Week 2 HW:Dna Read Write and Edit
Part 1 Benchling & In-silico Gel Art
Limbda Sequence Import


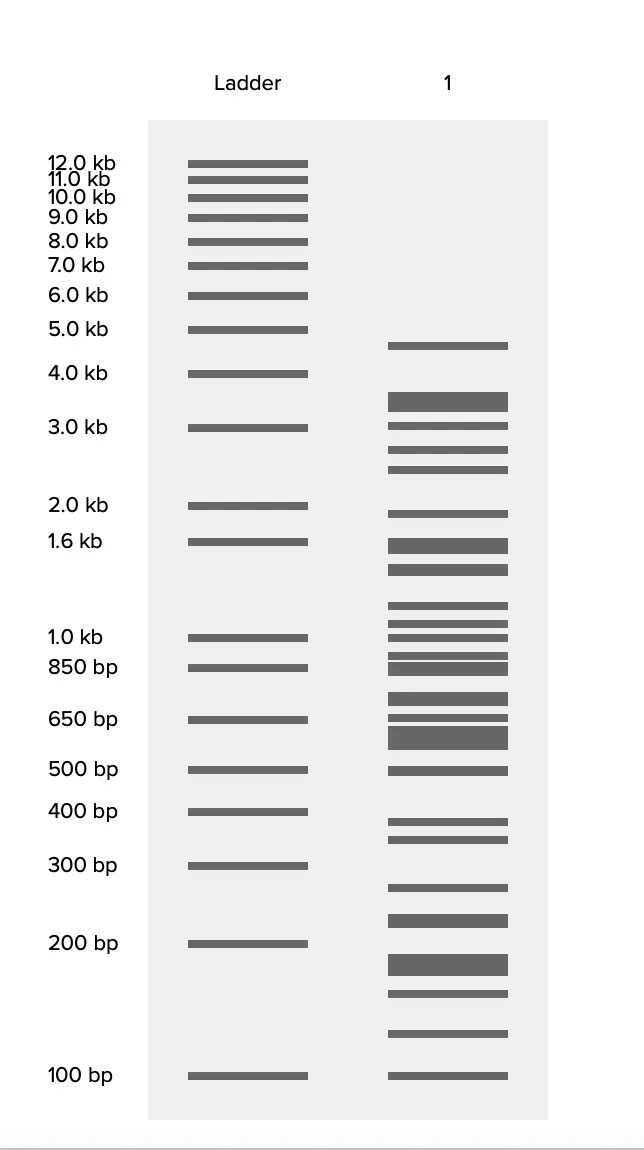
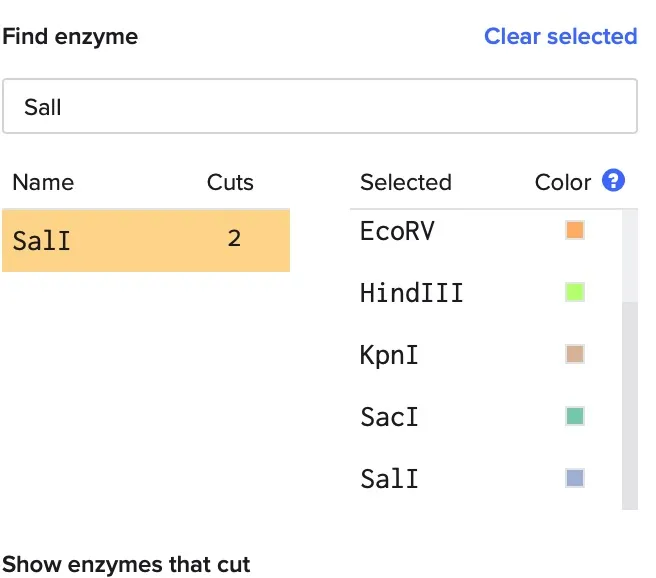
Part 2 Gel Art - Restriction Digests and Gel Electrophoresis
Limbda gel electrophoresis
The style of Paul Vanouse’s Latent Figure Protocol artworks


