Week 7 HW: Genetic Circuits Part II
Part 1. Intracellular Artificial Neural Networks
Q1. Advantages of IANNs over Traditional Boolean Genetic Circuits
A traditional genetic circuit works like a panel of on‑off light switches. Each gene is either fully expressed or completely silent, and the circuit’s output is a strict Boolean function of those binary inputs. An IANN, by contrast, behaves more like a set of dimmer switches connected through a mixing board. Each input can take any value within a continuous range, the connections have adjustable weights, and the final output is a smooth, graded signal instead of a hard 0 or 1.
This difference brings several benefits. Because IANNs are built from sequestrons that process signals in the analog domain, they can represent and compute with concentrations across a wide dynamic range. Boolean circuits squeeze all that richness into just two bins, but IANNs preserve it. A large enough IANN can in principle approximate any input‑output function, which is the biological version of the universal approximation theorem from machine learning. IANNs are also compact and scalable. Instead of layering many different logic gates, each with its own set of genetic parts, they use a single repeatable building block called a sequestron. The weights are set simply by adjusting DNA concentrations, so adding complexity means adding more copies of the same module rather than inventing new gate designs. Tuning the weights is like turning knobs on a mixing board: you change the ratio of plasmids, and the circuit’s behavior changes without needing to redesign any genetic parts. Finally, IANNs degrade gracefully. A small disturbance in the input causes only a small change in the output. Boolean circuits, on the other hand, can flip from the correct answer to the wrong one because of a tiny fluctuation near the switching threshold.
Q2. Applications
IANN could be designed to detect early tumor urinary tumor DNA (utDNA) in dogs by using CRISPR‑based DNA sensors that convert the presence of tumor‑specific mutations into transcriptional inputs for the IANN.
