Week 9 HW: Cell-Free Systems
Part A — General Questions
Q1: Advantages of Cell-Free Protein Synthesis over Traditional In Vivo Methods
Cell-free protein synthesis (CFPS) decouples protein production from cell viability, providing two structural advantages:
Flexibility — the reaction composition is fully user-controlled. Template DNA, cofactors, non-natural amino acids, detergents, redox buffers, and labeled substrates can be added directly at any concentration without membrane barriers or cell toxicity constraints. Reaction volumes can range from nanolitres (acoustic dispensing) to litres (batch bioreactor).
Control over experimental variables — Mg²⁺, K⁺, pH, temperature, ATP concentration, and reducing potential can be individually tuned between reactions in a plate-reader format. A single cell-free extract functions as a universal chassis: only the DNA template changes.
Two cases where CFPS outperforms cell production:
| Case | Why cell-free wins |
|---|---|
| Toxic or membrane-disrupting proteins (e.g., pore-forming toxins, antimicrobial peptides) | Expressing these in living cells kills the host before significant product accumulates. In CFPS the cell is already lysed — the protein can be produced without lethality. |
| Non-natural amino acid (ncAA) incorporation (e.g., azide-tagged, fluorescent, or photocrosslinking AAs) | CFPS allows amber stop-codon suppression with orthogonal tRNA/synthetase pairs and exogenously supplied ncAAs at defined concentrations — achieving near-100% ncAA incorporation at targeted sites without the metabolic dilution inevitable in living cells. |
Additional cases: rapid prototyping of genetic circuits (no cloning/transformation cycle), production of disulfide-bonded proteins in a controlled redox environment, and isotope-labelling for NMR without cell growth.
Q2: Main Components of a Cell-Free Expression System
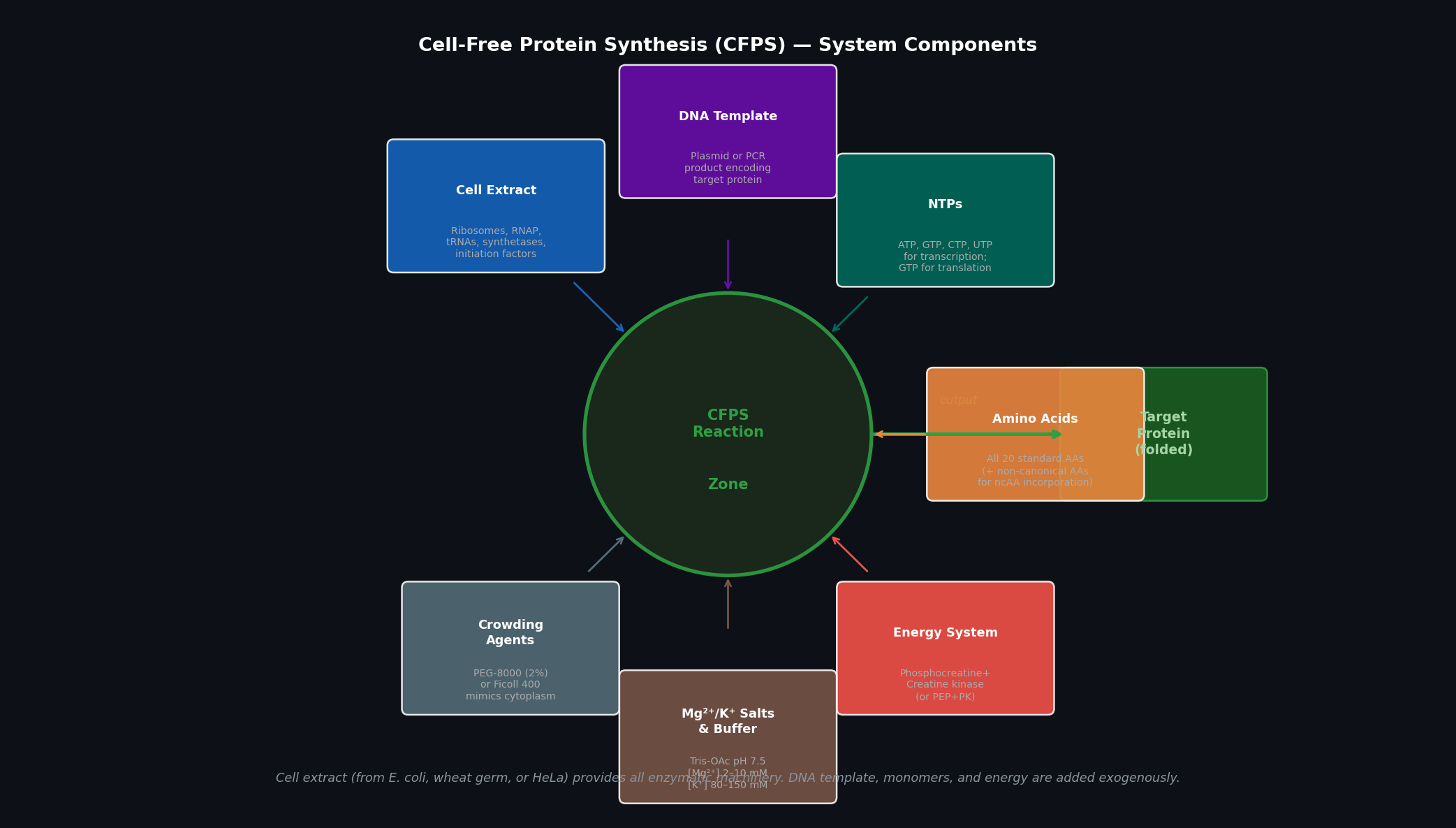
 Fig 1. Cell-free protein synthesis (CFPS) system components. Seven input categories feed into the central reaction zone; the output is folded target protein. The cell extract provides all enzymatic machinery; all remaining components are supplied exogenously and can be independently titrated.
