Week 6 HW: Genetic Circuits Part I

Image credit: Phase Genomics
Part 1. DNA Assembly
Q1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Components in Phusion High-Fidelity PCR Master Mix:
- DNA polymerase: a high-fidelity enzyme that synthesizes new DNA strands based on the template sequence.
- dNTPs: the molecular building blocks used to make the new DNA strands.
- Buffer: provides the proper chemical environment, such as pH and ionic strength, to maintain polymerase activity and stability.
- MgCl₂ (Mg²⁺): an essential cofactor for DNA polymerase and is required for its catalytic activity.
Q2. What are some factors that determine primer annealing temperature during PCR?
In PCR, primer annealing temperature (Ta) is primarily determined by the primer melting temperature (Tm), usually 2-5°C below the lowest Tm of the pair.
Key factors influencing this include:
- primer length
- guanine-cytosine (GC) content
- base sequence (nearest-neighbor interactions)
- salt concentration (e.g., Na⁺)
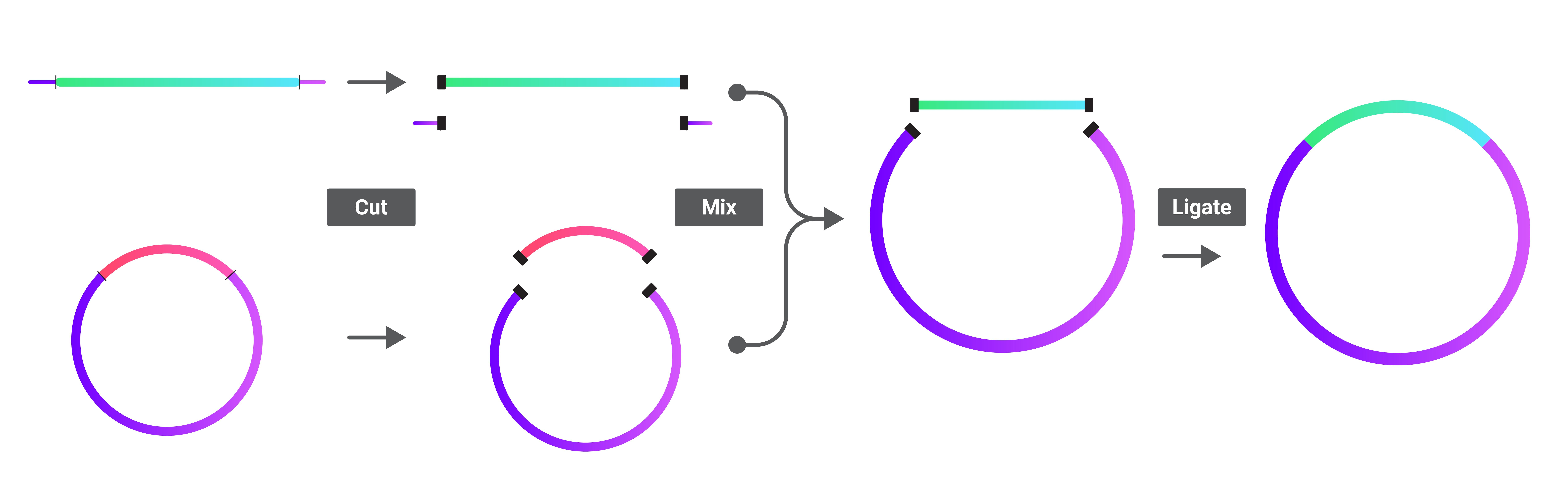
Q3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR amplifies a specific DNA sequence from a template using heat-stable polymerase and primers, while restriction enzymes (endonucleases) cut double-stranded DNA at specific recognition sites. PCR is ideal for generating large quantities of DNA from small amounts of starting material, while restriction digests are often used for cutting out known fragments from existing plasmids or genomic DNA.
