presence of mismatches: Tm decrease, if the primer is not perfectly complementary
3. Primer length: Longer primers tend to have higher Tm, but the optimal range is usually 18-22 bp.
4. Salt concentration: Especially Mg 2+ and monovalent cations (Na+, K+) in the buffer, higher concentrations stabilize binding and raise Tm.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
| Aspect | PCR | Restriction enzyme digestion |
|---|---|---|
| Protocol | Requires specific primers, master mix and thermocycler | Uses restrcition enzymes that cut DNA at specific sites |
| Flexibility | Generate fragments of virtually any sequence with appropriate primers. | Less flexible for generating custom fragments without suitable sites. |
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
The several conditions must be met:
- Purity: After PCR or digestion, DNA is purified to remove salts, enzymes, and residual primers.
- Proper concentration: DNA concentration must be quantified to calculate the correct molar ratio.
- Integrity: Verify by gel electrophoresis that fragments are not degraded.
5. How does the plasmid DNA enter the E. coli cells during transformation?
Plasmid DNA enters E. coli cells during transformation with artificially induced process:
Preparation of competent cells: bacteria are treated with calcium chloride
Heat shock : Exposing to 42 °C for a short time.
Passive diffusion: Plasmid DNA enters by diffusion through the pores.
6. Describe another assembly method in detail (such as Golden Gate Assembly)
1. Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
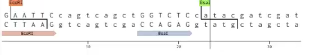
Golden Gate Assembly is a molecular cloning method that uses type IIS restriction enzymes, such as BsaI, which recognize non-palindromic DNA sequences but cut outside of their recognition sites. This unique property allows researchers to design custom overhangs that determine the precise order in which DNA fragments are assembled. By placing these recognition sites at the ends of multiple DNA fragments, several pieces can be joined together in a single, one-pot restriction-ligation reaction. During the process, the restriction enzyme cuts the DNA while a ligase simultaneously joins the fragments, enabling seamless and scarless assembly. This method is highly efficient and reduces the need for multiple cloning steps compared to traditional techniques. Golden Gate Assembly is widely used for constructing gene libraries, assembling synthetic biological circuits, and combining multiple genetic elements into a destination vector. However, one limitation is that internal restriction sites matching the enzyme used must be removed or avoided to prevent unwanted cutting
2. Model this assembly method with Benchling or Asimov Kernel!

Assignment: Asimov Kernel
Create a Repository for your work
Create a blank Notebook entry to document the homework and save it to that Repository
Explore the devices in the Bacterial Demos Repo to understand how the parts work together by running the Simulator on various examples, following the instructions for the simulator found in the “Info” panel (click the “i” icon on the right to open the Info panel)
Create a blank Construct and save it to your Repository
Recreate the Repressilator in that empty Construct by using parts from the Characterized Bacterial Parts repository
Search the parts using the Search function in the right menu
Drag and drop the parts into the Construct
Confirm it works as expected by running the Simulator (“play” button) and compare your results with the Repressilator Construct found in the Bacterial Demos repository
Document all of this work in your Notebook entry - you can copy the glyph image and the simulator graphs, and paste them into your Notebook
Build three of your own Constructs using the parts in the Characterized Bacterials Parts Repo
Explain in the Notebook Entry how you think each of the Constructs should function
Run the simulator and share your results in the Notebook Entry
If the results don’t match your expectations, speculate on why and see if you can adjust the simulator settings to get the expected outcome
REFERENCES:
- PromptS ChatGPT
- Explain Golden Gate Assembly in detail, including how type IIS restriction enzymes function, why they cut outside their recognition sites, and how this enables seamless DNA assembly. Provide a step-by-step explanation and include advantages over traditional cloning methods.
- Describe how Golden Gate Assembly can be used to construct synthetic genetic circuits. Include an example with multiple DNA fragments (e.g., promoter, coding sequence, terminator) and explain how their order is controlled during assembly.