Week 7 HW: Genetic Circuits Part II — Neuromorphic Circuits
Part 1 — Intracellular Artificial Neural Networks (IANNs)
Q1: Advantages of IANNs over Boolean genetic circuits
Traditional genetic circuits implement Boolean logic: each node is either “on” or “off,” and the circuit computes AND/OR/NOT/NAND operations over binary input signals. This is powerful for simple decision logic but breaks down for complex, real-world biological classification tasks.
IANNs offer the following concrete advantages:
| Property | Boolean Genetic Circuits | Intracellular ANNs |
|---|---|---|
| Output resolution | Binary (on/off) | Continuous, graded (analog) |
| Input integration | Gate-by-gate, combinatorial | Weighted sum over all inputs simultaneously |
| Decision boundaries | Only hyperplane-separable (linear combinations) | Non-linear (sigmoidal / Hill-function activation) |
| Trainability | Manually designed; no learning | Can be parameterised to fit data distributions |
| Number of parts for complexity | Exponential scaling with function complexity | Compact: depth × width rather than truth-table enumeration |
| Noise handling | Binary thresholds amplify noise near threshold | Continuous activation averages over molecular noise |
| Multi-class classification | Requires one circuit per class | Single network with multiple output nodes |
Key biological advantage: Real cellular environments present analog signals — metabolite concentrations, protein levels, pH — that exist on a continuum. IANNs can directly classify these graded inputs using Hill-function activation (analogous to the sigmoid in digital ANNs), without requiring a pre-amplification step to binarise the signal.
Key engineering advantage: For a Boolean circuit to compute an arbitrary function of N inputs, it requires up to 2^N gates. A single-layer perceptron with N inputs computes the same class of functions with only N+1 parameters (weights + bias), making design and genetic implementation far more compact.
Q2: Application for an IANN — Multi-Signal Cancer Biomarker Classifier
Application: Intracellular tumour-microenvironment classifier for targeted drug release
Goal: Engineer a therapeutic E. coli Nissle 1917 that senses three cancer-associated signals inside a tumour and releases a cytotoxic payload only when the weighted combination of signals exceeds a threshold — mimicking a trained classifier, not a simple AND gate.
Input/Output Behaviour
| Signal | Biological source | Intracellular proxy | Weight (conceptual) |
|---|---|---|---|
| X₁: Lactate | Warburg-effect tumour metabolism | Lld promoter (lactate-inducible) | w₁ = +0.6 |
| X₂: Hypoxia (HIF-1α) | O₂-depleted tumour core | Nar promoter (nitrate/anaerobic) | w₂ = +0.5 |
| X₃: Low pH (< 6.8) | Tumour acidosis | AcidR sensor promoter | w₃ = +0.4 |
Hidden layer (IANN neuron): Each promoter drives a different concentration of an RNA-binding protein (e.g., MS2 coat protein variants). The proteins bind different numbers of MS2 stem-loops on a synthetic mRNA encoding a pro-drug converting enzyme (e.g., carboxypeptidase G2, CPG2). The more stem-loops are occupied, the stronger the translational output — computing a weighted, analog sum.
Activation function: Hill-function binding kinetics of each RNA-binding protein to its cognate stem-loop, providing a smooth sigmoid-like threshold.
Output: CPG2 enzyme is produced only when the weighted sum exceeds the threshold. CPG2 converts a systemically administered pro-drug (e.g., ZD2767P) into a locally cytotoxic drug within the tumour, sparing healthy tissue.
Limitations
- Orthogonality: RNA-binding protein variants must not cross-react with each other’s stem-loops or with endogenous mRNAs — limited by the availability of engineered orthogonal systems.
- Weight programmability: Biological “weights” (promoter strength, RBS efficiency, stem-loop affinity) are fixed at construction time. Re-training requires re-engineering the DNA.
- Speed: Transcription/translation timescales (~minutes to hours) are far slower than silicon ANN inference; unsuitable for rapid signalling tasks.
- Metabolic load: Expressing multiple RNA-binding proteins, synthetic mRNAs, and CPG2 simultaneously imposes a fitness cost, selecting for circuit loss over many cell generations.
