Week 2 HW: DNA Read, Write, and Edit
Assignment Pre-Lecture 2
Professor Jacobson
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
- The error rate for polymerase is 1:106. The human genome is ~3.2 billion base pairs (3.2 x 109). Biology deals with this discrepancy by bridging the gap using a multi-step quality control system to lower the error rate to 1:10^9.
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
- The average human protein is ~1036 base pairs long, which is around 345 amino acids. Since 61 codons encode 20 amino acids, there are approximately 3 codons per amino acid on average. The number of ways to code would be 3^345. Some of the reasons these different codes don’t work include codon bias (different organisms differ in their abundance of specific tRNAs) and GC content (extreme GC content can make DNA difficult to synthesize chemically or unstable for the cell to maintain
Dr. LeProust
What’s the most commonly used method for oligo synthesis currently?
- The industry standard is phosphoramidite chemistry.
Why is it difficult to make oligos longer than 200nt via direct synthesis?
- For a 200nt oligo, the yield is significantly reduced because synthesis is a cyclic process and the efficiency drops exponentially per step. The result would be a mixture dominated by truncated failure sequences.
Why can’t you make a 2000bp gene via direct oligo synthesis?
- A 2000bp strand is impossible based on the yield equation (Yield = EfficiencyN). It would effectively be zero since 0.9952000 = 0.004%.
Dr. Church
- [Using Google & Prof. Church’s slide #4] What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
- The 10 Essential Amino Acids:
- Arginine
- Histidine
- Isoleucine
- Leucine
- Lysine
- Methionine
- Phenylalanine
- Threonine
- Tryptophan
- Valine
- The Lysine Contingency is a concept from Jurassic Park where the dinosaurs were genetically engineered to be “Lysine dependent,” assuming if they escaped they would die. However, there is a crucial caveat: lysine is an essential amino acid. This means that animals cannot synthesize it naturally and must obtain it from their diet. Since lysine is abundant in nature, an escaped dinosaur could easily survive by eating standard food in the wild.
- The 10 Essential Amino Acids:
Assignment Post-Lecture 2
Part 1: Benchling & In-silico Gel Art
See the Gel Art: Restriction Digests and Gel Electrophoresis protocol for details. Overview:
- Make a free account at benchling.com
- Import the Lambda DNA.
- Simulate Restriction Enzyme Digestion with the following Enzymes:
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
- Assignees for the following sections
MIT/Harvard students Required Committed Listeners Optional (for those with Lab access)
Perform the lab experiment you designed in Part 1 and outlined in the Gel Art: Restriction Digests and Gel Electrophoresis protocol.
Part 3: DNA Design Challenge
3.1. Choose your protein. In recitation, we discussed that you will pick a protein for your homework that you find interesting. Which protein have you chosen and why? Using one of the tools described in recitation (NCBI, UniProt, google), obtain the protein sequence for the protein you chose.
Phosphite Dehydrogenase (ptxD) from Pseudomonas stutzeri; This protein is crucial to my proposal for deep space habitation since it catalyzes the oxidation of phosphite (PO33-) into phosphate (PO43-). By deleting native phosphate transporters and inserting ptxD gene, it is possible to achieve synthetic auxotrophy. Ensuring the engineered organisms can only survive if fed an artificial phosphite preventing planetary contamination.
sp|O69054|PTXD_STUST Phosphite dehydrogenase OS=Stutzerimonas stutzeri OX=316 GN=ptxD PE=1 SV=1 MLPKLVITHR VHDEILQLLA PHCELMTNQT DSTLTREEIL RRCRDAQAMM AFMPDRVDAD FLQACPELRV VGCALKGFDN FDVDACTARG VWLTFVPDLL TVPTAELAIG LAVGLGRHLR AADAFVRSGE FQGWQPQFYG TGLDNATVGI LGMGAIGLAM ADRLQGWGAT LQYHEAKALD TQTEQRLGLR QVACSELFAS SDFILLALPL NADTQHLVNA ELLALVRPGA LLVNPCRGSV VDEAAVLAAL ERGQLGGYAA DVFEMEDWAR ADRPRLIDPA LLAHPNTLFT PHIGSAVRAV RLEIERCAAQ NIIQVLAGAR PINAANRLPK AEPAAC
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
3.3. Codon optimization.
Once a nucleotide sequence of your protein is determined, you need to codon optimize your sequence. You may, once again, utilize google for a “codon optimization tool”. In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
Optimization Organism: Bacillus subtilis
Rationale: I have chosen to optimize the sequence for Bacillus subtilis because it serves as the primary “Bio-Pharmacy” chassis in my habitat proposal. Since the habitat’s nutrient system will be standardized on a phosphite-based supply, all biological components (including the B. subtilis pharmaceutical production units) must efficiently express the ptxD protein to survive.
