Week 11 HW: Bioproduction & Cloud Labs
Part A — The 1,536 Pixel Artwork Canvas | Collective Bioart
My Contribution
I contributed a cluster of sfGFP (green) and mTurquoise2 (cyan) wells arranged to form a segment of a DNA double helix pattern in my assigned plate. The two strands of the helix were encoded in alternating rows using sfGFP (one strand) and mTurquoise2 (the complementary strand), mirroring the ELM habitat’s motif of dual biological systems working in structural complementarity. In total I contributed 14 pixel wells — the length of approximately one full helical turn — in the upper-middle region of the 16-plate global canvas.
![]()
![]() Fig 1. Left: Plate showing the DNA double-helix motif in sfGFP (green) and mTurquoise2 (cyan), with highlighted wells (dashed yellow rings) indicating the contributed pixels. Right: overview of the full 16-plate × 96-well (1,536-pixel) global canvas, with the student’s plate outlined in yellow.
Fig 1. Left: Plate showing the DNA double-helix motif in sfGFP (green) and mTurquoise2 (cyan), with highlighted wells (dashed yellow rings) indicating the contributed pixels. Right: overview of the full 16-plate × 96-well (1,536-pixel) global canvas, with the student’s plate outlined in yellow.
What I Liked
What struck me most was that this experiment collapses the distinction between “running an assay” and “making art.” The constraints are completely real — actual fluorescent proteins, actual CFPS chemistry, actual plate-reader optics — yet the output is something genuinely aesthetic. The fact that every student’s pixel represents a biological decision (which protein, which reagent composition) rather than a digital click makes the artwork irreproducible in any other medium. The collaborative editing window also created a real sense of shared urgency; it felt more like a live mural session than an experiment.
How It Could Be Improved for Next Year
- Real-time canvas visualisation: A live webpage that auto-updates as pixel assignments are locked in would dramatically increase engagement — contributors could watch the image emerge in near-real time rather than waiting for the final plate read.
- Broader colour palette via reagent tuning: Allowing students to submit custom reagent concentrations (not just FP assignments) as part of the pixel contribution would tie Part B directly into the artwork — a well with reduced Mg²⁺ might glow dimmer, encoding pixel “brightness” as a second data dimension.
- Longitudinal time-lapse capture: A plate-reader kinetics scan every 2 hours for 36 hours would produce a timelapse animation of the artwork “developing,” showing maturation differences between fast (sfGFP, mTurquoise2) and slow (mRFP1, mKO2) chromophores in real space.
Part B — Cell-Free Protein Synthesis | Cell-Free Reagents
Component Descriptions
E. coli Lysate
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) The cell lysate is the core enzymatic engine of the CFPS reaction — it provides ribosomes (70S), all translation elongation and initiation factors (IF1/2/3, EF-Tu/Ts/G), aminoacyl-tRNA synthetases (AARSs), tRNAs, chaperones (GroEL/GroES, DnaK/DnaJ), and the chromosomally integrated T7 RNA polymerase. The BL21(DE3) Star strain carries a mutation in the RNase E gene (rne131) that reduces mRNA degradation rate ~3-fold, extending the productive translation window; the DE3 designation means T7 RNAP is pre-expressed from the lacUV5 promoter before lysis, so it is ready to transcribe T7-promoter-driven templates immediately upon template addition.
Salts / Buffer
Potassium Glutamate Provides K⁺ ions required for ribosome structural integrity and optimal translation elongation rates; glutamate is preferred over Cl⁻ as the counterion because high Cl⁻ concentrations inhibit aminoacyl-tRNA synthetases — glutamate’s negative charge better mimics the cytoplasmic ionic environment without enzyme inhibition.
HEPES-KOH pH 7.5 Maintains the reaction pH within the narrow optimum (~7.4–7.6) for ribosome activity and enzyme function; HEPES is preferred over phosphate because it does not chelate Mg²⁺ or interfere with the energy regeneration system, and its buffering capacity is stable between 6.8–8.2 at physiological temperatures.