- In vivo tunability: Tumour microenvironments vary between patients; a fixed-weight circuit trained on population averages may mis-classify in individual patients.
Q3: Multilayer Intracellular Perceptron Diagram
The assignment provides a reference single-layer perceptron in which:
- X₁ input (Csy4 DNA) is transcribed → Csy4 mRNA → translated → Csy4 endoribonuclease protein (the “neuron”)
- X₂ input (FP DNA) is transcribed → FP mRNA → the Csy4 protein cleaves the FP mRNA in a concentration-dependent (analog) manner → translation → FP output
The diagram below shows the reference single-layer architecture first, then extends it to a multilayer design.
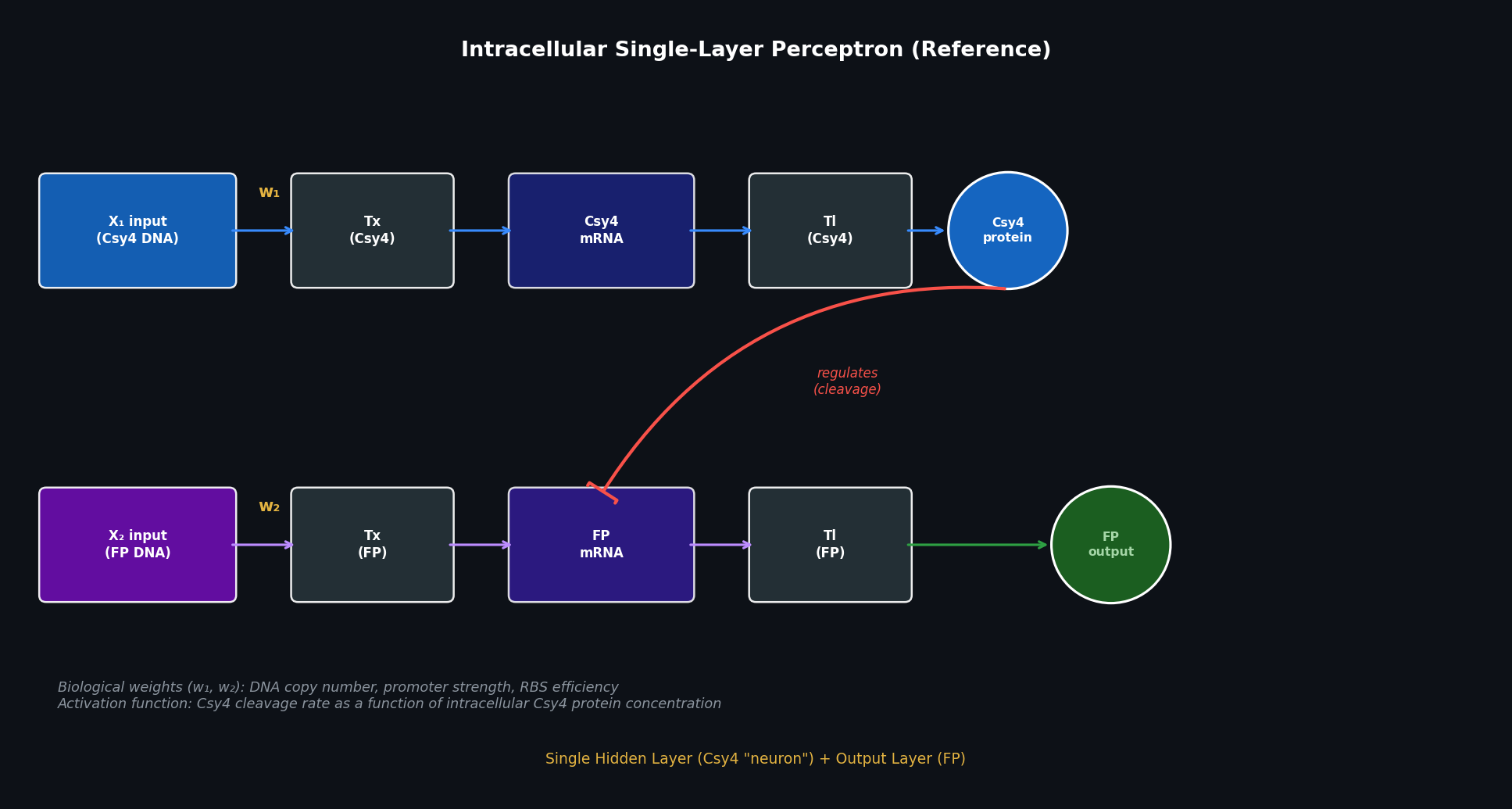
 Fig 1. Reference single-layer intracellular perceptron. X₁ DNA encodes Csy4 endoribonuclease (the “neuron”); X₂ DNA encodes the fluorescent protein output whose mRNA is regulated by Csy4 in an analog, concentration-dependent manner. Biological weights are set by DNA copy number, promoter strength, and RBS efficiency.