Fig 1. Cell-free protein synthesis (CFPS) system components. Seven input categories feed into the central reaction zone; the output is folded target protein. The cell extract provides all enzymatic machinery; all remaining components are supplied exogenously and can be independently titrated.
| Component | Role |
|---|---|
| Cell extract | Source of ribosomes (~70S or ~80S), RNA polymerase, tRNAs, aminoacyl-tRNA synthetases, initiation/elongation/termination factors, chaperones, and metabolic enzymes. Most commonly E. coli S30 extract (crude) or S12 (clarified). |
| DNA template | Encodes the protein of interest. Can be a plasmid (circular, more stable) or a linear PCR product (faster to produce; susceptible to exonucleases — protect with chi-site sequences or RecBCD inhibitors). Typically under a T7 or SP6 promoter. |
| NTPs (ATP, GTP, CTP, UTP) | Substrates for RNA polymerase during transcription; GTP is additionally consumed by elongation factors (EF-Tu) and ribosomes during translation. Starting concentrations: 1–2 mM each. |
| Amino acids (all 20) | Substrates for aminoacyl-tRNA synthetases; incorporated into the growing polypeptide. Typical concentration: 1–2 mM each. Non-canonical AAs can be added here for genetic code expansion. |
| Energy regeneration system | ATP and GTP are consumed rapidly; regeneration systems (phosphocreatine + creatine kinase, or PEP + pyruvate kinase) recycle ADP → ATP to maintain the energy charge. Critical for reaction duration beyond ~1 h. |
| Mg²⁺/K⁺ salts and buffer | Mg²⁺ (2–12 mM, titrated) stabilises ribosome structure and is a polymerase cofactor; K⁺ (80–150 mM) maintains the ionic environment for translation. Buffer (Tris-OAc or HEPES, pH 7.4–7.6) maintains pH. |
| Crowding agents | PEG-8000 (2%) or Ficoll-400 mimics the macromolecular crowding of the cytoplasm (~300–400 g/L total macromolecule concentration), boosting effective enzyme concentrations and translation rates. |
Q3: Why Energy Regeneration is Critical — Ensuring Continuous ATP Supply
Why it matters: Transcription consumes NTPs stoichiometrically (one ATP/GTP/CTP/UTP per base incorporated). Translation is even more energy-intensive: each amino acid incorporation requires 4 high-energy phosphate bonds (2 ATP for aminoacylation, 1 GTP for EF-Tu delivery, 1 GTP for translocation). A 300-residue protein costs ~1,200 GTP equivalents. Without regeneration, the ATP pool (typically 1–2 mM at start) is exhausted in under 30 minutes.
How energy is regenerated:
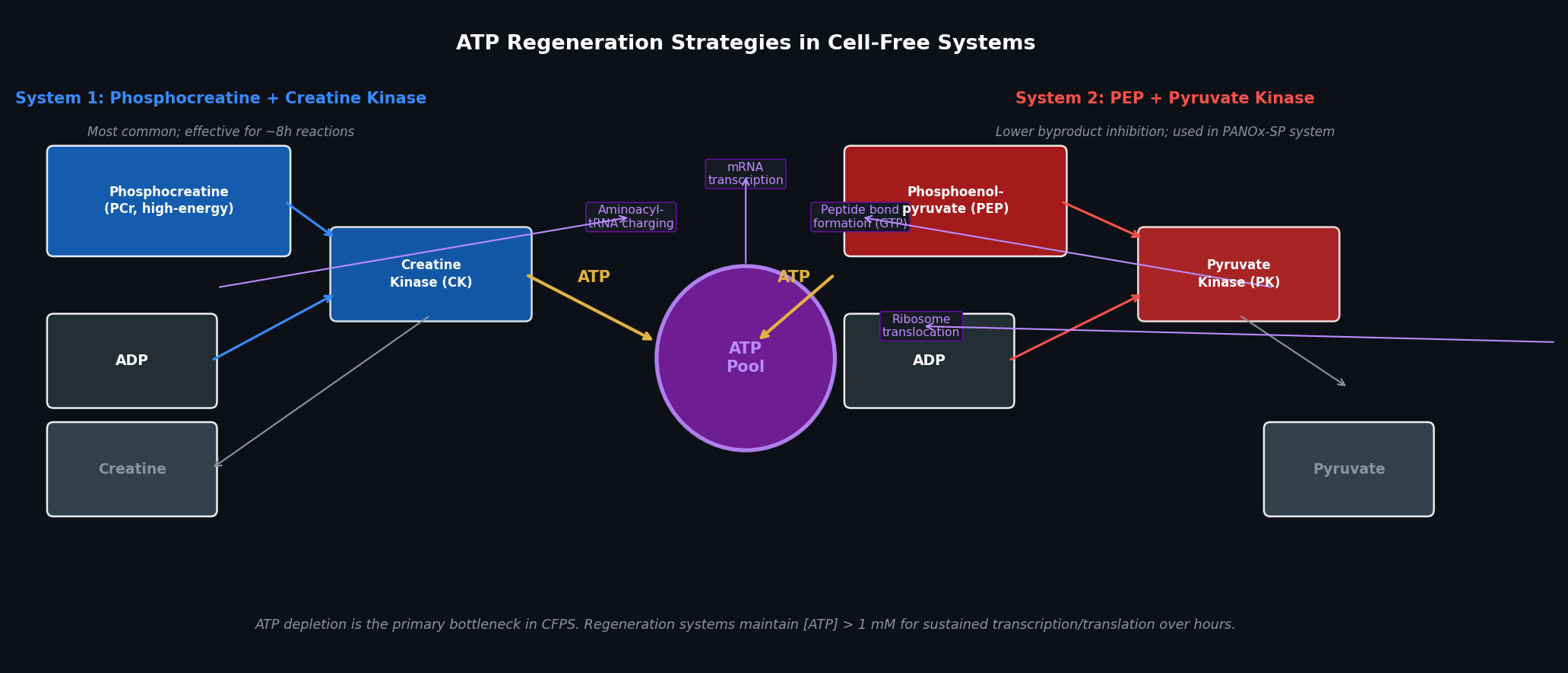
 Fig 2. Two principal ATP regeneration systems used in CFPS. Left: phosphocreatine (PCr) + creatine kinase (CK) — the most common system, effective for 6–8 h. Right: phosphoenolpyruvate (PEP) + pyruvate kinase (PK) — used in the PANOx-SP system (Jewett & Swartz, 2004); lower byproduct inhibition for reactions requiring >8 h. Both systems feed a central ATP pool consumed by transcription and translation.
