Week 3 Lab: Opentrons Art
Opentrons Artwork: Fluorescent Bacteria Pixel Art
Overview | Objective
In this two-day lab, you’ll program the Opentrons OT-2 pipetting robot to create stunning, glowing designs by depositing genetically engineered E. coli onto black (charcoal) agar plates. These bacteria express fluorescent proteins in vibrant colors, forming “bio-art” that comes to life under UV light. It’s your chance to turn cutting-edge biotech into a canvas for creativity!
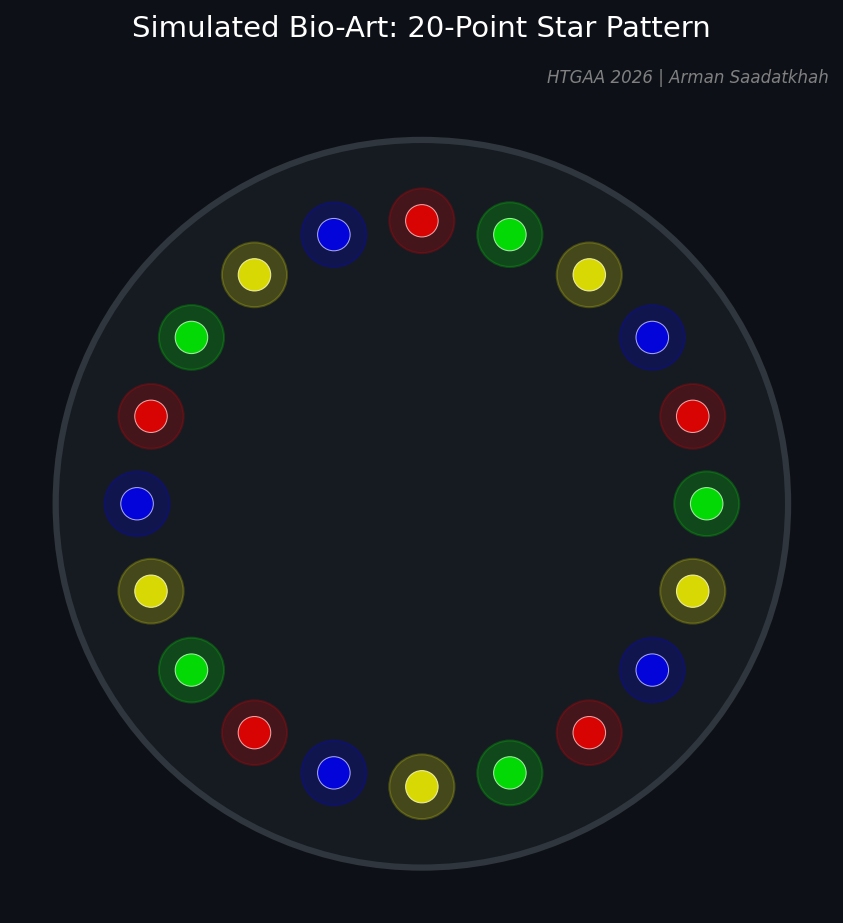
 Fig 1. Simulated 20-point star pattern generated via Python for the Opentrons robot.
Fig 1. Simulated 20-point star pattern generated via Python for the Opentrons robot.
Overview | Concepts Learned & Skills Gained
This week, you will be working with the Opentrons OT-2, a liquid handling robot used in various life-science laboratories. You will learn:
- How to incorporate automation into synthetic biology research.
- How to code a Python script using the Opentrons API.
- How to create agar plates, a basic tool in molecular biology.
