Week 6 Lab: Gibson Assembly
The Chromophore Color Cloning Quest
Overview | Objective
In this lab, you’ll be changing the color-generating chromophore of the purple Acropora millepora chromoprotein (amilCP) to a variety of orange, pink, and blue mutants.
First, we’ll prepare two polymerase chain reactions (PCR) to generate the necessary fragments for a Gibson assembly. Using the amilCP-encoding Addgene mUAV plasmid as a template, we will amplify:
- The Backbone Fragment: Containing the origin of replication, Chloramphenicol resistance, and the promoter/RBS.
- The Insert Fragment: Containing the chromophore region with intentional mutations for color variation.
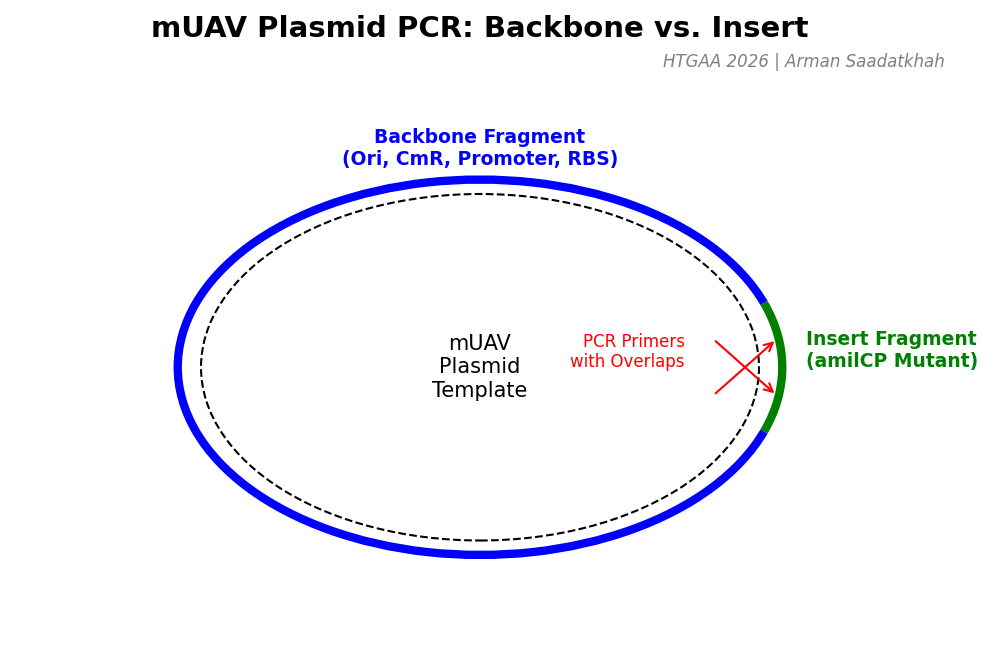
 Fig 1. PCR strategy for generating Backbone and Insert fragments from the mUAV template.
Fig 1. PCR strategy for generating Backbone and Insert fragments from the mUAV template.
Pre-Lab | Concepts
(1) Spectral Engineering of amilCP
The amilCP gene contains a chromophore (CP) region that can be mutated to express different colors. By changing the DNA sequence at the CP site (cagTGTCAGtac), we can engineer proteins that absorb light at different wavelengths.