Fig 2. Two principal ATP regeneration systems used in CFPS. Left: phosphocreatine (PCr) + creatine kinase (CK) — the most common system, effective for 6–8 h. Right: phosphoenolpyruvate (PEP) + pyruvate kinase (PK) — used in the PANOx-SP system (Jewett & Swartz, 2004); lower byproduct inhibition for reactions requiring >8 h. Both systems feed a central ATP pool consumed by transcription and translation.
Method for continuous ATP supply — PANOx-SP (preferred for long reactions):
In the PANOx-SP system (Jewett & Swartz, Biotechnol Bioeng 2004):
- PEP (phosphoenolpyruvate, 30 mM) donates its phosphate to ADP via pyruvate kinase, regenerating ATP
- NAD⁺ (0.33 mM) and oxalic acid extend reaction lifetime by reducing pyruvate accumulation (a feedback inhibitor of pyruvate kinase)
- Spermidine (1 mM) and putrescine (1.5 mM) stabilise ribosome structure
This system sustains productive synthesis for 10–14 hours, compared to ~3 hours for simple phosphocreatine systems. For the ptxD biosensor work in the ELM habitat, PANOx-SP is the preferred energy system because the phosphite dehydrogenase reaction (NAD⁺ reduction) can additionally be used to assess whether the expressed ptxD is enzymatically active in the cell-free context.
Q4: Prokaryotic vs. Eukaryotic Cell-Free Expression Systems
| Feature | Prokaryotic (E. coli S30) | Eukaryotic (Wheat Germ / Rabbit Reticulocyte / HeLa) |
|---|---|---|
| Extract preparation | Exponential-phase cells lysed by French press or bead milling | Plant embryo milling (WG) or reticulocyte lysis (RRL); more complex |
| Translational machinery | 70S ribosomes, σ70-based transcription (needs T7 RNAP added) | 80S ribosomes; supports cap-dependent translation |
| Post-translational modifications | No glycosylation; limited disulfide bonding | Glycosylation (HeLa extract), disulfide bonding, signal peptide cleavage |
| Yield | High: 0.5–2 mg/mL protein | Lower: 10–200 µg/mL protein |
| Cost and speed | Low cost, fast extract preparation | Higher cost, more variable |
Protein choice for prokaryotic system: ptxD (phosphite dehydrogenase, 24 kDa) ptxD is a soluble, cytoplasmic enzyme from Pseudomonas stutzeri with no disulfide bonds or glycosylation. E. coli CFPS produces it at high yield (>1 mg/mL) with correct folding and NAD⁺-dependent enzymatic activity. The system is compatible with direct addition of ¹⁵N-labelled amino acids for NMR structural validation.
Protein choice for eukaryotic system: Erythropoietin (EPO, 30 kDa glycoprotein) EPO requires N-linked glycosylation at Asn24, Asn38, and Asn83 for correct folding, receptor binding, and in vivo half-life. E. coli CFPS produces only aglycosylated EPO with sharply reduced activity. A HeLa-cell CFPS extract (or wheat germ + microsomes) provides the oligosaccharyltransferase machinery required for correct glycoform addition.
Q5: Designing a Cell-Free Experiment to Optimise Membrane Protein Expression
Membrane proteins account for ~30% of the proteome but are notoriously difficult to produce in conventional CFPS: their hydrophobic transmembrane helices cause aggregation in aqueous reaction buffers.