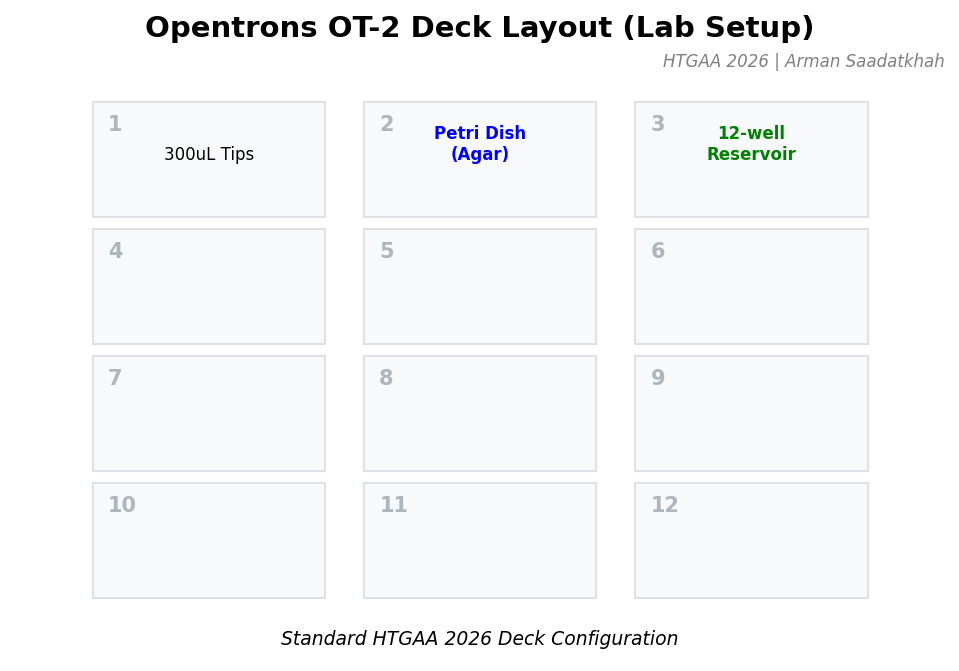

 Fig 2. Standard HTGAA 2026 Deck Configuration for the OT-2 robot.
Fig 2. Standard HTGAA 2026 Deck Configuration for the OT-2 robot.
Pre-Lab | Reading
The “Central Dogma” of Opentrons
Before programming, it’s important to understand the workflow that transforms an idea into a precise robotic procedure.
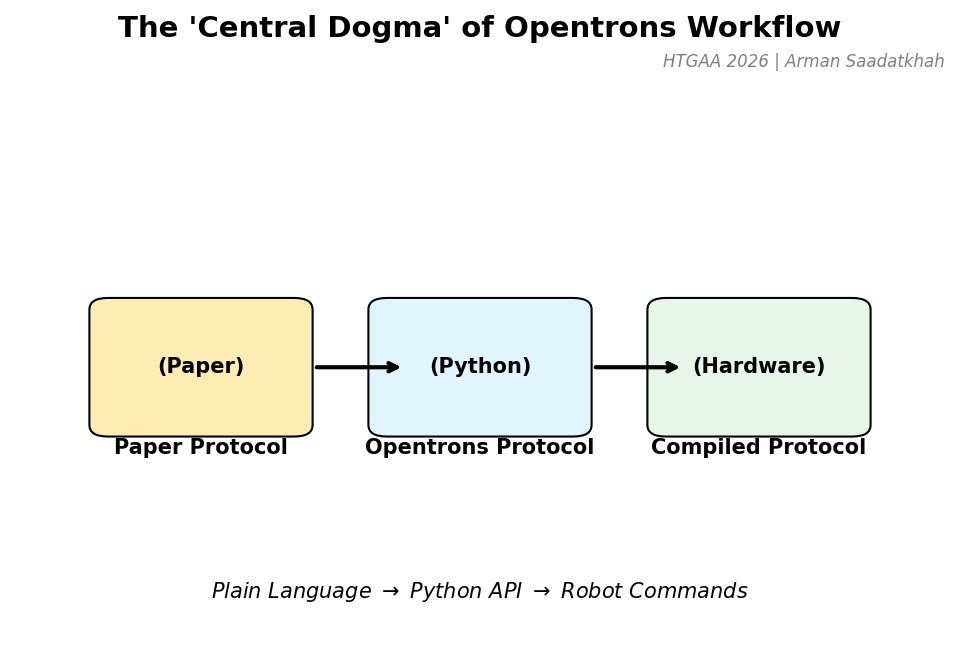
 Fig 3. The transformation from plain language instructions to hardware commands.