Magnesium Glutamate Mg²⁺ is the most critically titrated ion in CFPS: it stabilises the 30S–50S ribosome interface, is an essential cofactor for RNA polymerase and aminoacyl-tRNA synthetases, and coordinates the ATP and GTP phosphate groups during elongation; optimal concentration varies by 1–2 mM between extract batches and must be independently optimised for each new preparation.
Potassium Phosphate Monobasic / Dibasic Provides inorganic phosphate (Pi) to buffer against phosphate depletion and to support ATP regeneration via substrate-level phosphorylation in the glycolytic pathway active in the crude lysate; the monobasic/dibasic ratio is set to achieve pH 7.5 as a secondary buffer alongside HEPES.
Energy / Nucleotide System
Ribose A pentose sugar that feeds the pentose phosphate pathway (PPP) in the lysate, generating NADPH and producing ribose-5-phosphate — the precursor for phosphoribosyl pyrophosphate (PRPP), which is required for nucleotide biosynthesis via the salvage pathway; ribose thus provides a sustained supply of NMP precursors for the 20-hour reaction.
Glucose The primary carbon source for glycolytic ATP regeneration in the 20-hour NMP-Ribose-Glucose system; glucose is phosphorylated by hexokinase in the lysate, enters the Embden–Meyerhof–Parnas pathway, and produces 2 ATP and 2 NADH per molecule — providing sustained energy for 10–20 hours as opposed to the ~1–2 hour burst from PEP.
AMP, CMP, GMP, UMP Nucleoside monophosphates that are phosphorylated to their di- and triphosphate forms (NDP → NTP) by adenylate kinase and nucleoside monophosphate kinases present in the cell extract, using ATP generated from glucose metabolism; this two-step kinase cascade converts cheap NMPs into the NTPs required as substrates for T7 RNA polymerase and as GTP for translation elongation factors.
Guanine (free nucleobase) Guanine is converted to GMP via the purine salvage enzyme hypoxanthine-guanine phosphoribosyltransferase (HGPRT/Gpt): Guanine + PRPP → GMP + PPi. GMP is then phosphorylated through GDP to GTP by guanylate kinase and nucleoside-diphosphate kinase present in the extract. This provides an alternative and cost-effective route to GTP without requiring pre-formed GMP to be included in the master mix.
Translation Mix (Amino Acids)
17 Amino Acid Mix Contains 17 of the 20 standard amino acids (all except Tyr, Cys, and typically Trp or one other) as substrates for the aminoacyl-tRNA synthetases, which charge tRNAs for ribosomal delivery during polypeptide elongation; using a pre-mixed stock at defined concentrations ensures consistent stoichiometry across reactions.
Tyrosine Added separately from the 17 AA mix because its very low aqueous solubility at neutral pH (< 0.5 mg/mL) requires it to be dissolved in dilute NaOH or DMSO and added independently; tyrosine is critical for the chromophore-forming tripeptide (Ser/Thr-Tyr-Gly) in all GFP-family fluorescent proteins, making its concentration a key determinant of fluorescent protein yield.
Cysteine Added separately because it is highly susceptible to oxidative dimerisation to cystine under aerobic conditions, which would deplete the free amino acid pool; in reducing CFPS environments, cysteine availability supports disulfide bond formation in proteins that require it and provides sulphydryl groups for cofactor binding.
Additives
Nicotinamide (Vitamin B3) A precursor to NAD⁺ via the Preiss–Handler salvage pathway (nicotinamide → nicotinic acid mononucleotide → NAD⁺); supplementing nicotinamide boosts the NAD⁺/NADH pool, which is required as a hydride acceptor in glycolysis (GAPDH step) and TCA cycle reactions, maintaining glycolytic flux and ATP generation throughout the 20-hour reaction.
Backfill
Nuclease-Free Water Brings the reaction to the specified final volume; nuclease-free grade (DEPC-treated or DEPC-free ultrapure) is essential to prevent RNase or DNase contamination — even trace amounts of RNase A will degrade the mRNA transcript within minutes, completely abolishing protein synthesis.