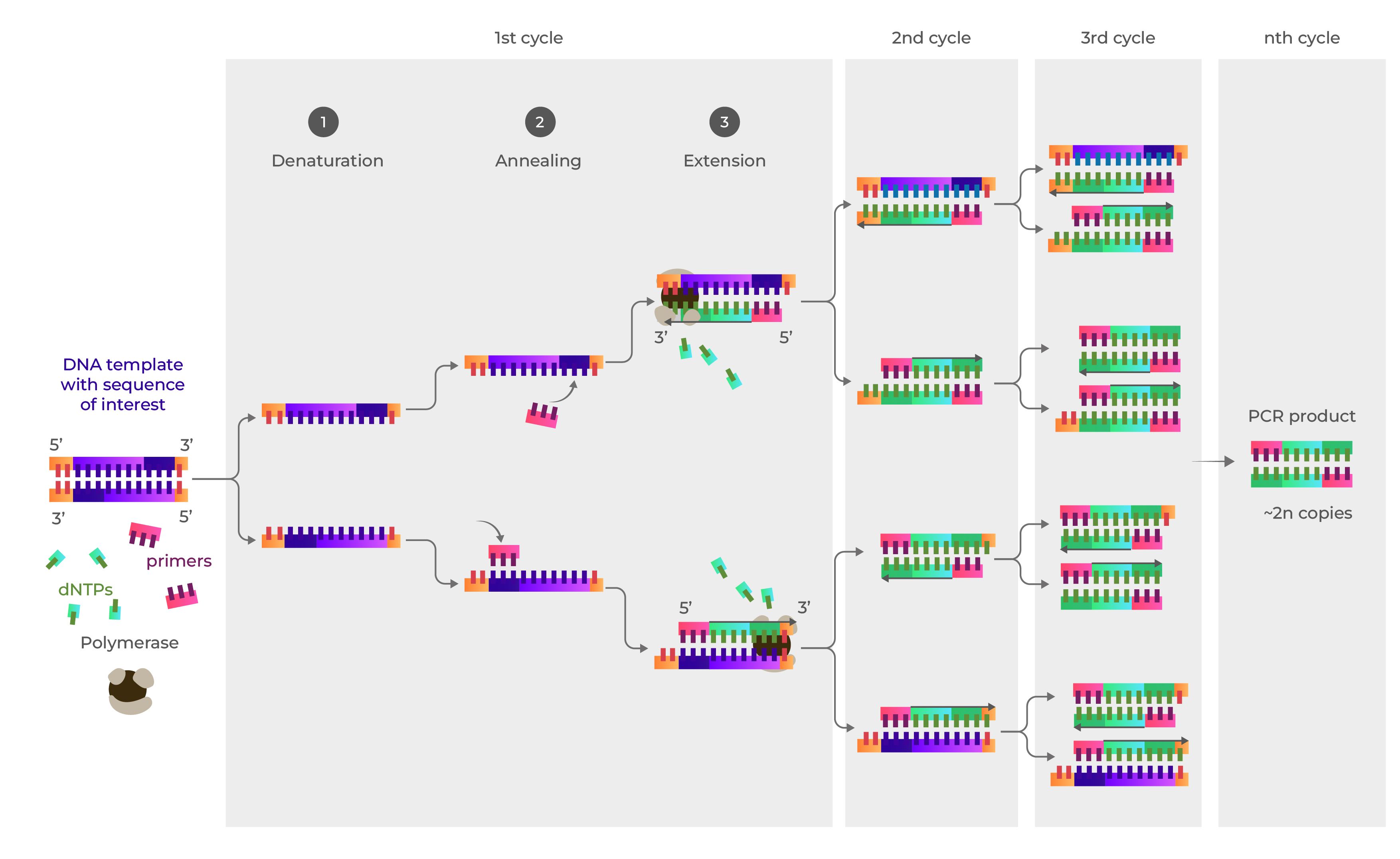
 Polymerase Chain Reaction (PCR) Overview
Polymerase Chain Reaction (PCR) Overview
Q4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
- The vector is properly linearized.