Challenges and solutions:
| Challenge | Strategy |
|---|---|
| Aggregation upon synthesis in aqueous environment | Supply nanodiscs (MSP + lipid mix) or lipid vesicles (liposomes) directly in the reaction — the nascent protein partitions into the bilayer co-translationally |
| Detergent requirement | Add DDM (n-dodecyl-β-D-maltoside, 0.1–1%) or LMNG to solubilise the protein and prevent aggregation; must titrate carefully (high detergent inhibits ribosomes) |
| Chaperone requirement | Supplement with GroEL/GroES or DnaK/DnaJ/GrpE to assist co-translational folding |
| Low yield | Use a dialysis-based cell-free system (continuous-exchange CFPS, CECF) — fresh substrates diffuse in, products diffuse out through a membrane, sustaining synthesis 10× longer than batch |
| Activity verification | For a channel or transporter, reconstitute expressed protein into proteoliposomes and measure ion flux (patch-clamp) or fluorescent substrate uptake |
Experimental design for a GPCR (e.g., beta-2 adrenergic receptor, B2AR):
- Prepare E. coli S30 extract supplemented with 0.1% DDM and pre-formed DMPC nanodiscs (1 µM MSP1D1)
- Add linear template encoding B2AR with a C-terminal FLAG tag (for pull-down) under T7 promoter
- Run CECF reaction at 30°C for 12 h (lower temperature reduces aggregation)
- Verify co-translational nanodisc insertion by size-exclusion chromatography (shift to ~200 kDa nanodisc peak)
- Titrate [DDM] and [nanodisc] across a 96-well plate to maximise folded yield (western blot with anti-FLAG)
- Confirm ligand binding (radioligand binding assay with ³H-dihydroalprenolol)
Q6: Troubleshooting Low Protein Yield in a Cell-Free System
| Reason | Diagnostic | Strategy |
|---|---|---|
| 1. DNA template quality or concentration suboptimal | Run SDS-PAGE after 2 h; compare mRNA levels by dot-blot with a probe against your transcript | Test a range of template concentrations (1–20 nM plasmid or 5–100 nM linear). Add linear template chi-sequence extensions or use a plasmid. Verify template by gel electrophoresis to confirm no degradation. |
| 2. Mg²⁺ concentration not optimised | Systematic grid: vary [Mg²⁺] from 2–16 mM in 1 mM steps across a 96-well plate; read yield by fluorescence (GFP) at 4 h | Mg²⁺ is the most sensitive parameter in E. coli CFPS — optimal [Mg²⁺] varies ±2 mM between extract batches. Peak yield often at 6–10 mM. Each new extract batch requires re-optimisation. |
| 3. Protein misfolding or degradation | Add protease inhibitor cocktail (PMSF, EDTA) to test for proteolysis; add molecular chaperones (GroEL/GroES) to test for folding defect; confirm via anti-His western vs. functional assay | Supplement with disulfide bond isomerase (DsbC) for disulfide-containing proteins; add GroEL/GroES (1 µM each); conduct reaction at 25°C instead of 37°C to slow folding kinetics and reduce aggregation |
Kate Adamala Question — Synthetic Minimal Cell Design
Design: “ELM Phosphite Sentinel”
1. Function
The Phosphite Sentinel is a synthetic minimal cell (SMC) that monitors extracellular phosphite concentration and communicates the habitat health status to surrounding B. subtilis reporter bacteria — serving as an analog biosensor and inter-cellular messenger system within the Multi-Trophic Myco-Foundry ELM habitat.
2. Input and Output
- Input: Extracellular phosphite (PO₃²⁻) concentration
- Output: IPTG released into the surrounding medium (when phosphite is HIGH → habitat healthy) or IPTG withheld (when phosphite is LOW → biocontainment stress, kill-switch risk)
- Downstream output: B. subtilis cells carrying P_lac::GFP fluoresce green when IPTG is present (phosphite abundant) and are dark when phosphite falls below threshold (alert state)
3. Could this be realised by cell-free Tx/Tl alone (without encapsulation)?
No. If IPTG were not encapsulated inside a vesicle, it would freely diffuse into the surrounding bacteria regardless of phosphite concentration, eliminating the switch behaviour. Encapsulation is essential: the SMC only releases IPTG when phosphite-responsive α-hemolysin (aHL) pores are expressed and assembled into the membrane, physically gating IPTG efflux.
4. Could this be realised by a genetically modified natural cell?
Yes, in principle: a B. subtilis cell could be engineered to express lacZ or sfGFP under a ptxD-responsive promoter. However, this would require transforming every B. subtilis reporter cell with the ptxD sensing construct, and the sensing and reporting functions would be co-located in the same cell. The SMC architecture separates these roles — any downstream cell type (including mammalian cells) can be used as the reporter without modification, making the SMC a universal phosphite-sensing actuator.
5. Desired Outcome
In the presence of sufficient phosphite (>1 mM), α-hemolysin is expressed inside the SMC, assembles into the lipid bilayer, and releases pre-encapsulated IPTG into the habitat medium. Surrounding B. subtilis cells induce GFP expression, confirming that: (a) phosphite is available, (b) the ptxD-dependent biocontainment system is functioning, and (c) the synthetic cell is intact and metabolically active.