Fig 1. Reference single-layer intracellular perceptron. X₁ DNA encodes Csy4 endoribonuclease (the “neuron”); X₂ DNA encodes the fluorescent protein output whose mRNA is regulated by Csy4 in an analog, concentration-dependent manner. Biological weights are set by DNA copy number, promoter strength, and RBS efficiency.
Multilayer Design
To build a multilayer perceptron, I introduce a hidden layer between the input DNA concentrations and the final fluorescent protein output:
- Layer 1 (Input → Hidden): Three input DNA constructs (X₁, X₂, X₃) each drive transcription of a distinct mRNA. After translation, three different endoribonuclease-family proteins are produced (Csy4 homolog 1, RNase L1 variant, and an engineered ribozyme protein). Each protein’s expression level is a weighted function of the input DNA concentrations — forming the first set of “neurons.”
- Layer 2 (Hidden → Output): The three Layer 1 endoribonucleases converge on a single fluorescent protein mRNA. Each enzyme cleaves a different stem-loop structure within the FP mRNA 5’ UTR, collectively suppressing or permitting FP translation in proportion to the weighted combination of Layer 1 outputs.
- Output: The FP protein level represents the multilayer network’s final classification of the three-dimensional input.
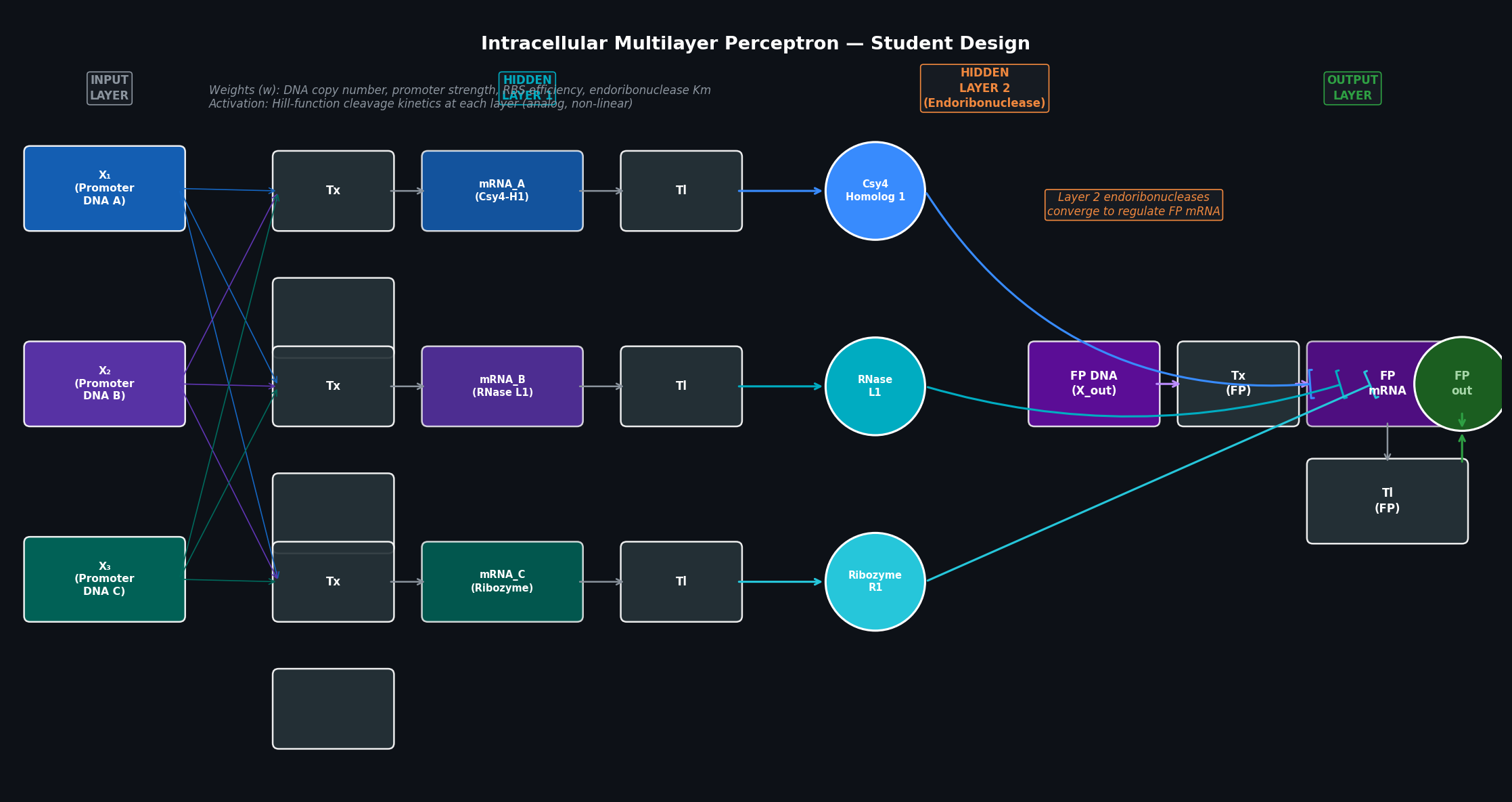
 Fig 2. Multilayer intracellular perceptron. Three DNA inputs (X₁–X₃) feed into three parallel transcription/translation units (Hidden Layer 1), each producing a distinct endoribonuclease. These three enzymes converge on the FP mRNA in Hidden Layer 2, regulating its stability and translation rate. The output (FP protein level) represents an analog weighted classification over all three inputs. Tx = transcription; Tl = translation.
Fig 2. Multilayer intracellular perceptron. Three DNA inputs (X₁–X₃) feed into three parallel transcription/translation units (Hidden Layer 1), each producing a distinct endoribonuclease. These three enzymes converge on the FP mRNA in Hidden Layer 2, regulating its stability and translation rate. The output (FP protein level) represents an analog weighted classification over all three inputs. Tx = transcription; Tl = translation.