- Each digested or PCR-amplified fragment has the correct 20–40 bp overlapping homology with the adjacent fragment.
- All DNA fragments are clean and of the expected size.
- After assembly, confirm the final construct by sequencing.
Q5. How does the plasmid DNA enter the E. coli cells during transformation?
Plasmid DNA enters E. coli when the cells are shocked by heat shock or electrical shock, which temporarily opens the cell membrane and allows the plasmid to enter by diffusion.
Q6. Describe another assembly method in detail (such as Golden Gate Assembly)
Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Golden Gate Assembly is a one-step cloning method that uses a Type IIS restriction enzyme and DNA ligase in the same reaction tube. Unlike standard restriction enzymes, Type IIS enzymes cut outside of their recognition sites, which allows researchers to design custom overhangs that determine the order of assembly. DNA fragments and the destination vector are designed with matching overhangs, so the pieces ligate together in a predefined sequence. Because the recognition sites are removed during the reaction, the correctly assembled product is typically scarless and is not re-cut by the enzyme. This also helps reduce the background from empty vector, since unassembled or re-ligated vector can be cut again during the cycling reaction. Golden Gate Assembly is especially useful for efficient multi-fragment and modular DNA assembly.
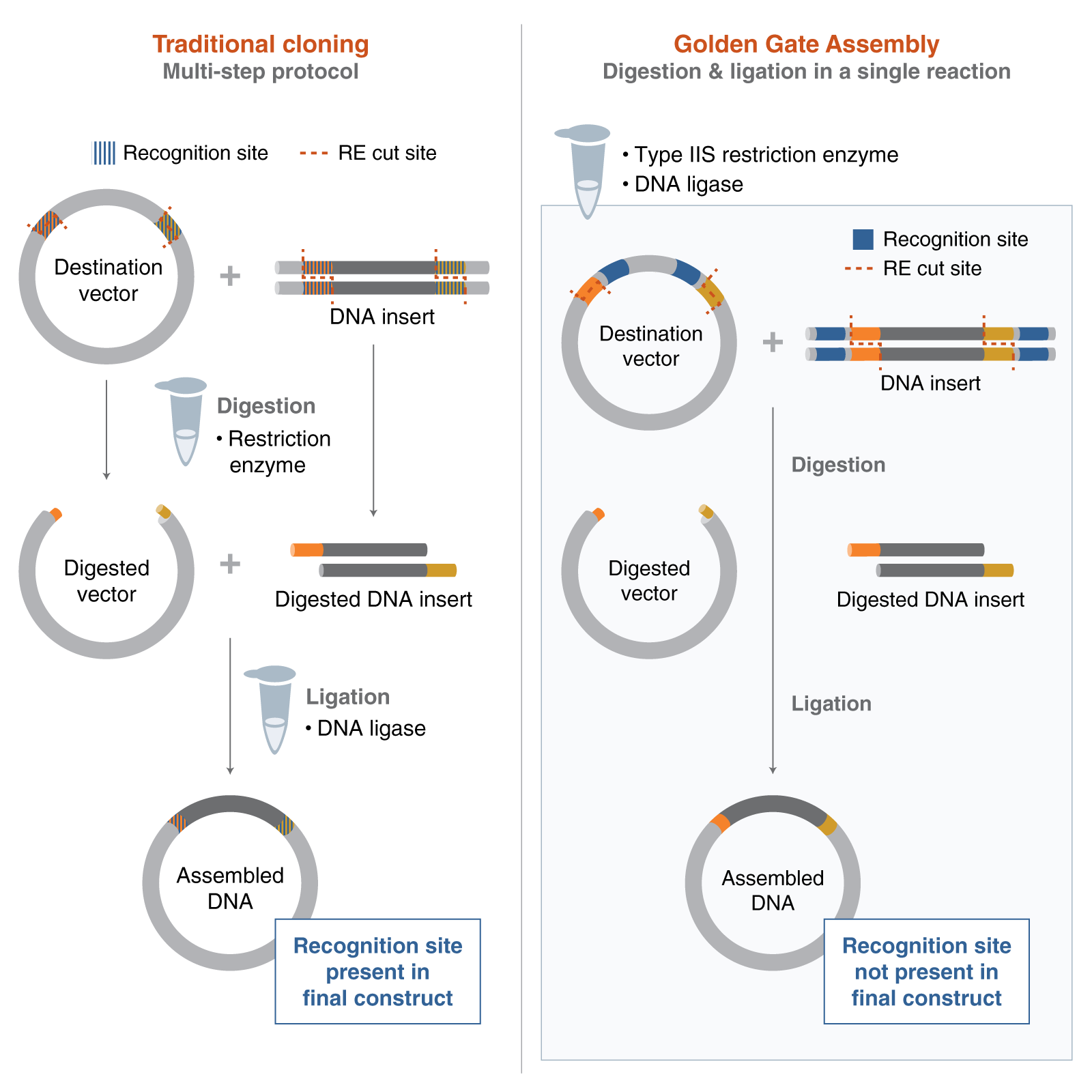
 Golden Gate Assembly is a Restriction Enzyme Cloning Method. Golden Gate Assembly vs. Traditional Cloning.
Golden Gate Assembly is a Restriction Enzyme Cloning Method. Golden Gate Assembly vs. Traditional Cloning.
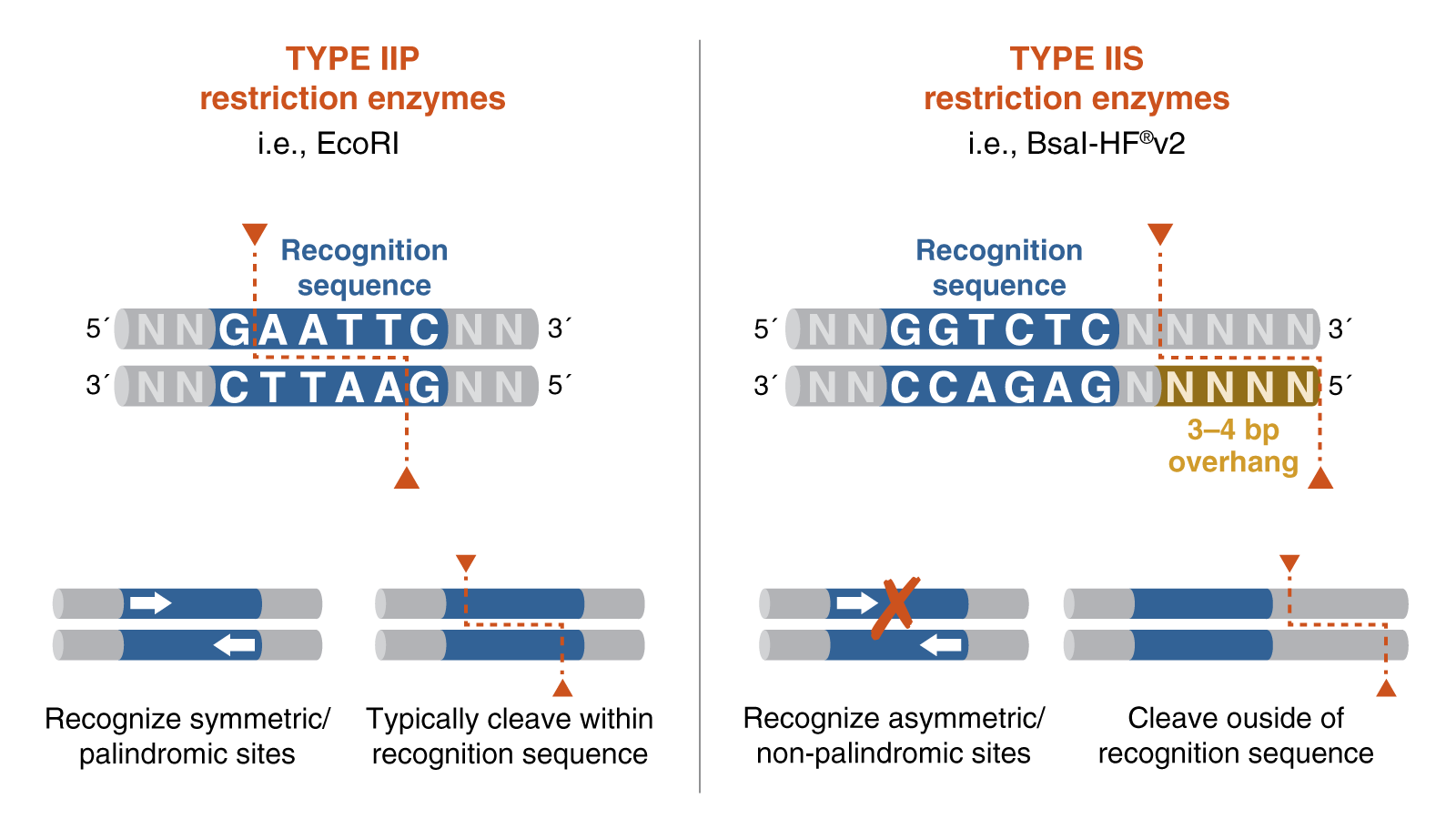
 Type IIS Restriction Enzymes Enable the Unique Features of Golden Gate Assembly Type IIP vs. Type IIS Restriction Enzymes
Type IIS Restriction Enzymes Enable the Unique Features of Golden Gate Assembly Type IIP vs. Type IIS Restriction Enzymes
- Model this assembly method with Benchling or Asimov Kernel!
Part 2. Asimov Kernel
Methodology & Logic Implementation