Part 3 DNA Design Challenge
Protein choose: Oxytocin-neurophysin 1 (Prairie Vole)
[

Why: I chose this protein from the prairie vole (Microtus ochrogaster) because these rodents are famous models in neuroscience for studying monogamy and social bonding. Oxytocin is the key molecular driver of their lifelong pair-bonding behavior. Sequence (FASTA): sp|E0V840|OXYT_MICOC Oxytocin-neurophysin 1 OS=Microtus ochrogaster OX=10031 GN=Oxt PE=3 SV=1 MACPSLACCLLGLLALTSACYIQNCPLGGKRAALDLDTRKCLPCGPGGKGRCFGPNICCADELGCFVGTAEALRCQEENYLPSPCQSGQKPCGSGGRCAAAGVCCSPDGCRMDPACDPESAFSER
I use bioinformatic to reverse translate the protein by bioinformatic
Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence
gcgtgcgatgaatttggccatattaaactgatgaacccgcagcgcagcaccgtgtggtat taaatggcgggcccgagcctggcgtgctgctttctgggcctgctggcgctgaccagcgcg tgctatattcagaactgcccgctgggcggcaaacgcgtgctggatctggatgtgcgcacc tgcccgccgtgcggcccgggcggccgcggccgctgctttggcccgagcatttgctgcgcg gatgaactgggctgctttgtgggcaccgcggaagcgctgcgctgccaggaagaaaactat ctgccgagcccgtgccagagcggccagaaagcgtgcggcagcggcggccgctgcgcggcg ctgggcctgtgctgcagcccggatggctgccatgcggatccggcgtgcgatgcggaagcg acctttagccagcgc
Codon Optimization I have chosen Escherichia coli (E. coli) as the host organism for optimization.
Reason: E. coli is the gold standard “workhorse” in synthetic biology and biotechnology. It has a very rapid growth rate, its genetics are thoroughly understood, and it is highly cost-effective for large-scale recombinant protein production. By optimizing the Prairie Vole oxytocin sequence for E. coli, we ensure that the bacteria can efficiently “read” the instructions and produce high concentrations of the hormone for research or industrial use
Sequence:
gcgtgcgatgaatttggccatatcaagctgatgaacccgcagcgttccaccgtgtggtat taaatggcgggtccgagcctggcgtgctgcttcctgggcctgctggcgctgaccagcgcg tgctacatccagaactgcccgctgggcggcaaacgtgtgctggatctggatgtgcgcacc tgcccgccgtgcggtccgggcggtcgtggccgttgctttggcccgagcatctgctgcgcg gatgaactgggctgctttgtgggcaccgcggaagcgctgcgttgccaggaagaaaactac ctgccgagcccgtgccagagcggccagaaagcgtgcggtagcggcggtcgttgcgcggcg ctgggcctgtgctgcagcccggatggctgccatgcggatccggcgtgcgatgcggaagcg accttcagccagcgt
https://benchling.com/s/seq-8tGivOKSZJUUVEjLY7cE?m=slm-y1fK0BVnbJ3uF2UYnwf2
Part 4: Prepare a Twist DNA Synthesis Order
Build MY DNA Insert Sequence
- I imported Oxytocin-neurophysin 1 (Prairie Vole) using Benchling and created an annotation to describe each piece.


- And then, in the Twsit, the DNA sequence was virtually combined with the vector and exported as a Genbank document.


- This is the plasmid i just built with my expression cassette included. Congratulations on building my first plasmid!