Key design principles:
- Orthogonal endoribonuclease families (Csy4 and its engineered homologs) ensure each hidden-layer neuron targets only its designated mRNA stem-loop
- The FP mRNA carries three distinct stem-loop architectures in its 5’ UTR, each specifically bound by one Layer 2 enzyme
- Weight magnitudes are set at construction time by promoter strength and RBS efficiency; they are fixed (unlike trained digital ANNs)
Part 2 — Fungal Materials
Q1: Examples of Existing Fungal Materials
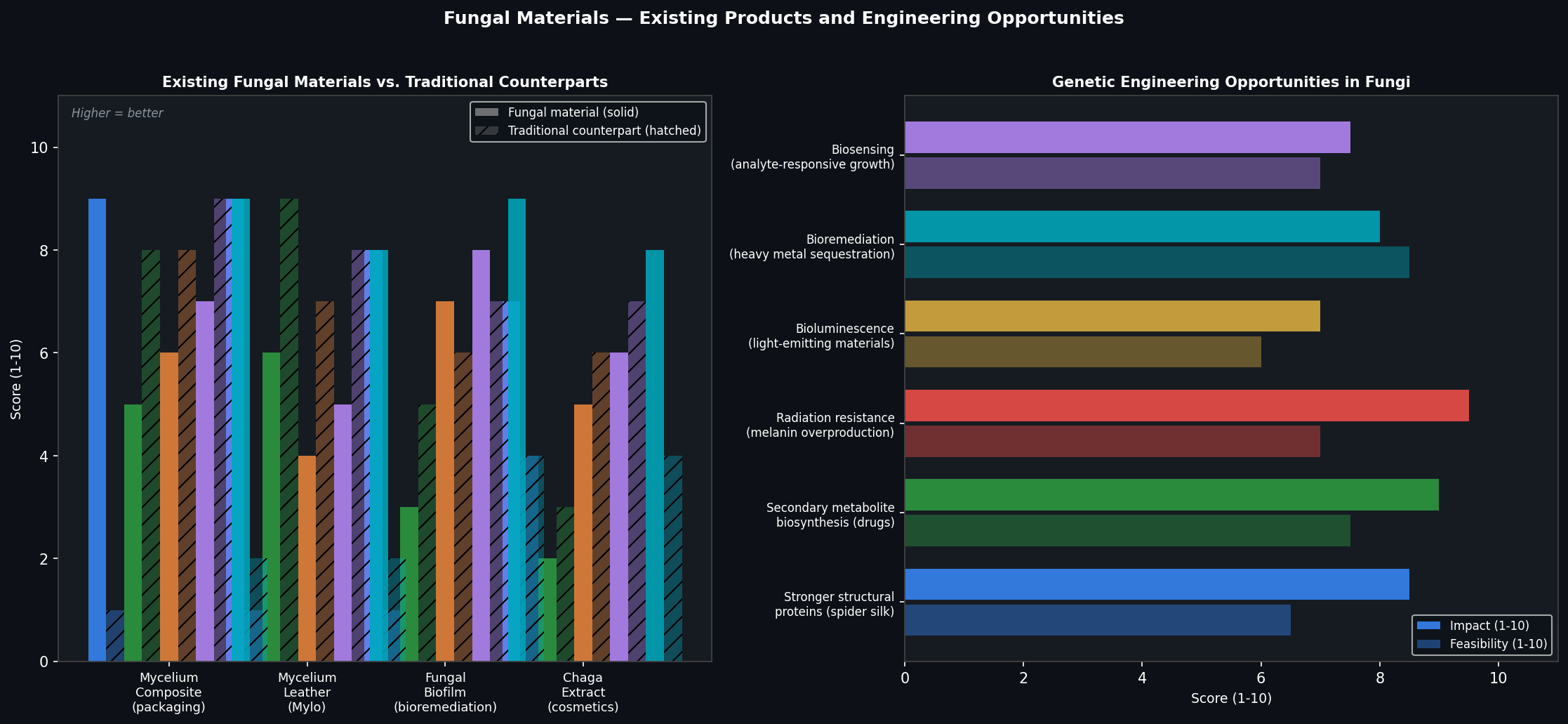
 Fig 3. Left: comparative property scores for existing fungal materials vs. traditional counterparts across five dimensions. Right: impact and feasibility scores for six genetic engineering opportunities in fungal systems.
Fig 3. Left: comparative property scores for existing fungal materials vs. traditional counterparts across five dimensions. Right: impact and feasibility scores for six genetic engineering opportunities in fungal systems.