Fig 3. The transformation from plain language instructions to hardware commands.
- Paper Protocol: Instructions written in plain language (e.g., “Pipette 100 uL into A1”).
- Opentrons Protocol: The Python script that translates these steps using the Opentrons API.
- Compiled Protocol: The Opentrons App compiles the script into commands controlling the robot hardware.
GFP and Friends: The Science of Glow
Green Fluorescent Protein (GFP) is a protein that glows green when illuminated with UV light. “Fluorescing” involves absorbing light at one wavelength (UV) and re-emitting it at another (Visible Green).
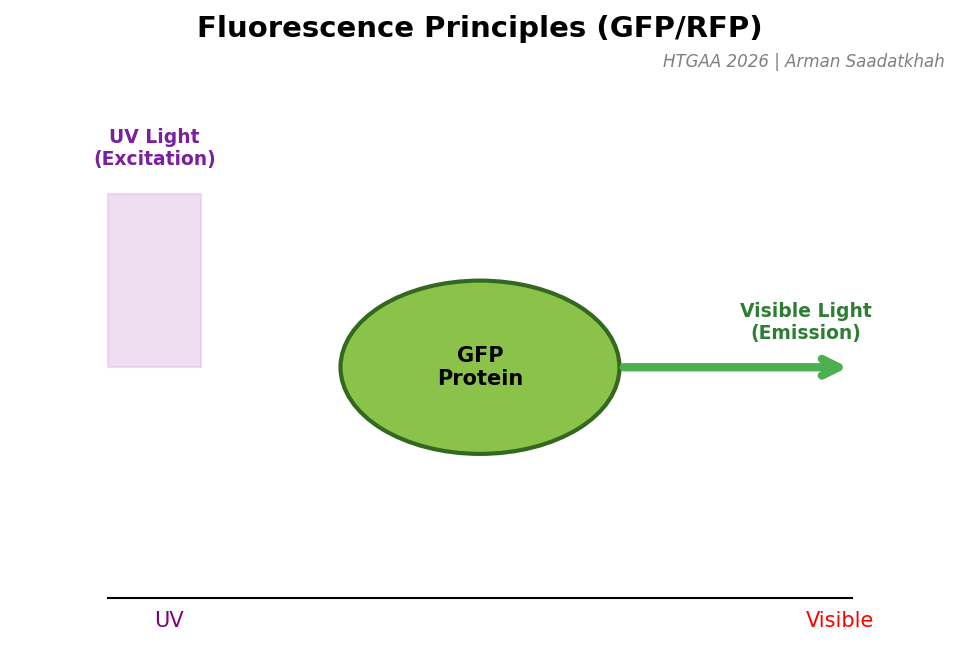
 Fig 4. The mechanism of absorption and emission in fluorescent proteins.
Fig 4. The mechanism of absorption and emission in fluorescent proteins.
For this lab, we use E. coli spliced with R/G/B/C/YFP genes. We mix charcoal powder into the agar to make it black, enhancing the visibility of the glowing designs.
Protocol | Part 1: Fluorescent Bacteria & Black Agar Script
Time Estimate: 2 Hours
Artistic Concept — “ELM Habitat Cross-Section”
My design uses the 96-well plate as a canvas to depict a cross-sectional schematic of the Multi-Trophic Myco-Foundry — the engineered living material (ELM) habitat proposed in Week 1.