In this practice, I manually selected individual components from the Characterized Bacterial Parts repository to construct a functional Repressilator. The core logic was implemented by cross-linking three inhibitory units in a cyclic negative feedback loop: pTet → LambdaCI, pLambdaCI → LacI, and pLacI → TetR.
My primary focus was to explore how specific biological factors impact the simulation outcome, including:"
RNAP & Ribosome Flux: Assessing the transcriptional and translational efficiency at each junction.
RNA & Protein Concentrations: Monitoring the temporal dynamics of gene expression.
Component Strength: Evaluating if the specific Characterized Parts selected (such as the B3 RBS) are kinetically compatible with the desired oscillatory behavior.
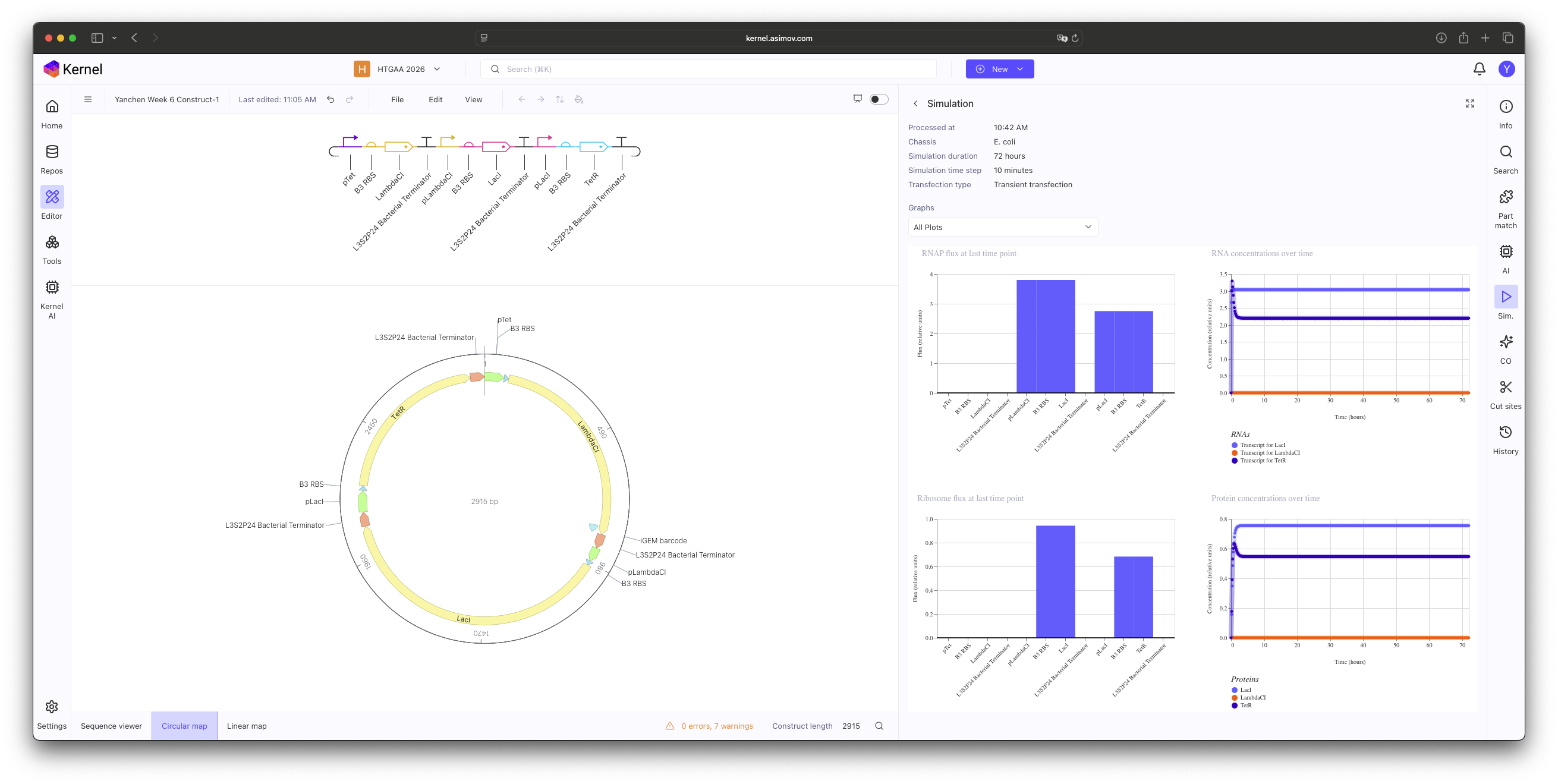
 Construct 1: Promoter mismatch.
Construct 1: Promoter mismatch.
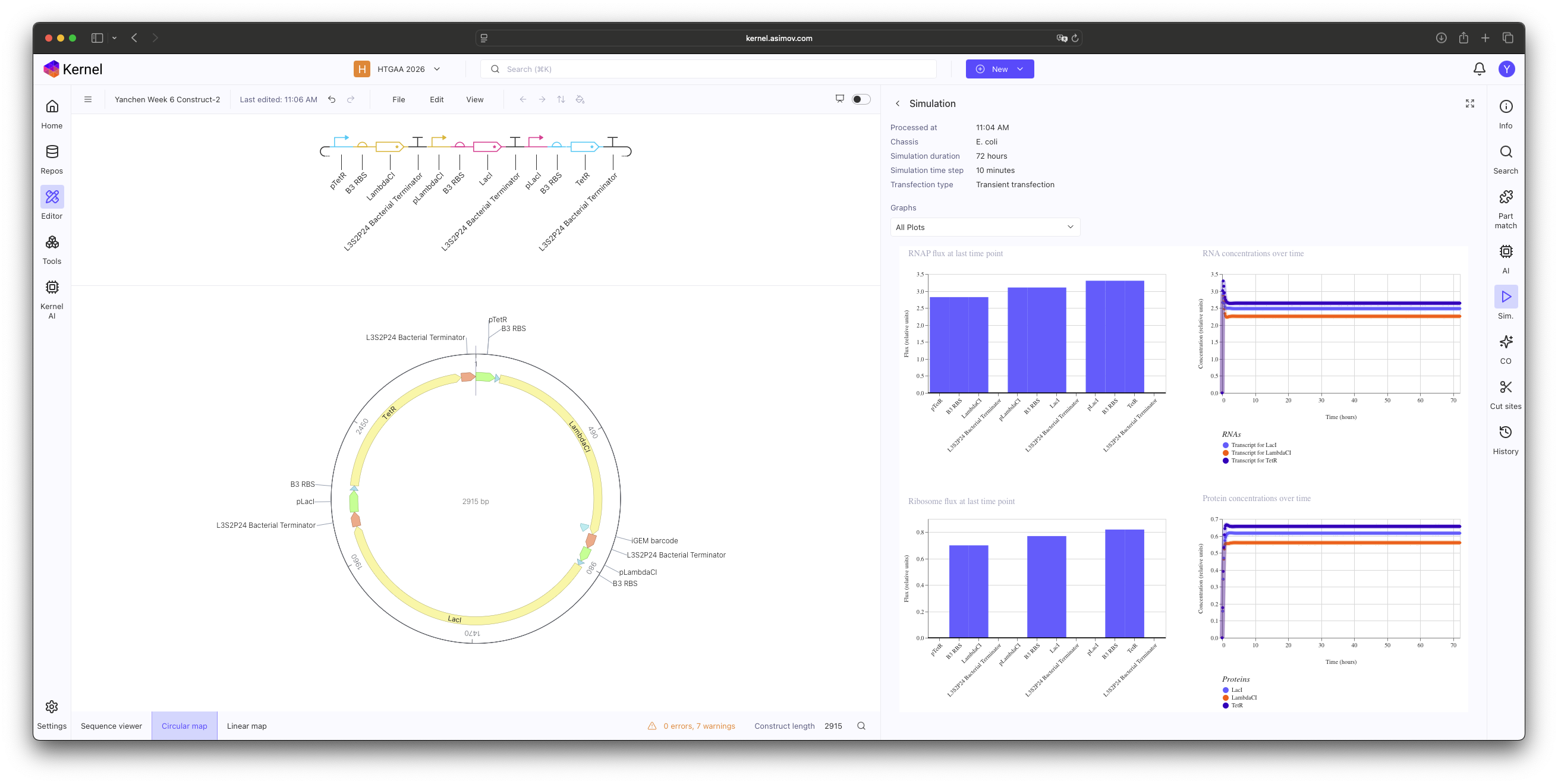
 Construct 2: Ribosome binding site (B3 RBS). Initial simulations showed no oscillations; this is likely due to the B3 RBS being too potent, which leads to an excessive accumulation of repressor proteins that the system cannot degrade quickly enough to maintain a periodic rhythm.
Construct 2: Ribosome binding site (B3 RBS). Initial simulations showed no oscillations; this is likely due to the B3 RBS being too potent, which leads to an excessive accumulation of repressor proteins that the system cannot degrade quickly enough to maintain a periodic rhythm.