Pre-processing (in vitro): Three CRISPR-Cas13a sensors, each with mutation-specific crRNAs, detect BRAF V595E, TP53 mutations, and aberrant methylation in urine cfDNA. The collateral cleavage activity de-represses synthetic promoters proportionally to how much mutant DNA is present — converting molecular detection into analog transcriptional signals.
Computation (in vivo, HEK293 cells): A two-layer IANN built from sequestrons receives those three graded promoter signals as endoribonuclease inputs. Layer 1 integrates the BRAF and TP53 channels; Layer 2 combines Layer 1’s output with the methylation signal to produce a final weighted decision.
Output: mCherry fluorescence intensity acts as a continuous “cancer probability score” — low for healthy, moderate for single-mutation/early-stage, high for multi-mutation/advanced disease.
The analogy throughout is a team of sniffer dogs reporting to a handler — each dog gives a graded intensity signal for its specific scent (mutation), and the handler weighs them all to decide whether to raise the alarm.
The limitations section covers the real practical hurdles: sensitivity floor of CRISPR sensors for dilute utDNA, the sensor-to-cell interface challenge, transfection variability, the 650 ng DNA budget, temporal lag, incomplete mutation panels, gaps in canine methylome data, and lack of tissue-of-origin discrimination.
CRISPR-to-IANN canine utDNA detection system
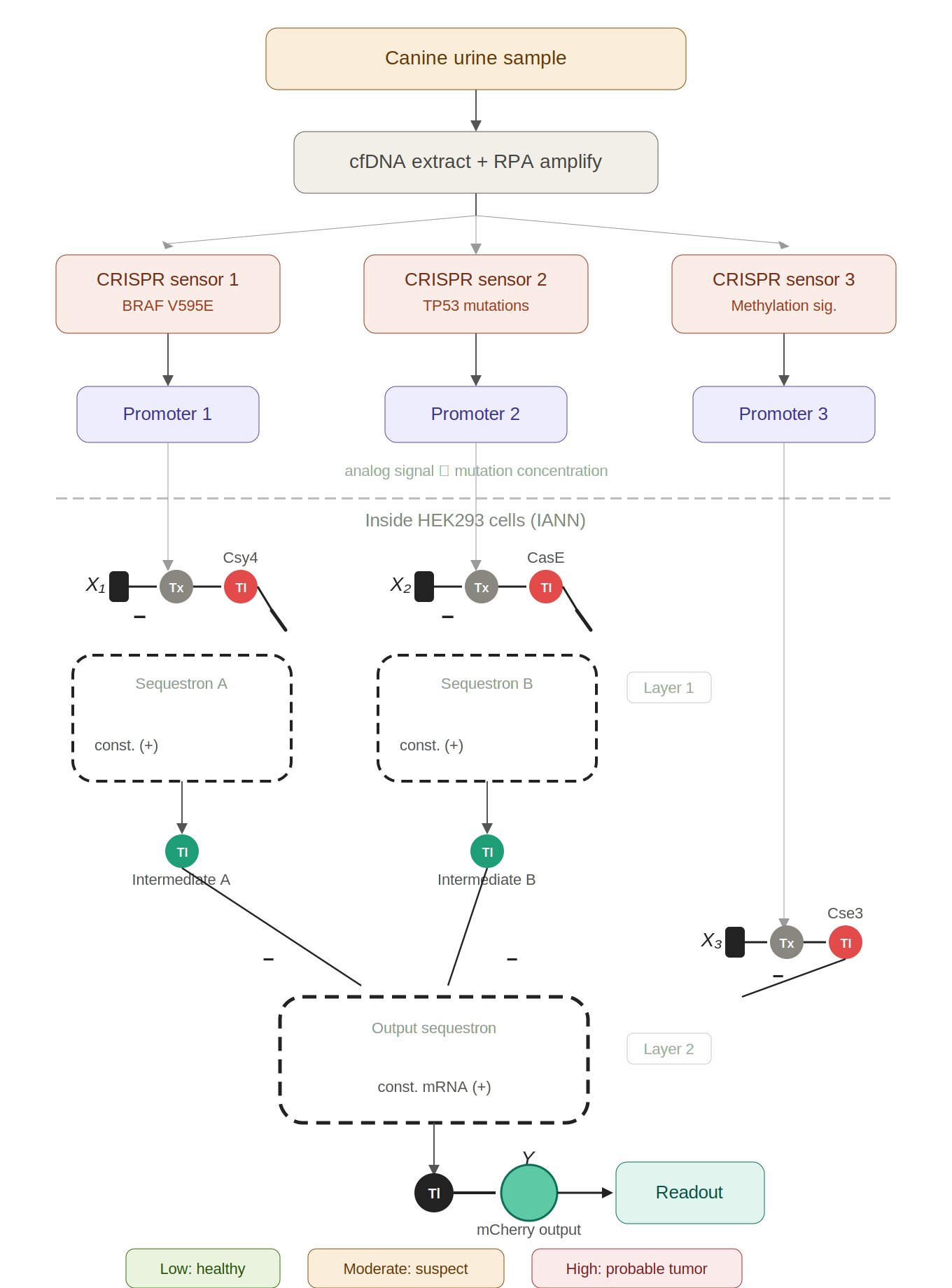

Figure 1: Early canine tumor detection via CRISPR-to-IANN biosensor. Urine cfDNA is amplified and split across three CRISPR-Cas13a sensors targeting BRAF V595E, TP53 mutations, and methylation signatures. Each sensor de-represses a promoter proportionally to mutant utDNA concentration. Inside HEK293 cells, these analog signals feed a two-layer IANN built from sequestrons. The mCherry fluorescence output serves as a continuous cancer probability score.
Q3 Single-layer intracellular perceptron
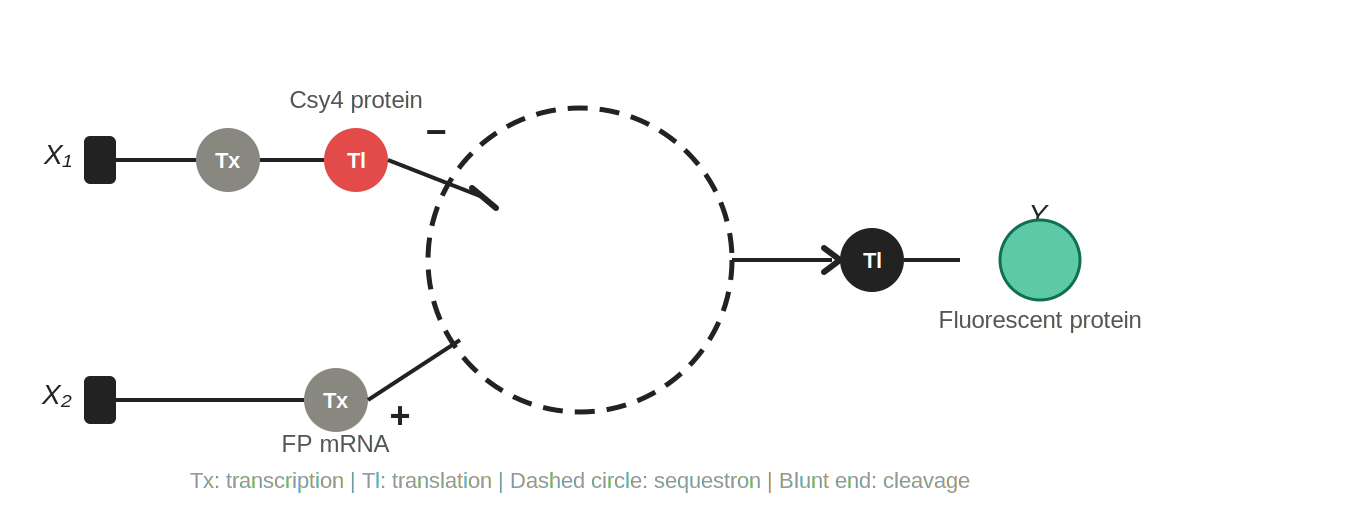

Figure 2: Single-layer intracellular perceptron. X₁ is DNA encoding the Csy4 endoribonuclease; X₂ is DNA encoding a fluorescent protein whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation. The dashed circle represents the sequestron, where Csy4 (−) cleaves the fluorescent protein mRNA (+), and surviving mRNA is translated into the output Y.