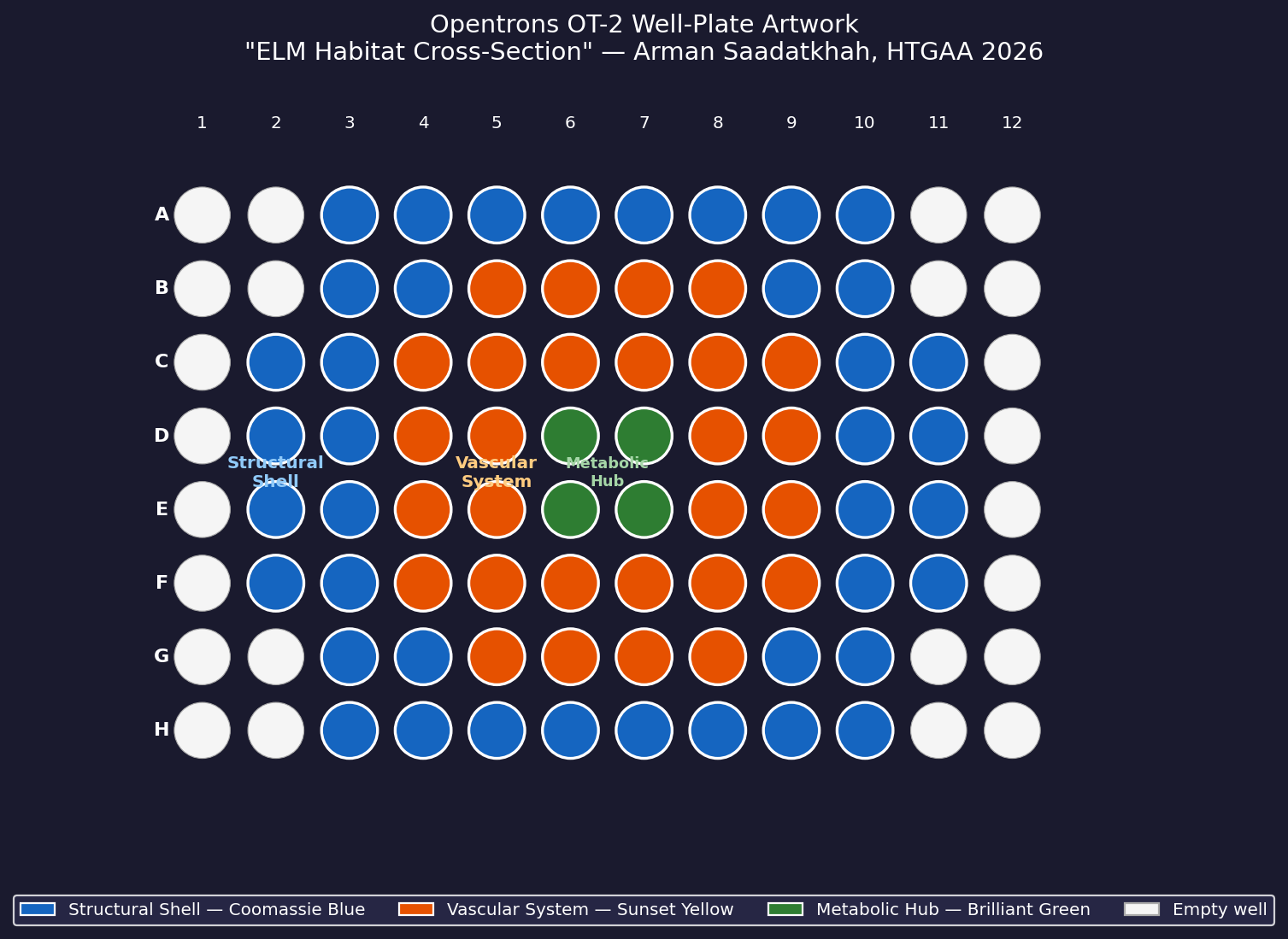
 Fig 5. Well-plate layout for “ELM Habitat Cross-Section.” Blue = Structural Shell, Orange = Vascular System, Green = Metabolic Hub.
Fig 5. Well-plate layout for “ELM Habitat Cross-Section.” Blue = Structural Shell, Orange = Vascular System, Green = Metabolic Hub.