3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
In-vivo (Cell-Dependent Expression): I would clone the DNA into an expression vector (plasmid) containing a strong B. subtilis promoter and transform it into the living cells. The cell’s natural machinery would use RNA polymerase for transcription, and ribosomes for translation that would produce the enzyme as the cells grow.
Cell-Free Protein Synthesis (CFPS): I could use a cell-free “extract” containing all the necessary molecular machinery (ribosomes, tRNAs, polymerases) in a test tube, we can produce in a specific protein without overhead of maintaining a living culture.
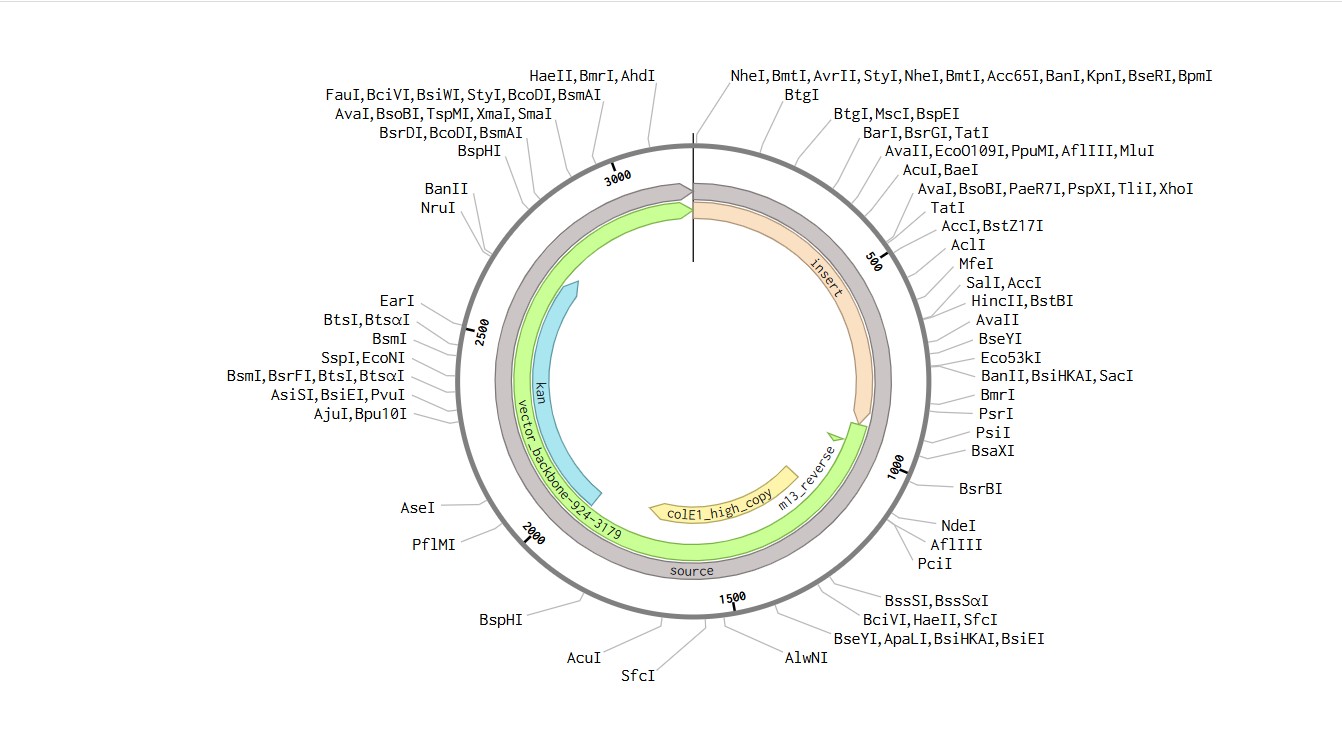
Part 4: Prepare a Twist DNA Synthesis Order

Part 5: DNA Read/Write/Edit Assignees for the following
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank) 1.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
- Oxford Nanopore Technologies (ONT) MinION
(iii) Is your method first-, second- or third-generation or other? How so?