Current commercial and research fungal materials:
| Material | Product / Company | Application | Fungal species |
|---|---|---|---|
| Mycelium composite | Ecovative Design (EcoCradle, AirMycelium) | Packaging, insulation, replacing EPS foam | Ganoderma spp. |
| Mycelium leather | Bolt Threads (Mylo), MycoWorks (Reishi) | Textile / fashion leather alternative | Ganoderma lucidum, Phanerochaete |
| Fungal biofilm mat | Research stage (e.g., Colorifix, Biocouture) | Bioremediation, dye production | Aspergillus niger, Trichoderma |
| Chitosan film | Various (food packaging, wound dressings) | Antimicrobial barrier | Rhizopus spp., Mucor |
| Ergot alkaloids / penicillin | Pharmaceutical industry | Drug production | Claviceps, Penicillium |
Advantages over traditional counterparts:
- Biodegradability: Mycelium composites decompose in soil within 30–90 days; polystyrene foam persists > 500 years
- Carbon footprint: Grown on agricultural waste (hemp hurds, corn stalks); net CO₂-negative lifecycle vs. petroleum-derived foam
- No toxic inputs: No petrochemical feedstock; no solvent processing
- Structural tunability: Growth conditions (humidity, CO₂, substrate) allow tuning of mechanical properties (density, compressive strength) without changing the genetic composition
Disadvantages:
- Lower mechanical strength: Mycelium composites have compressive strength of ~0.1–0.5 MPa vs. ~20–80 MPa for structural polymers — unsuitable for load-bearing applications without reinforcement
- Moisture sensitivity: Ungrown mycelium rehydrates and loses structural integrity above ~70% relative humidity
- Production time: 4–10 days of growth vs. hours for injection-moulded plastics
- Batch variability: Biological growth introduces variability not present in synthetic manufacturing
- Regulatory burden: Novel fungal biomaterials face extensive safety testing; some mycelium-derived products still lack GRAS status
Q2: Genetic Engineering of Fungi — Motivations and Advantages over Bacteria
What to engineer and why:
1. Structural protein integration (spider silk / amyloid fibres) Engineer Ganoderma or Aspergillus to secrete MaSp1/MaSp2 spider silk proteins that self-assemble along hyphal walls during growth. This would create mycelium composites with tensile strength approaching Kevlar (~3.5 GPa), enabling structural aerospace components grown from agricultural waste — directly applicable to the ELM deep-space habitat.
2. Secondary metabolite production (pharmaceuticals) S. cerevisiae and A. niger already produce penicillin, statins, and artemisinic acid. Engineering B. subtilis-equivalent productivity into mycelium-forming fungi would allow pharmaceutical payloads to be embedded directly within the structural habitat material — the central goal of the ELM Multi-Trophic Myco-Foundry.
3. Radiation-melanin overproduction Radiotrophic fungi (Cladosporium sphaerospermum, Cryptococcus neoformans) use melanin as an energy-harvesting radiation shield. Engineering this pathway into the structural shell species of the ELM habitat would provide passive radiation protection on the Mars surface.
4. Bioluminescent indicators Fungal bioluminescence (Neonothopanus nambi 4-gene cassette) can be introduced to provide visual health readouts of the living material system — a glow indicates metabolic activity; dimming indicates stress.
Advantages of synthetic biology in fungi vs. bacteria:
| Feature | Fungi | Bacteria |
|---|---|---|
| Macroscopic 3D structure | Mycelium forms centimetre-to-metre-scale architectures naturally | Require scaffolding or biofilm engineering for 3D form |
| Post-translational modifications | Eukaryotic: glycosylation, disulfide bonds, correct protein folding | Prokaryotic: lack glycosylation; many human proteins misfold |
| Secondary metabolite pathways | Rich natural biosynthetic gene clusters (PKS, NRPS); many drugs already from fungi | More limited secondary metabolism |
| Mechanical output | Structural mycelium = built-in material scaffold | No equivalent self-assembling macrostructure |
| Gene regulation complexity | Introns, histone modification, chromatin remodelling allow nuanced control | Primarily operon-based; less epigenetic layering |
| Containment | Non-sporulating mutants are easily contained; growth stops when substrate is consumed | Horizontal gene transfer is a significant biocontainment concern |
Key limitation of fungal SynBio: Genetic tools are less mature than for E. coli. CRISPR-Cas9 transformation efficiency in Ganoderma and other basidiomycetes remains 10–100× lower than in S. cerevisiae, and reliable promoter libraries are sparse. However, this gap is closing rapidly (Kück & Hoff, 2010; Wenderoth et al., 2017).
Part 3 — First DNA Twist Order
Aim 1 Draft
Title: Phosphite Auxotrophy Biocontainment System for ELM Deep-Space Habitats
Aim 1: Engineer and validate a synthetic phosphite auxotrophy kill switch in Bacillus subtilis by integrating the Pseudomonas stutzeri ptxD gene (phosphite dehydrogenase) under a constitutive promoter and deleting the endogenous phoA alkaline phosphatase locus, such that engineered cells grow only in phosphite-supplemented media and are inviable in standard phosphate media or uncontrolled environments.