Python Script (Opentrons API v2.14)
The robot run starts without any tips. Fresh tips are used for every color to prevent cross-contamination.
Protocol | Part 2: Automation Plan for Final Project
Goal: Use lab automation to screen and validate the phosphite auxotrophy biocontainment system.
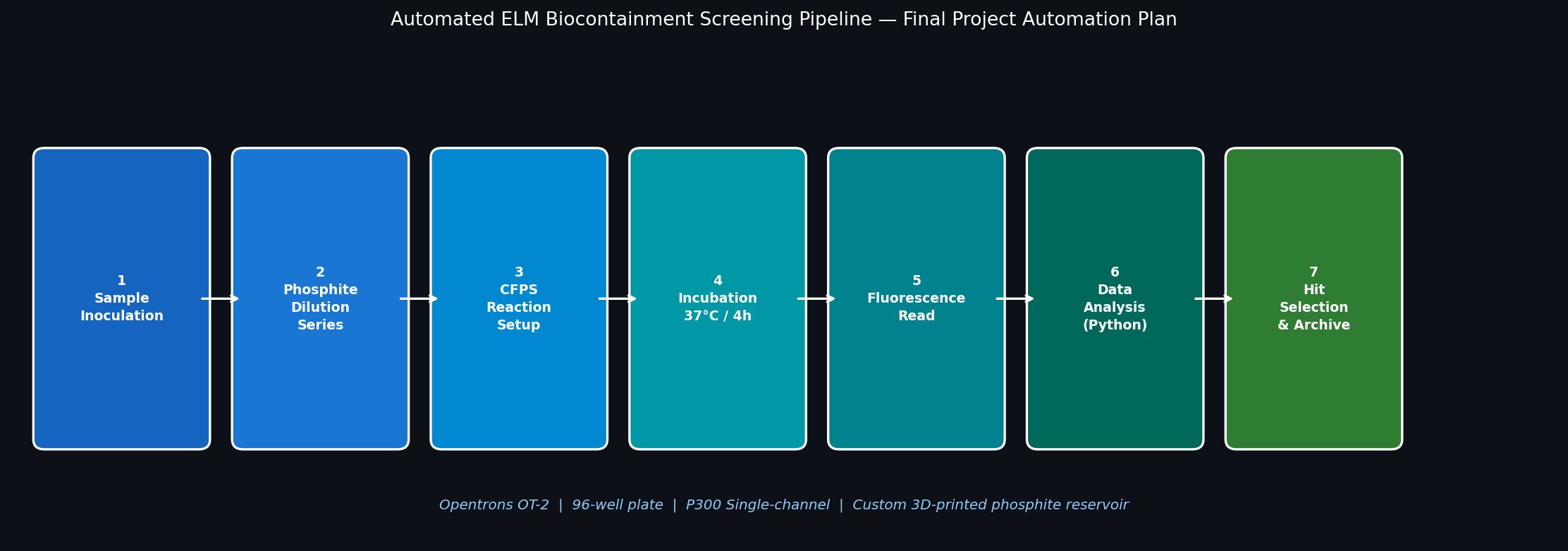
 Fig 6. Seven-step Opentrons automation pipeline for screening the ELM phosphite auxotrophy kill switch.
Fig 6. Seven-step Opentrons automation pipeline for screening the ELM phosphite auxotrophy kill switch.
The OT-2 will run a 12-condition × 8-replicate growth screen, testing the ptxD-based kill switch across a 2-fold phosphite dilution series. This allows for rapid calculation of the IC₅₀ value, essential for the safety case of deployment.
Post-Lab | Troubleshooting & Results
Alice Cai’s final plate (2023) demonstrates the transition from a dark charcoal agar to a vibrant glowing masterpiece under UV light. Simple geometric shapes often yield the most precise results on the OT-2.
HTGAA 2026 | Arman Saadatkhah | Reference: Opentrons OT-2 Documentation