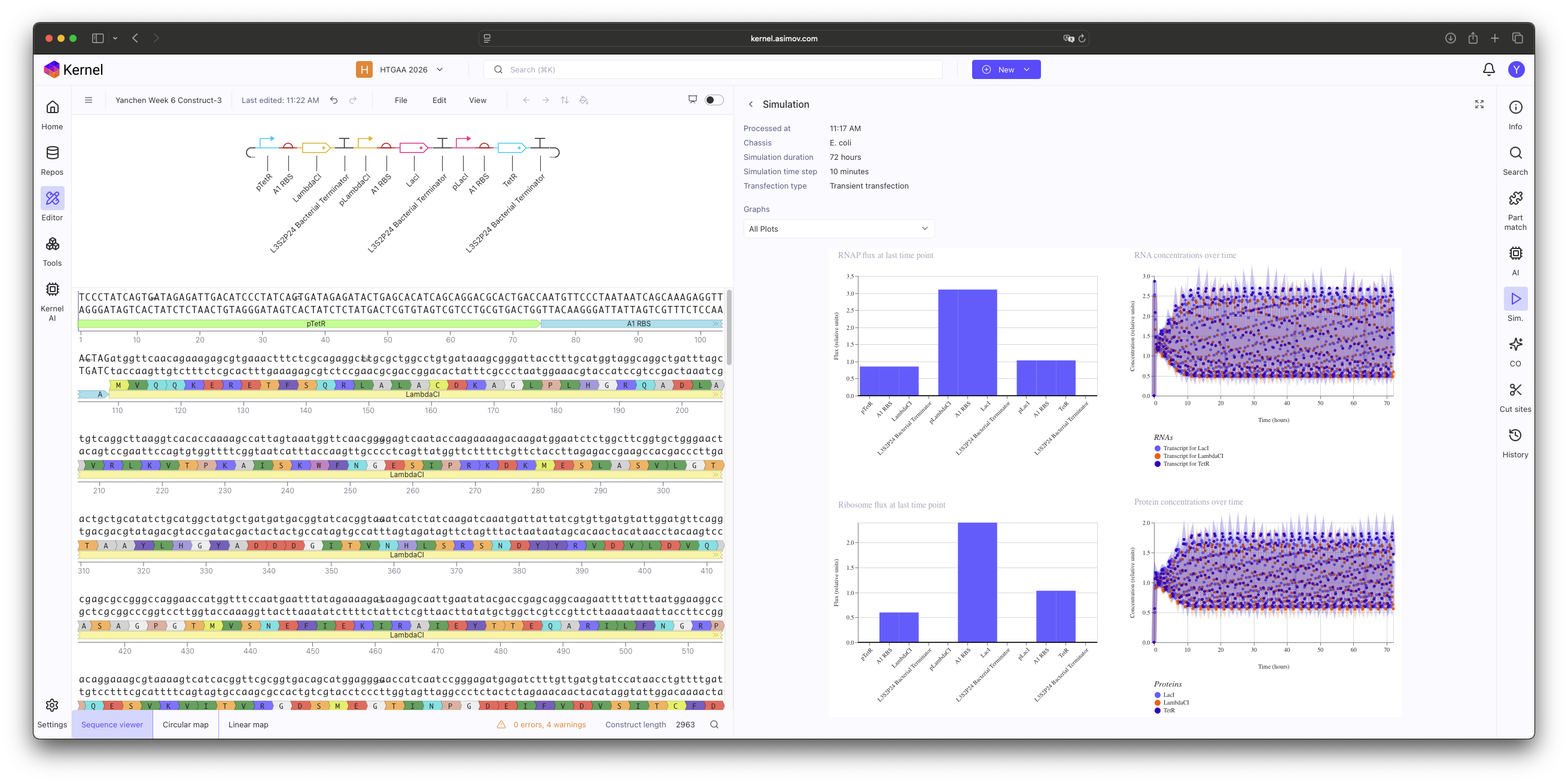
 Construct 3: Change Ribosome binding site (B3 RBS) to A1 RBS.
Construct 3: Change Ribosome binding site (B3 RBS) to A1 RBS.
Results & Analysis
The simulation initially resulted in stable, non-oscillatory steady states. Through this exercise, I identified that using standard Characterized Parts requires more than just correct logic; it requires kinetic tuning. Specifically, the B3 RBS proved to be too potent, leading to a protein synthesis rate that overwhelmed the system’s degradation capacity. This ’locked’ the Protein and RNA concentrations at a high-level equilibrium, preventing the periodic ‘dip’ necessary for oscillation.