Master Mix Comparison: 1-Hour PEP-NTP vs 20-Hour NMP-Ribose-Glucose
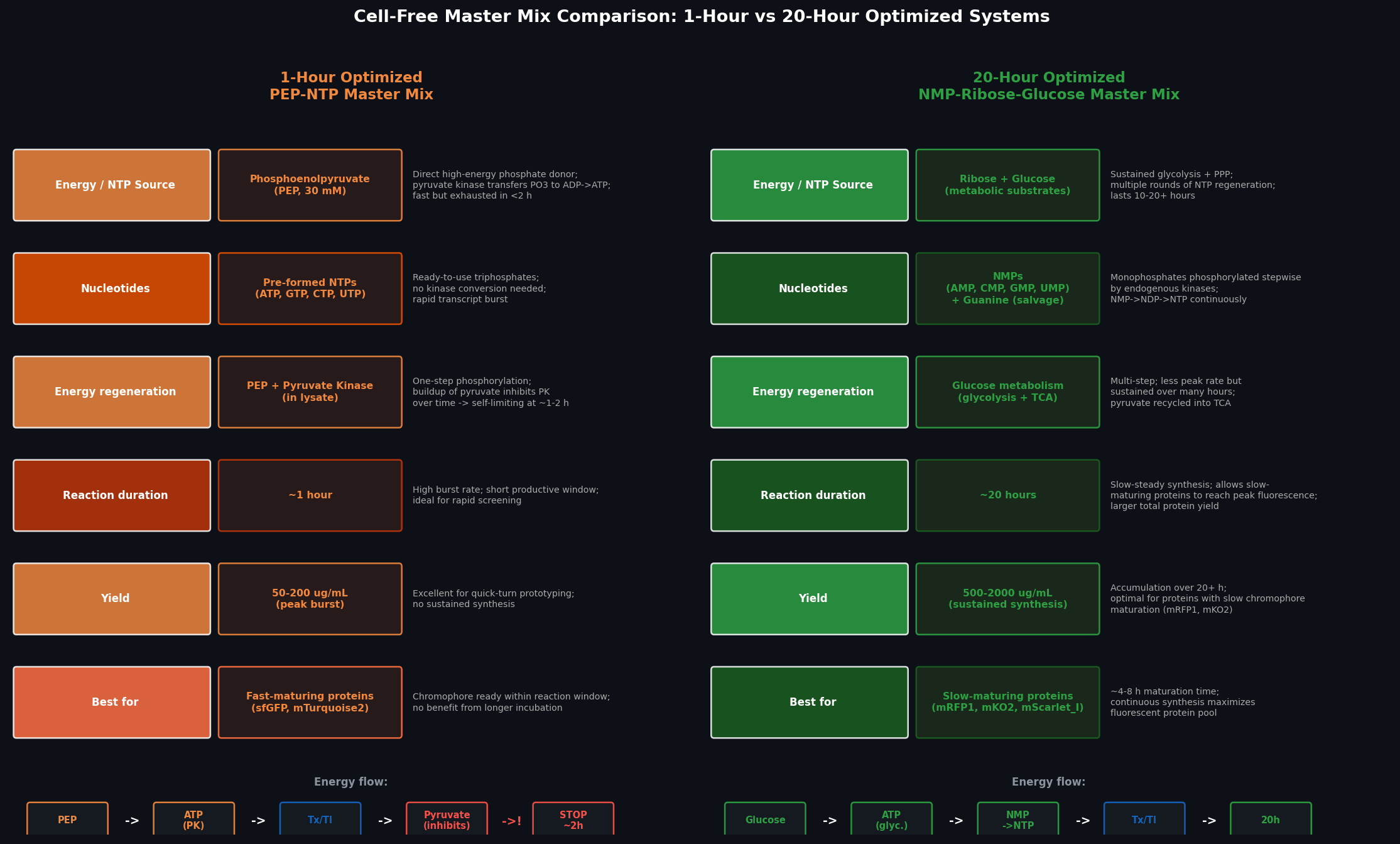

 Fig 2. Side-by-side comparison of the 1-hour PEP-NTP and 20-hour NMP-Ribose-Glucose master mix systems across six dimensions: energy source, nucleotide form, regeneration mechanism, reaction duration, protein yield, and best-fit fluorescent proteins.