6. Component Design
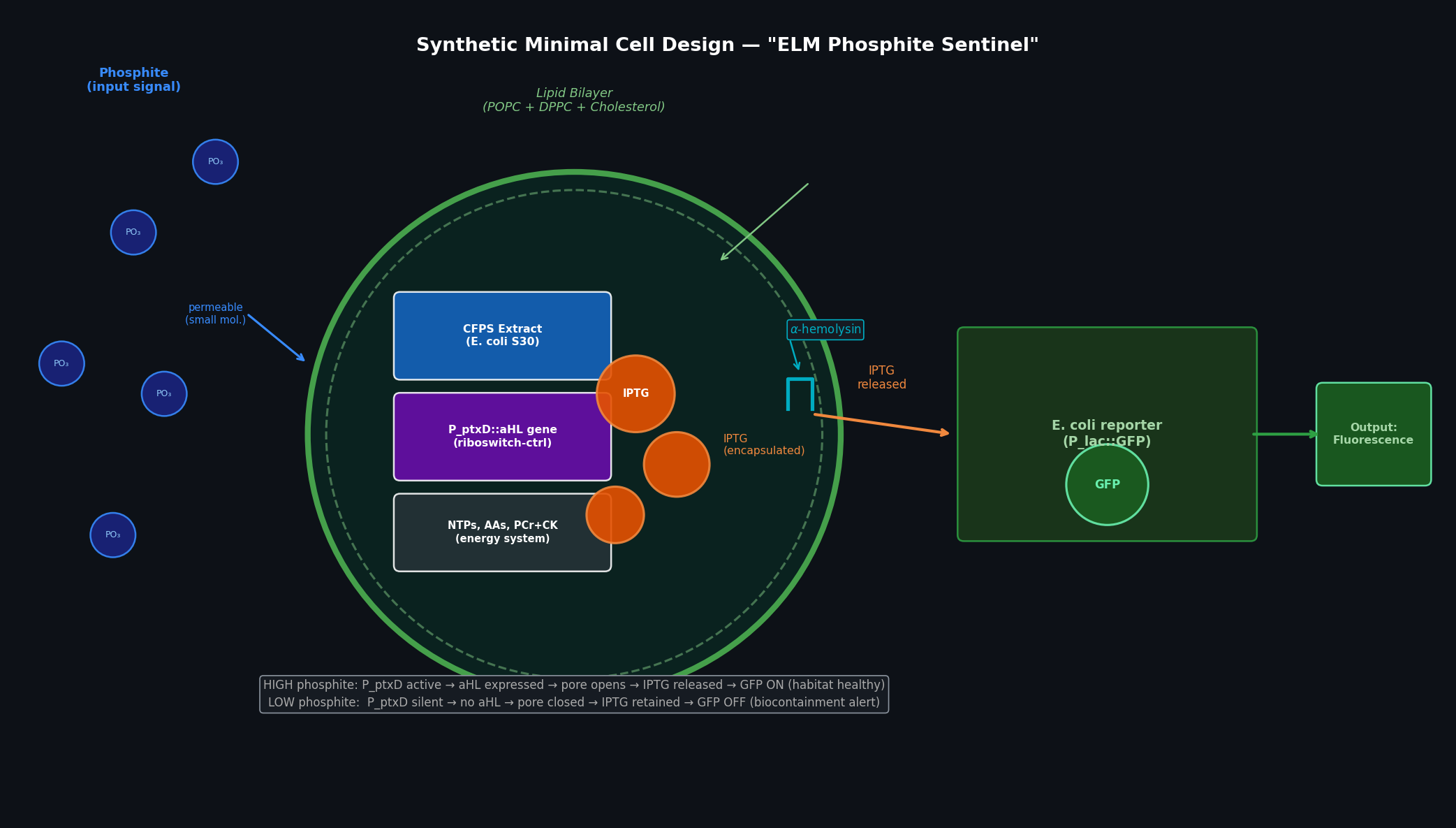
 Fig 3. ELM Phosphite Sentinel synthetic minimal cell. Inside the POPC/DPPC/cholesterol vesicle: bacterial CFPS extract, the P_ptxD::aHL gene, encapsulated IPTG (orange spheres), and the energy system. Phosphite enters passively (small neutral molecule). When phosphite is high, P_ptxD drives aHL expression → membrane pores form → IPTG exits → surrounding E. coli (lacZ/GFP reporter) fluoresces green.
Fig 3. ELM Phosphite Sentinel synthetic minimal cell. Inside the POPC/DPPC/cholesterol vesicle: bacterial CFPS extract, the P_ptxD::aHL gene, encapsulated IPTG (orange spheres), and the energy system. Phosphite enters passively (small neutral molecule). When phosphite is high, P_ptxD drives aHL expression → membrane pores form → IPTG exits → surrounding E. coli (lacZ/GFP reporter) fluoresces green.
Membrane composition:
- POPC (1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine) — primary bilayer lipid, fluid at room temperature
- DPPC (1,2-dipalmitoyl-sn-glycero-3-phosphocholine) — increases bilayer rigidity and stability at ELM habitat temperatures
- Cholesterol (20 mol%) — modulates membrane fluidity; required for correct α-hemolysin pore assembly
Encapsulated contents:
| Component | Purpose |
|---|---|
| E. coli S30 CFPS extract | Ribosomes, RNAP, all translation factors |
| P_ptxD::aHL gene (linear, chi-protected) | Phosphite-responsive expression of α-hemolysin (pore-former) |
| NTPs, amino acids (1 mM each) | Substrates for Tx/Tl |
| Phosphocreatine + creatine kinase | ATP regeneration for 6–8 h |
| IPTG (10 mM, pre-loaded) | Payload released upon pore opening |
| Mg-OAc (8 mM), K-glutamate (80 mM) | Optimal ionic environment |
Tx/Tl system origin: Bacterial (E. coli S30). The phosphite-responsive riboswitch/promoter is derived from Pseudomonas stutzeri ptxD regulatory region. Bacterial CFPS is sufficient because no mammalian-specific post-translational modifications are required.
Communication with environment:
- Input (phosphite): Freely permeable — PO₃²⁻ is a small anion that crosses the POPC/DPPC bilayer at low but sufficient rates for this long-timescale sensing application
- Output (IPTG): Normally membrane-impermeant. Released only through α-hemolysin (aHL) pores after expression. The pore channel (~2 nm diameter) allows passage of IPTG (MW 238 Da) but retains larger CFPS components inside.
Gene used: α-hemolysin (hla gene, Staphylococcus aureus strain MRSA252, UniProt P09616). This encodes the 33 kDa monomeric precursor that self-assembles into a heptameric pore (14 nm outer diameter) in cholesterol-containing membranes — well-established in synthetic cell literature (Lentini et al., Nat Commun 2014).