Deliverable: Characterise the phosphite IC₅₀ using an Opentrons-automated 96-well growth screen (Week 3 automation pipeline) and confirm zero-growth in phosphate-only controls over 72 h.
Final Project Summary
The Multi-Trophic Myco-Foundry (ELM Habitat) is an engineered living material proposed as a self-sustaining deep-space habitat module for long-duration crewed Mars missions. The system integrates three biological layers — a radiation-shielding fungal structural shell, a bacterial vascular network for nutrient transport, and a pharmaceutical biosynthesis core — into a single co-cultivated mycelium composite. Key engineering goals include:
- Biocontainment: Phosphite auxotrophy + quorum-sensing kill switch (Week 6 Construct 1 + 2)
- Radiation resilience: Melanin-overproducing Ganoderma shell + SOS-CRISPR repair in bacteria (Week 6 Construct 3)
- Pharmaceutical production: CFPS biosensor-validated drug synthesis in B. subtilis core (Week 4 ESMFold/ProteinMPNN design)
- Structural integrity: Spider silk protein co-expression in mycelium growth phase
Insert Sequence Design for DNA Twist
The insert ordered from DNA Twist encodes the phosphite-inducible ptxD-sfGFP biosensor described in the Week 6 Golden Gate Assembly Benchling model. This serves as the first experimental validation module for the ELM habitat’s phosphite auxotrophy system.

 Fig 4. Top: linear part map of the 1,933 bp DNA Twist insert showing all functional elements and Golden Gate-compatible BsaI overhangs. Bottom: key sequence features at the 5’ junction (BsaI site, AATG overhang, P_ptxD promoter core, RBS B0034, ptxD ATG start) and backbone/cloning summary table.
Fig 4. Top: linear part map of the 1,933 bp DNA Twist insert showing all functional elements and Golden Gate-compatible BsaI overhangs. Bottom: key sequence features at the 5’ junction (BsaI site, AATG overhang, P_ptxD promoter core, RBS B0034, ptxD ATG start) and backbone/cloning summary table.
Insert specification:
| Parameter | Value |
|---|---|
| Total insert length | 1,933 bp |
| Backbone vector | pSEVA221 (pBBR1 ori, KanR, broad-host-range) |
| Assembly method | Golden Gate (BsaI-HFv2, NEB) |
| 5’ overhang | AATG (joins to backbone left flank) |
| 3’ overhang | GTGC (joins to backbone right flank) |
| Synthesis vendor | DNA Twist (standard turnaround, codon-optimised for B. subtilis) |
| Cloning host | E. coli DH5α |
| Expression host | B. subtilis 168 (ATCC 23857) |
Insert part structure (5’ → 3’):
Representative 5’ junction sequence (first 80 bp of insert):
(P_ptxD synthetic core + RBS B0034 + ptxD initiation; full sequence in Benchling shared folder)
Backbone: pSEVA221
pSEVA221 was selected because:
- Broad-host-range replication (pBBR1 ori): Functions in both E. coli (cloning) and B. subtilis (expression) without re-cloning
- Kanamycin resistance: Orthogonal to the ampicillin selection used in other constructs in the lab; reduces cross-contamination risk
- Minimal backbone: 4.2 kb, low metabolic load, well-characterised MCS
- SEVA standard compliance: Compatible with the Standard European Vector Architecture modular cloning system used by collaborators
Connection to final project:
This insert is the first physically synthesised component of the ELM Habitat project. Upon successful cloning and verification (colony PCR + Sanger sequencing of the insert junctions), the construct will be:
- Transformed into B. subtilis 168 for phosphite-inducible GFP validation
- Used as the template for the Opentrons-automated phosphite IC₅₀ screen (Week 3 automation pipeline, Q2)
- Integrated into the ptxD knock-in locus (amyE::ptxD) for chromosomal auxotrophy validation
AI Disclosure
Claude Sonnet 4.6 (Anthropic) was used to assist with IANN mechanism descriptions, multilayer perceptron diagram design, fungal materials literature synthesis, ODE model structure for IANN applications, and figure generation code. The ELM habitat project concept, final project aims, DNA insert design rationale, and biological engineering connections were developed by the student.