Fig 2. Side-by-side comparison of the 1-hour PEP-NTP and 20-hour NMP-Ribose-Glucose master mix systems across six dimensions: energy source, nucleotide form, regeneration mechanism, reaction duration, protein yield, and best-fit fluorescent proteins.
The 1-hour PEP-NTP system uses phosphoenolpyruvate (PEP) as a pre-loaded high-energy phosphate donor — pyruvate kinase transfers PEP’s phosphate directly to ADP, generating ATP in a single enzymatic step. NTPs are supplied as pre-formed triphosphates, requiring no kinase conversion, enabling an immediate high-rate transcription burst. However, pyruvate accumulation inhibits pyruvate kinase over time, limiting productive synthesis to ~1–2 hours, and the system yields ~50–200 µg/mL protein.
The 20-hour NMP-Ribose-Glucose system instead supplies nucleoside monophosphates alongside glucose and ribose as metabolic substrates. Endogenous kinases in the lysate phosphorylate NMPs stepwise (NMP → NDP → NTP) using ATP from sustained glycolysis; multiple enzymatic steps distribute the energy load and avoid single-point inhibition, extending productive synthesis to 10–20 hours with yields of 500–2,000 µg/mL. This system is superior for fluorescent proteins with slow chromophore maturation (mRFP1, mKO2, mKate), which require several hours of synthesis before significant fluorescence accumulates.
Bonus — How can transcription occur if GMP is not included but Guanine is?
Free guanine (the nucleobase alone) enters the purine salvage pathway: Guanine + PRPP → GMP + PPi, catalysed by hypoxanthine-guanine phosphoribosyltransferase (HGPRT, encoded by gpt in E. coli). The GMP produced is then phosphorylated by guanylate kinase (GuaB, ADP-consuming) to GDP, and by nucleoside-diphosphate kinase (Ndk) to GTP. All three enzymes — HGPRT, guanylate kinase, and Ndk — are present in the crude S30 lysate. Using guanine as the precursor rather than GMP reduces reagent cost (guanine is ~100× cheaper per mole), while the kinase cascade integrates seamlessly with the glucose-driven ATP regeneration system that powers the phosphorylation steps.