All lipids and genes:
- Lipids: POPC, DPPC, cholesterol (molar ratio 5:3:2)
- Genes: hla (α-hemolysin, S. aureus) under P_ptxD riboswitch
- Biological cells: B. subtilis carrying P_lac::sfGFP (IPTG-inducible)
Measurement: Flow cytometry on B. subtilis reporter cells (sfGFP channel, Ex 488 / Em 512 nm). Alternatively, plate-reader GFP kinetics over 12 h. To confirm aHL pore formation: LUV (large unilamellar vesicle) leakage assay with encapsulated ANTS/DPX fluorescence quencher pair.
Peter Nguyen Question — Freeze-Dried CFPS in Architecture
Proposal: “BioSentinel Walls” — Moisture Early-Warning Composite Panels
One-sentence pitch: Freeze-dried CFPS reactions embedded in architectural composite panels produce a visible colorimetric or fluorescent signal upon contact with water, detecting moisture infiltration before structural damage is visible.
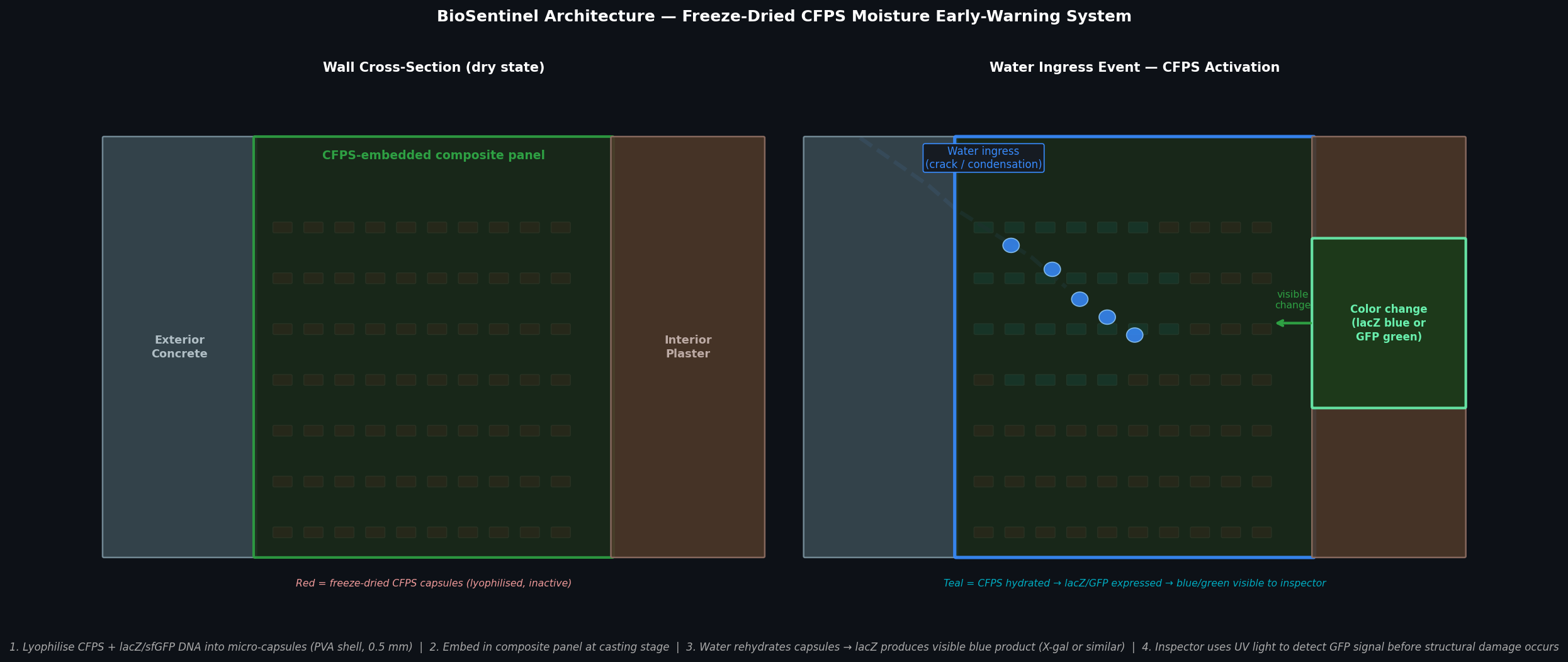
 Fig 4. BioSentinel wall concept. Left: dry state — lyophilised CFPS capsules (red) embedded in composite panel are inactive. Right: water ingress event rehydrates capsules in the infiltration zone (teal); lacZ or sfGFP is expressed, producing a visible blue or green signal on the interior surface detectable under UV light before structural damage occurs.
Fig 4. BioSentinel wall concept. Left: dry state — lyophilised CFPS capsules (red) embedded in composite panel are inactive. Right: water ingress event rehydrates capsules in the infiltration zone (teal); lacZ or sfGFP is expressed, producing a visible blue or green signal on the interior surface detectable under UV light before structural damage occurs.