Part 5: DNA Read/Write/Edit
DNA Read
What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
Target DNA: The frozen cell DNA of endangered or extinct species (such as the Bucardo Pyrenean goat or the mammoth) stored in biological repositories like the San Diego Zoo’s “Frozen Zoo”. +2 Reason: These DNA molecules are not merely chemical entities; they are carriers of “Salvage Genomics”. By reading these DNA sequences, we can transform the organism into a “biological information unit”, which is the first step towards achieving “functional de-extinction”, that is, identifying the key genetic characteristics (Functional Traits) that define the species.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Is your method first-, second- or third-generation or other? How so?
What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
What is the output of your chosen sequencing technology?
Technology: Illumina short-read sequencing (NGS) combined with PacBio or Oxford Nanopore (long-read sequencing). Reason: Biological sample libraries (such as cells frozen for several decades) may be subject to degradation. Illumina offers high-precision base coverage, and long-read sequencing is helpful in reconstructing the complete genomic map of extinct species without a reference genome, through De Novo Assembly.
Input and Preparation Steps (Library Preparation):
The genomic DNA extracted from the Frozen Zoo samples, which have been frozen for several decades. Key steps:
Fragmentation: Break down the extracted long-chain DNA. Adapter Ligation: This process connects the specific primers required for sequencing with the sample labels. Library enrichment: The target fragment is amplified through PCR (if it is a second-generation method).
Core steps of sequencing and base identification (Base Calling):
Step: Synthesize fluorescently labeled bases on the flow cell (for sequencing by synthesis method). Base Calling: The machine captures the color of the fluorescence signal produced by each cycle, and converts it into an A, T, C, G sequence through an algorithm.
What is the output?
Output: FASTQ format file containing sequence information and quality scores
DNA Write (i) What DNA would you want to synthesize (e.g., write) and why?
Objective: To synthesize DNA fragments encoding specific functional traits of mammoths (such as cold-resistant hemoglobin genes or thick fur genes). Reason: During the process of extinction, we cannot directly “revive” the old life forms. Instead, by writing these specific sequences, we use them as “designable materials” to create a new organism capable of performing ecological functions (as Novak described it as an ecological substitute). +1
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
Technology: Inkjet-based (inkjet-based) high-throughput DNA synthesis or enzymatic synthesis. Reason: The enzymatic synthesis method is more environmentally friendly and can synthesize longer DNA fragments, making it suitable for synthesizing complex and large-scale genetic circuits like that of a mammoth.
DNA Edit (i) What DNA would you want to edit and why?
Objective: Use the genome of Asian elephant stem cells as a template to remove the specific sites that do not match those of the mammoth. +2
Reason: This editing behavior exemplifies the “deconstruction of species essentialism” as mentioned in RP - species are no longer fixed boundaries, but rather a set of “programmable function libraries” that can be modified, stored, and recombined.
(ii) What technology or technologies would you use to perform these DNA edits and why?
How does your technology of choice edit DNA? What are the essential steps?
What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
What are the limitations of your editing methods (if any) in terms of efficiency or precision?
What technology is being used?
Technology: CRISPR-Cas9 gene editing technology. Reason: It can precisely and efficiently perform multi-site editing within the vast elephant genome, which is the core technology currently adopted by companies like Colossal.
How to conduct editing? Key steps
Design: Design sgRNAs that specifically recognize the genetic loci of mammoth animals. Cutting: The Cas9 protein cuts the double-stranded DNA under the guidance of the gRNA. Repair: Introduce synthetic mammoth DNA template and use homologous recombination repair (HDR) to insert the new sequence into the genome.
Preparation and Input
Input: Asian elephant fibroblast cells, Cas9 protein, designed guide RNA (gRNA), and donor DNA template containing the mammoth sequence.
Limitations of Technology
Off-target effects: May accidentally damage the key survival genes in the Asian elephant genome. Efficiency issue: It is extremely difficult to carry out thousands of gene edits simultaneously within a living cell. Ethical challenges: As you stated in your report, this kind of editing blurs the boundaries between “nature” and “design”, and may give rise to entirely new and difficult-to-define “new life (Neolife)”.