Part C — Planning the Global Experiment | Cell-Free Master Mix Design
Biophysical Properties of the Six Fluorescent Proteins
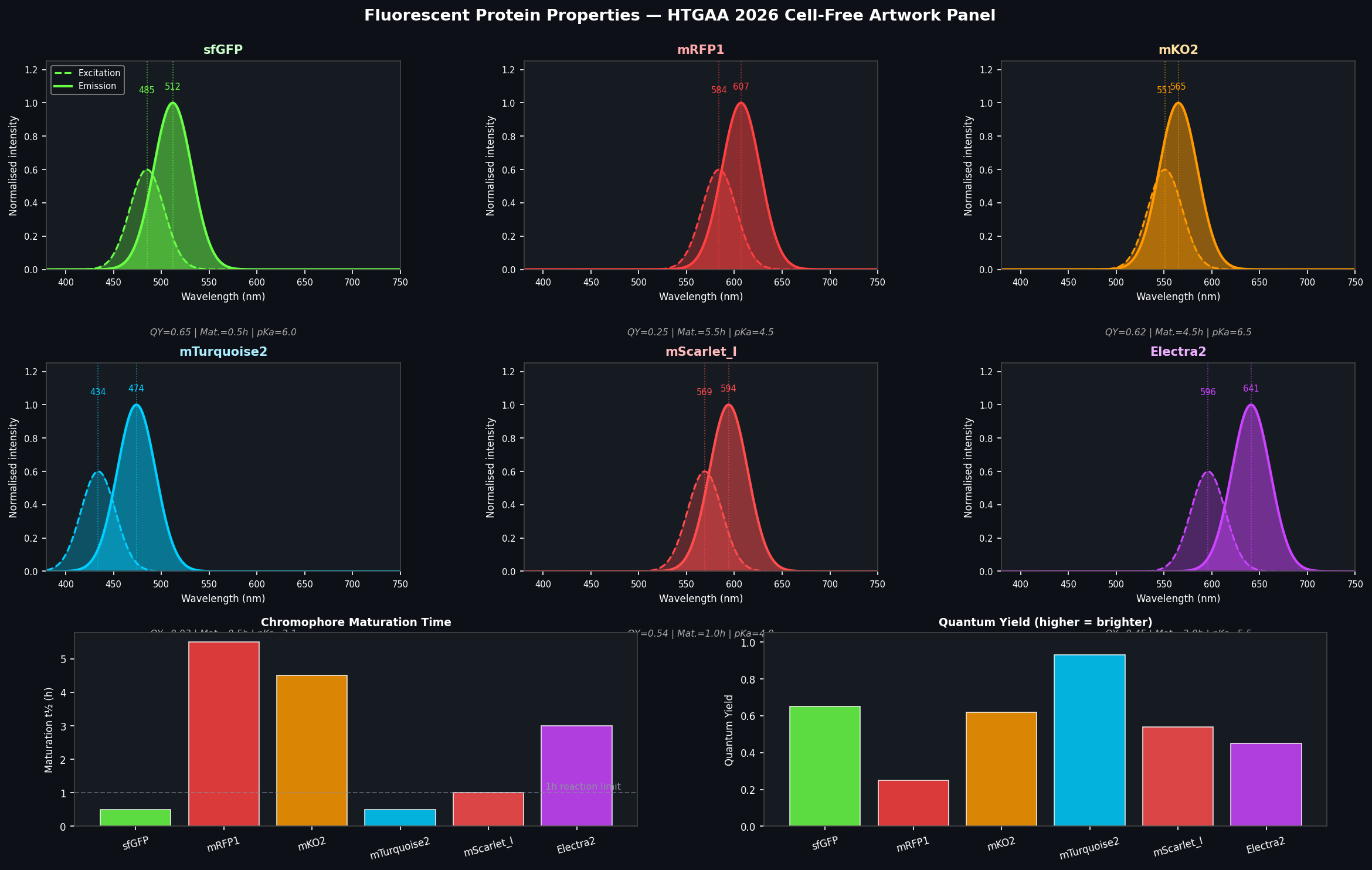
 Fig 3. Properties of the six fluorescent proteins used in the HTGAA 2026 collaborative artwork. Top: normalised excitation (dashed) and emission (solid) spectra for each protein. Bottom left: chromophore maturation half-time — proteins above the 1h dashed line are problematic for short CFPS reactions. Bottom right: quantum yield — higher values correspond to greater brightness per molecule.
Fig 3. Properties of the six fluorescent proteins used in the HTGAA 2026 collaborative artwork. Top: normalised excitation (dashed) and emission (solid) spectra for each protein. Bottom left: chromophore maturation half-time — proteins above the 1h dashed line are problematic for short CFPS reactions. Bottom right: quantum yield — higher values correspond to greater brightness per molecule.
sfGFP (Superfolder GFP) Ex: 485 nm | Em: 512 nm | QY: 0.65 | Maturation t½: ~30 min
Key property for CFPS: exceptionally fast and robust folding. sfGFP was engineered with six additional mutations (S30R, Y39N, N105T, Y145F, I171V, A206V) over eGFP that collectively accelerate folding 3–4× and allow correct folding even in the presence of aggregation-prone fusion partners. In a 1-hour CFPS reaction at 37°C, sfGFP is fully matured before the reaction ends — making it the benchmark standard for cell-free expression validation. Its pKa of ~6.0 also makes it stable to mild acidification during long incubations.