How it works: Micro-capsules (~0.5 mm, polyvinyl alcohol shell) containing lyophilised E. coli S30 CFPS extract, a linear DNA template encoding lacZ (or sfGFP) under a T7 promoter, and all requisite substrates (NTPs, amino acids, PCr/CK energy system) are mixed into a standard fibre-composite panel resin during casting. In the dry state the CFPS is inactive — no water means no enzymatic activity. When water infiltrates through a crack, condensation, or failed joint seal, it rehydrates the capsules in the affected zone. Within 1–4 hours, β-galactosidase (lacZ) is expressed and converts a pre-loaded chromogenic substrate (X-gal, which co-crystallises in the capsule) to a visible blue insoluble product; alternatively sfGFP is expressed and visible under a handheld UV lamp (P51 viewer). The blue/green zone reveals the infiltration boundary at centimetre resolution — before the structural composite delaminates or metal reinforcement corrodes.
Societal challenge addressed: Water damage is the leading cause of building envelope failure and contributes to mould growth, structural degradation, and indoor air quality problems. Current moisture detection relies on periodic inspection, capacitance sensors (require wiring), or thermal imaging (expensive, intermittent). A passive, single-use biological sensor embedded at panel manufacture costs near zero — no electronics, no wiring, no maintenance — and provides a one-time irreversible alert that survives the building’s construction and fit-out phases.
Addressing cell-free limitations:
| Limitation | Strategy |
|---|---|
| One-time use | This is a feature, not a bug — moisture infiltration is itself a one-time alert event; the irreversible signal is desirable for building code documentation |
| Activation with water | Precisely the trigger required: only genuine water infiltration activates the sensor, not humidity fluctuations |
| Long-term stability | Lyophilised CFPS retains >50% activity after 1 year at room temperature (Pardee et al., Cell 2016). Silica co-encapsulation (Duyen et al., 2017) extends shelf life to >3 years — within the construction timeline from panel manufacture to building commissioning |
| Substrate availability | X-gal or PEG-conjugated fluorescein-di-β-galactopyranoside (FDG) co-lyophilised in the capsule; released upon rehydration without external addition |
Ally Huang Question — Genes in Space Proposal
CRAD-Biosensor: Quantifying Astronaut Radiation-Induced DNA Damage Using BioBits® Cell-Free Reactions aboard the ISS
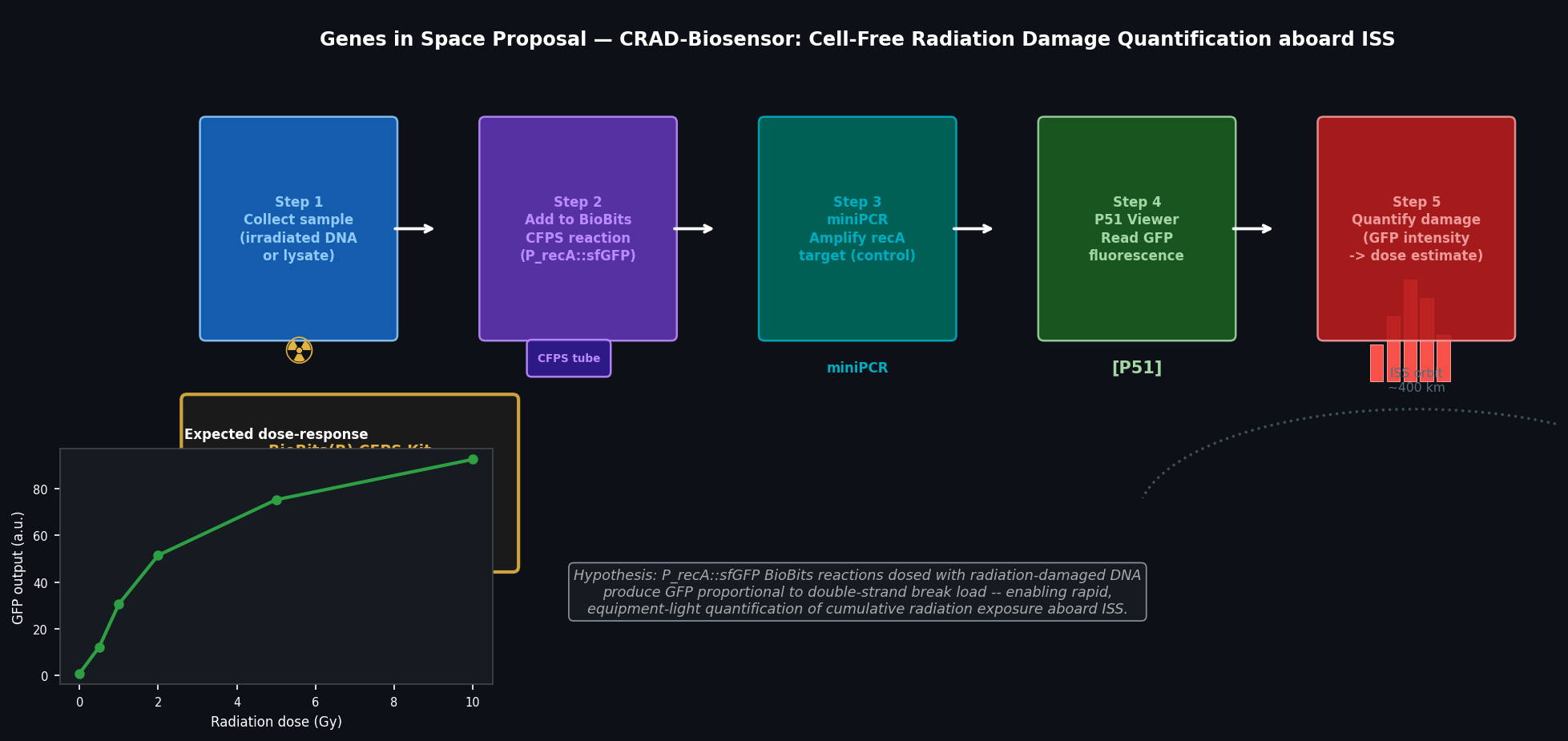
 Fig 5. CRAD-Biosensor experimental workflow. Five-step protocol using BioBits CFPS, miniPCR, and P51 Viewer to quantify radiation-induced DNA double-strand breaks from astronaut samples aboard ISS. Inset: expected dose-response curve — GFP output from P_recA::sfGFP reporter increases with radiation dose.