- Third-generation since it performs single-molecule, real-time sequencing without the need for PCR amplification.
(iv) What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
- Input: Pure sample extracted from exoplanet (soil, ice, or liquid)
- Analysis: Design Primers for DNA that is best optimizied for sample data. Produce with Baseline (ELM strain) and customize Living Material in accordance to exoplant sample analysis.
- Fragmentation: Minimal shearing to maintain “Long Reads”.
- End-Repair/A-Tailing: Preparing DNA ends for adapter attachment.
- Adapter Ligation: Attaching motor proteins and sequencing adapters to guide DNA into the pore.
(v) What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
- DNA is taken through a microscopic protein nanopore embedded in an electrically resistant membrane, and when each nucleotide passes through, it causes a specific measurable disruption in the ionic current, and the algorithms translates these info into DNA bases.
(vi) What is the output of your chosen sequencing technology?
- FastQ files containing long-read sequences which allow for the assembly of unknown microbial genomes with no reference template.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
- I want to synthesize a customized ptxD expression cassette designed with modular “adapter regions.” Allowing the ptxD gene (the phosphite-based biocontainment lock) to be rapidly swapped into different host organisms depending on the destination’s gravity and radiation profile.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
- Phosphoramidite Synthesis (Si-based)
(iii) What are the essential steps of your chosen sequencing methods?
- De-blocking: Removing the protective DMT group from the 5’ hydroxyl of the first nucleotide.
- Coupling: Activating and adding the next phosphoramidite nucleotide to the chain.
- Capping: Acetylating any unreacted chains to prevent “deletion” errors.
- Oxidation: Converting the unstable phosphite triester into a stable phosphate triester.
(iv) What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
- Speed: Chemical cycles take minutes per base, making long-gene synthesis time-consuming.
- Accuracy: Error rates increase with length; chemical synthesis typically plateaus around 200bp, requiring enzymatic assembly for larger cassettes.
- Scalability: While high-throughput on silicon chips, the cost scales linearly with length, making whole-genome “writing” prohibitively expensive for remote deployment.
5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?Colossal, Biosciences Inc., biotechnology company that leverages genetic engineering to working to de-extinct various historic animals, such as the woolly mammoth.
- I would edit the chaperone protein genes in the foundry’s microbial population, and by replacing the native promoters with environmentally responsive promoters, it is possible to achieve “self-tuning” meaning the ecosystem would increase metabolic activity or thicken walls of habitat in response to specific atmospheric stressors of exoplanet.
(ii) What technology or technologies would you use to perform these DNA edits and why?
- Targeting: A Guide RNA (gRNA) is designed to match the specific DNA locus of the chaperone promoter.
- Binding: The gRNA directs the Cas9 nuclease to the target site.
- Cleavage: Cas9 induces a Double-Strand Break (DSB).
- Repair: The cell uses Homology-Directed Repair (HDR) to incorporate a provided donor DNA template (the new responsive promoter).
(iii) What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
Preparation & Input:
- Design: Computational modeling of gRNAs to maximize binding efficiency.
- Input: Cas9 protein (or mRNA), gRNA, and the donor DNA template delivered via plasmid or viral vector.
(iv) What are the limitations of your editing methods (if any) in terms of efficiency or precision?
Limitations:
- Efficiency: HDR is naturally less frequent than the error-prone NHEJ pathway, leading to low success rates in certain fungi.
- Precision: Risk of Off-target effects, where the Cas9 cuts at unintended locations, potentially compromising the “Kill Switch” integrity.
Disclaimer: Artificial Intelligence was used in this assignment to assist with conceptual brainstorming, technical copywriting, and formatting. The core scientific concept and final submission were curated by the student.