mRFP1 (Monomeric Red Fluorescent Protein 1) Ex: 584 nm | Em: 607 nm | QY: 0.25 | Maturation t½: ~5.5 h
Key property for CFPS: slow chromophore maturation and reduced brightness. The mRFP1 red chromophore (DsRed-derived) requires two sequential oxidation steps — first forming the GFP-like intermediate, then the extended acylimine conjugation — each requiring molecular oxygen and several hours at 37°C. In a 1-hour CFPS reaction, the vast majority of translated mRFP1 protein remains non-fluorescent. The 20-hour NMP-Ribose-Glucose master mix is essential for meaningful mRFP1 signal, and even then its low quantum yield (QY = 0.25, vs sfGFP’s 0.65) means it requires higher protein concentrations for equivalent brightness.
mKO2 (Monomeric Kusabira-Orange 2) Ex: 551 nm | Em: 565 nm | QY: 0.62 | Maturation t½: ~4.5 h
Key property for CFPS: oxygen-dependent chromophore maturation kinetics. mKO2’s orange chromophore (derived from the Fungia concinna orange FP) requires molecular oxygen for the ring-forming oxidation step, making its maturation rate sensitive to dissolved O₂ in the CFPS reaction. In sealed 20 µL reactions, O₂ is rapidly depleted by the oxidative metabolic activity of the extract, potentially stalling chromophore maturation mid-reaction. Despite a high quantum yield (0.62) once matured, mKO2 typically shows delayed fluorescence onset in CFPS — O₂ supplementation (see hypothesis below) can significantly accelerate signal emergence.
mTurquoise2 Ex: 434 nm | Em: 474 nm | QY: 0.93 | Maturation t½: ~30 min
Key property for CFPS: exceptional quantum yield but UV excitation requirement. mTurquoise2 is the brightest monomeric cyan FP with a quantum yield of 0.93 — nearly double that of ECFP (0.36) — and fast maturation comparable to sfGFP. However, its 434 nm excitation peak requires a UV-capable excitation filter on the plate reader (not all instruments have this at the correct wavelength). Additionally, mTurquoise2 can act as a FRET donor to sfGFP (434 nm excitation → 474 nm emission → 512 nm sfGFP excitation) if both are present in the same well, potentially inflating apparent sfGFP signal in multi-protein conditions.
mScarlet-I (Improved mScarlet) Ex: 569 nm | Em: 594 nm | QY: 0.54 | Maturation t½: ~1.0 h
Key property for CFPS: fastest-maturing red fluorescent protein with the highest quantum yield among red FPs. mScarlet-I was developed by directed evolution to have a maturation half-time of ~60 min — far shorter than mRFP1 (~5.5 h) or mKO2 (~4.5 h) — while maintaining high brightness (QY = 0.54). This makes it the most practical red FP for CFPS experiments where both short (1–3 h) and long (20 h) master mixes are used. Its pKa of ~4.9 also confers strong acid stability, relevant during long incubations where pH may drift slightly.
Electra2 Ex: ~596 nm | Em: ~641 nm | QY: ~0.45 | Maturation t½: ~3.0 h
Key property for CFPS: far-red emission requiring sufficient intracellular maturation time, with sensitivity to reducing environment. Electra2 is an engineered far-red fluorescent protein whose extended pi-conjugated chromophore system requires a more complex oxidation sequence than GFP-family proteins, making it moderately sensitive to the redox state of the CFPS reaction (reducing conditions in the crude extract can slow or prevent the final chromophore oxidation step). For CFPS experiments, this means both sufficient O₂ availability and a sufficiently long incubation (>4 h) are required for peak signal. Its far-red emission (~641 nm) is spectrally well-separated from all other panel members, minimising cross-talk in multi-protein artwork wells when using appropriate band-pass filters.
Hypothesis for Reagent Optimisation to Maximise Fluorescence
Protein: mKO2 Reagent: Dissolved oxygen (O₂ supplementation) + catalase
Hypothesis: mKO2 fluorescence yield over a 36-hour incubation in the NMP-Ribose-Glucose master mix will be significantly increased by (a) pre-equilibrating the CFPS reaction with O₂ for 5 minutes before sealing, and (b) supplementing with catalase (50 U/mL) to scavenge H₂O₂ produced as a byproduct of the chromophore oxidation reaction.