Part 2. Fungal Materials
1.What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Mycelium packaging, such as Ecovative’s Mushroom Packaging, is made by growing mycelium on agricultural waste inside a mold. After a few days the material is heat‑killed and dried, producing a rigid, lightweight foam that can replace Styrofoam for protective packaging. It is completely biodegradable, grows on waste feedstocks, and uses little energy to manufacture. However, it has lower compressive strength than Styrofoam, is sensitive to moisture, and is slower to produce at scale.
Mycelium leather, like Bolt Threads’ Mylo and MycoWorks’ Reishi, is grown as a pure sheet in controlled fermentation, then tanned and finished much like animal leather. It is used in fashion and accessories. It requires no animal farming and has a much lower water and land footprint, and its thickness and texture can be tuned. On the downside, it is still expensive at small scale, its durability and aging properties are still being improved, and it needs chemical post‑processing to match the flexibility of animal leather.
Mycelium‑based building insulation is grown on straw or wood shavings and can be used as thermal and acoustic panels. It has heat insulation similar to synthetic foams and is naturally fire‑retardant. It is non‑toxic, sequesters carbon, and resists fire better than petroleum‑based foams. But it is not strong enough for structural uses and can degrade if it gets wet.
Mycoprotein foods like Quorn and Meati come from fermenting filamentous fungi to make high‑protein, fibrous biomass that feels like meat. These products are high in protein, have a complete amino acid profile, and produce far fewer greenhouse gases than animal farming. Still, some people are allergic, the feedstocks are sugar‑based, and the taste and texture are not yet exactly the same as real meat.
Mycelium automotive parts were explored by Ford and Ecovative for interior pieces like dashboards, door panels, and seat cushions, taking advantage of the material’s sound absorption and impact resistance. They are lightweight, need no adhesive because the mycelium acts as the binder, and can be composted at the end of their life. However, they are sensitive to water, there is not yet much data on long‑term durability, and they are not yet cost‑competitive with synthetic foams at automotive scale.
2. What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Bacteria are like bicycles: fast, simple, and cheap, good for making small proteins and simple chemicals. Fungi are like trucks. They are eukaryotes, so they have the cellular machinery to fold complex proteins, add post‑translational modifications, and build intricate three‑dimensional structures that bacteria simply cannot make. One can make mycelium materials that are naturally stronger, more flexible, or more water‑resistant without needing chemical post‑processing. You can also make fungi produce high‑value proteins while they are growing. They could secrete antimicrobial peptides, fire‑retardant proteins, or pigments directly into the material, creating functional composites in a single step. You could create self‑healing materials by engineering dormant spores into dried mycelium composites that reactivate when water enters a crack, similar to how skin heals. You could also enhance bioremediation by engineering white‑rot fungi to produce extra versions of the enzymes that break down plastics, pesticides, or industrial dyes.
Compared to bacteria, fungi have many advantages for synthetic biology. They fold and modify proteins correctly because they have an endoplasmic reticulum and Golgi apparatus, which bacteria lack. Their hyphal growth lets them naturally form sheets, foams, and composites; bacteria only grow as single cells in liquid. Fungi can grow on cheap, unprocessed plant waste because they secrete powerful enzymes to break down cellulose and lignin, while most bacteria need processed sugars. Their eukaryotic compartments let them keep toxic intermediates separate and run incompatible pathways at the same time, which helps them make complex natural products. Genetic tools like CRISPR‑Cas9, promoter libraries, and selectable markers are now well developed in model fungi, making engineering much easier. And because fungi are multicellular, they can form different tissue types such as aerial hyphae or fruiting bodies, opening up possibilities for spatial organization and layered material architectures that are impossible with single‑celled bacteria.