Fig 5. CRAD-Biosensor experimental workflow. Five-step protocol using BioBits CFPS, miniPCR, and P51 Viewer to quantify radiation-induced DNA double-strand breaks from astronaut samples aboard ISS. Inset: expected dose-response curve — GFP output from P_recA::sfGFP reporter increases with radiation dose.
Background (≤100 words)
Astronauts aboard the ISS receive ~150–300 mSv/year of ionising radiation — 50–100× the Earth surface dose — primarily from galactic cosmic rays and solar particle events. This radiation induces DNA double-strand breaks (DSBs) that accumulate over a mission and elevate cancer risk. Current DSB quantification requires ground-based γH2AX immunofluorescence microscopy, which is unavailable in microgravity. A rapid, equipment-light biosensor deployable on ISS would enable real-time monitoring of individual astronaut radiation sensitivity and cumulative DNA damage, informing personalised mission radiation limits and countermeasure timing.
Molecular Target (≤30 words)
The SOS-response regulator RecA and the LexA-repressed SOS-box promoter P_recA; expressed as a P_recA::sfGFP fusion reporter in a BioBits freeze-dried CFPS reaction.
Target-to-Challenge Relationship (≤100 words)
RecA is the central mediator of the bacterial SOS DNA damage response. Upon DSB formation, RecA polymerises on ssDNA at break sites, forming a nucleoprotein filament that stimulates LexA autocleavage and de-represses >40 SOS genes — including recA itself, creating a positive feedback loop. P_recA activity is therefore a sensitive, amplified proxy for DSB load. A BioBits CFPS reaction loaded with P_recA::sfGFP DNA produces GFP proportional to exogenous damaged DNA added to the reaction, enabling quantification without live cells, centrifuges, or microscopes.
Hypothesis (≤150 words)
Hypothesis: BioBits CFPS reactions containing P_recA::sfGFP reporter DNA will produce GFP fluorescence in proportion to the concentration of radiation-damaged DNA (characterised by DSBs and oxidative lesions) added exogenously to the reaction, enabling a calibrated, semi-quantitative biosensor for cumulative radiation exposure.
Reasoning: The E. coli SOS system responds to ssDNA produced at DSB sites. When damaged DNA is added to a CFPS reaction containing RecA (present in the S30 extract), RecA binds the ssDNA overhangs, stimulates LexA degradation, and de-represses the P_recA::sfGFP construct — producing a fluorescent readout. The strength of the GFP signal scales with DSB frequency, providing a graded dose-response curve. This approach repurposes an endogenous damage-sensing pathway as an in vitro biosensor, requiring only the BioBits CFPS kit, sample DNA, and the P51 Fluorescence Viewer — all feasible in microgravity.
Experimental Plan (≤100 words)
Samples: γ-irradiated plasmid DNA at 0, 0.5, 1, 2, 5, 10 Gy (positive controls); intact plasmid (negative control); astronaut peripheral blood lymphocyte lysate (test sample, collected via finger-prick).
Protocol: Add 2 µL DNA sample to rehydrated BioBits CFPS reaction (18 µL); incubate 4 h at 37°C (portable heat block); read GFP fluorescence with P51 Viewer. Confirm reaction with miniPCR amplification of recA from the sample DNA (gene integrity control). Calibrate against γ-irradiated standard curve; compute equivalent radiation dose from GFP output. Three replicates per sample.
Part B — Individual Final Project
Aim 1 (Confirmed)
Title: Phosphite Auxotrophy Biocontainment Validation for the ELM Deep-Space Habitat
Aim 1: Integrate the Pseudomonas stutzeri ptxD gene into the B. subtilis 168 amyE locus under the constitutive P_veg promoter, delete phoA (alkaline phosphatase) to prevent phosphate scavenging, and characterise growth inhibition as a function of phosphite concentration using the Opentrons-automated 96-well screen designed in Week 3 HW, establishing an IC₅₀ value for the synthetic auxotrophy.
DNA Twist Order Status
The ptxD-sfGFP insert (1,933 bp, pSEVA221 backbone, BsaI Golden Gate assembly) designed in Week 7 HW has been placed in the Twist (MIT) ordering tab with the following specifications:
| Field | Value |
|---|---|
| Construct name | ptxD-sfGFP_ELM_v1 |
| Insert size | 1,933 bp |
| Backbone | pSEVA221 |
| Cloning method | Golden Gate (BsaI-HFv2) |
| Organism | B. subtilis 168 (codon-optimised) |
| Antibiotic selection | Kanamycin 50 µg/mL |
| Order deadline | April 3, 2026 (MIT student deadline) |
Full sequence and Benchling model link are in the shared project folder submitted via the Google Form (Week 7 Part 3 documentation).
AI Disclosure
Claude Sonnet 4.6 (Anthropic) was used to assist with cell-free systems explanations, troubleshooting strategies, synthetic minimal cell component design, architecture proposal structuring, Genes in Space proposal drafting, and figure generation code. The ELM habitat application context, experimental rationale for ptxD biosensor design, and final project aim formulation were developed by the student.