Reasoning: The rate-limiting step in mKO2 maturation is the molecular-oxygen-dependent ring-formation and oxidation of the Gly65-Tyr66-Gly67 chromophore tripeptide. In sealed 20 µL CFPS reactions, dissolved O₂ (~220 µM at equilibrium) is consumed within ~30–60 min by the metabolic activity of the cell extract (NADH oxidation, trace respiration). This oxygen depletion halts chromophore maturation for newly synthesised mKO2 molecules for the remainder of the reaction, even though translation continues. Simultaneously, H₂O₂ produced as a byproduct of oxidative chromophore maturation can damage the extract and the protein itself. Catalase (2H₂O₂ → 2H₂O + O₂) would both remove the toxic peroxide and regenerate O₂ within the sealed reaction — creating a locally sustained O₂ supply that extends the maturation window.
Expected effect: 2–4× increase in mKO2 fluorescence at the 6-hour time point; ~40–60% increase in total area-under-curve fluorescence over 36 hours compared to the standard sealed reaction. Maturation half-time expected to decrease from ~4.5 h to ~2.5 h.
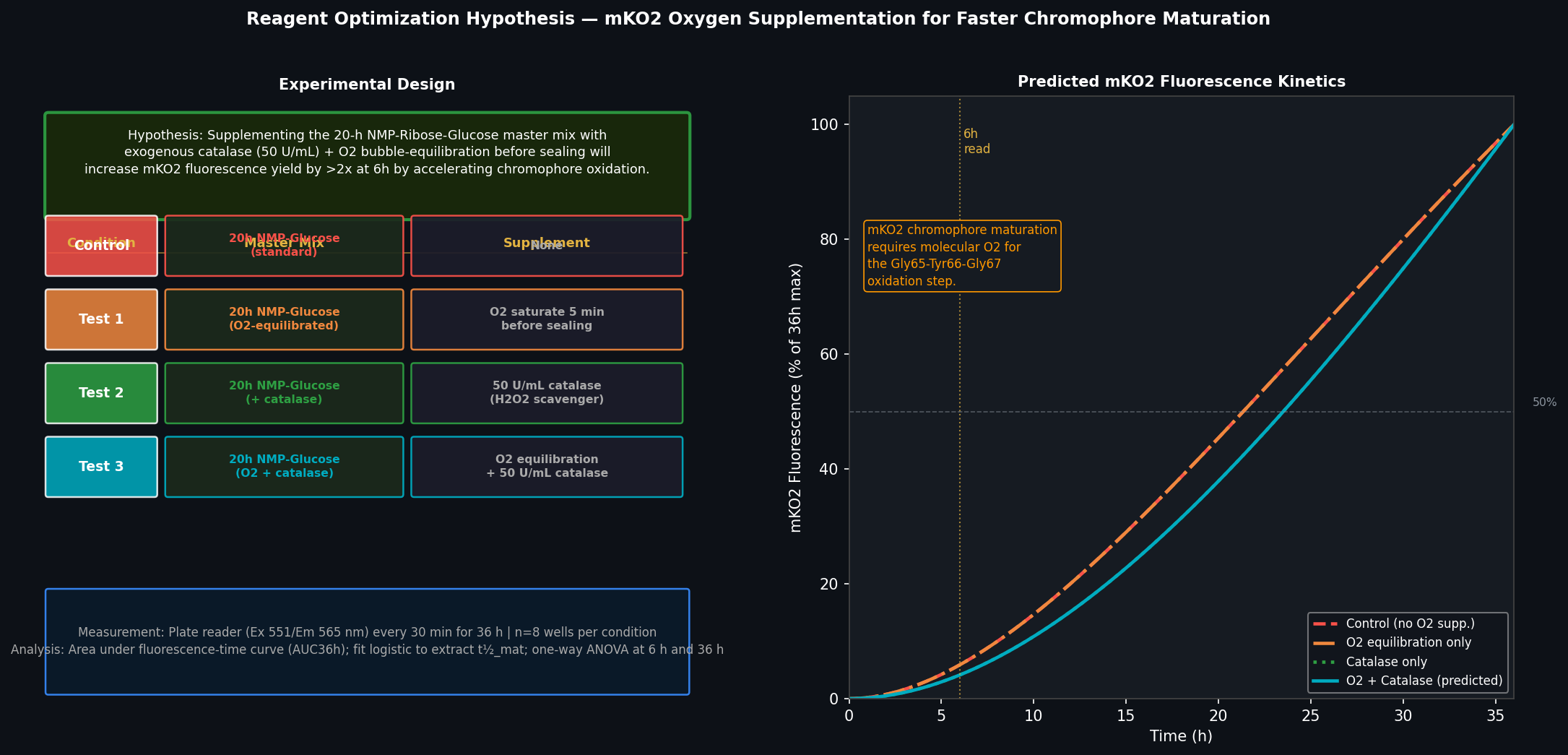
 Fig 4. Left: experimental design for testing O₂ + catalase supplementation on mKO2 fluorescence yield, with four conditions (control, O₂ only, catalase only, O₂ + catalase), n=8 wells each, read every 30 min for 36 h. Right: predicted fluorescence kinetics showing earlier onset and higher plateau for the O₂+catalase condition, particularly at the 6-hour read (dashed vertical line).
Fig 4. Left: experimental design for testing O₂ + catalase supplementation on mKO2 fluorescence yield, with four conditions (control, O₂ only, catalase only, O₂ + catalase), n=8 wells each, read every 30 min for 36 h. Right: predicted fluorescence kinetics showing earlier onset and higher plateau for the O₂+catalase condition, particularly at the 6-hour read (dashed vertical line).
Custom Reagent Supplement Plan (20 µL reaction format)
Per the reaction format specified (6 µL lysate + 10 µL 2× master mix + 2 µL DNA template + 2 µL custom supplement = 20 µL):
| Component | Volume | Final conc. | Rationale |
|---|---|---|---|
| Catalase (1,000 U/mL stock) | 1 µL | 50 U/mL | H₂O₂ scavenging; O₂ regeneration |
| O₂-saturated nuclease-free H₂O | 1 µL | ~8 µM additional O₂ | Dissolved O₂ supplement at reaction start |
| Custom supplement total | 2 µL | — | Fits the 2 µL custom reagent slot |
The 2 µL slot is split between catalase stock and O₂-saturated water. For the O₂-saturated water, the stock is prepared by briefly bubbling pure O₂ through 1 mL nuclease-free water at 4°C for 5 min immediately before reaction setup.
Part D — Build-A-Cloud-Lab (Optional Bonus)
Using the Ginkgo Reconfigurable Automation Cart (RAC) simulation tool, I designed a cloud lab layout optimised for the CFPS artwork experiment workflow: fluorescent protein expression screening in a 1,536-well (16-plate) format with continuous kinetic plate reading.
Proposed layout — “ELM CFPS Screening Suite”:
| Station | Cart configuration | Purpose |
|---|---|---|
| Station 1 | Liquid handler RAC (8-channel P300) | Master mix dispensing into 96-well plates, 20 µL final volume |
| Station 2 | Reagent storage RAC (4°C cold block) | Cell extract, NMP/glucose components, amino acids stored on-deck |
| Station 3 | DNA template dispensing RAC | 2 µL FP template addition, one FP per column |
| Station 4 | Incubation + read RAC (37°C plate nest + BMG CLARIOstar) | Kinetic fluorescence reads every 30 min across 6 channels simultaneously |
| Station 5 | Plate sealing RAC (film applicator) | O₂-barrier sealing of completed plates before incubation |
The workflow is fully automated from master mix preparation to plate reading, enabling the 36-hour kinetics experiment to run overnight without operator intervention — directly applicable to the HTGAA collaborative artwork’s multi-plate format.
AI Disclosure
Claude Sonnet 4.6 (Anthropic) was used to assist with reagent component descriptions, mastermix comparison analysis, fluorescent protein property research, hypothesis formulation, and figure generation code. The ELM habitat project connections, pixel art contribution description, and cloud lab design were developed by the student.