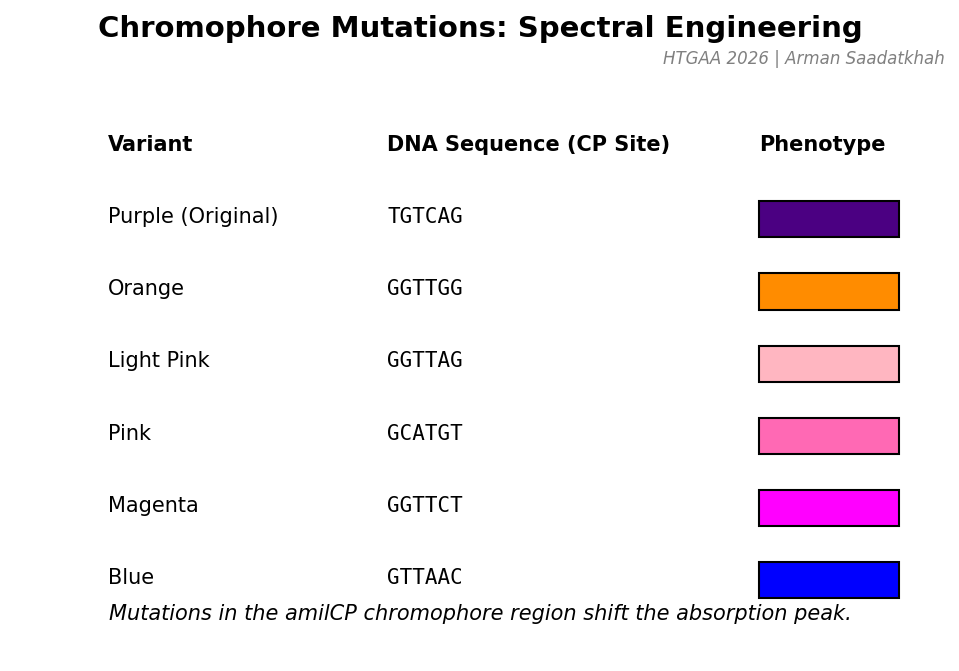

 Fig 2. DNA sequences and predicted phenotypes for various amilCP mutants.
Fig 2. DNA sequences and predicted phenotypes for various amilCP mutants.
(2) Gibson Assembly Mechanism
Gibson Assembly is a “one-pot” isothermal reaction that allows multiple DNA fragments to be joined together. It relies on overlapping sequences (20-40 bp) at the ends of the fragments.
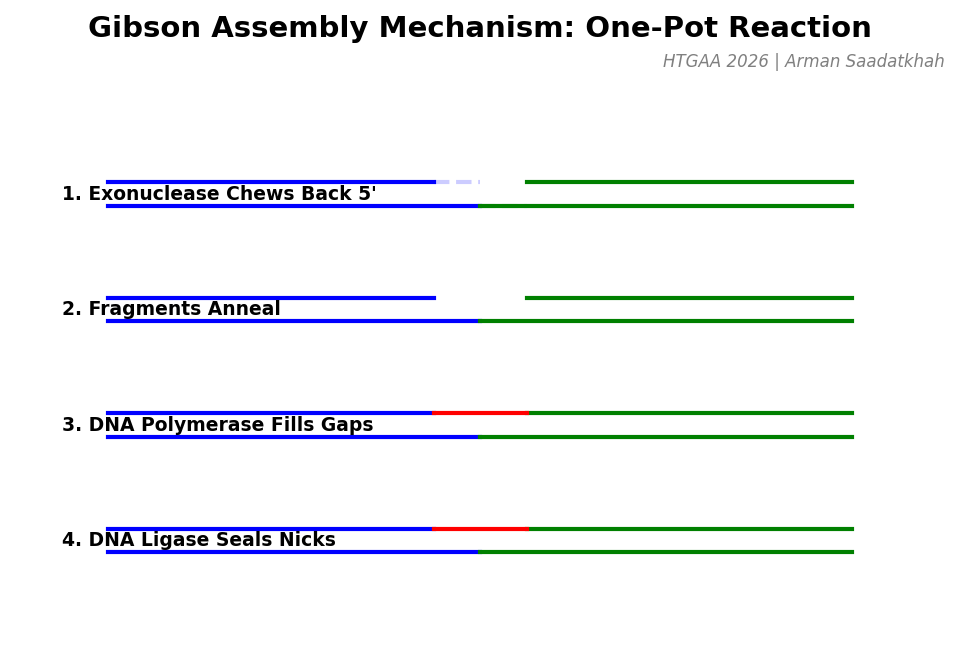

 Fig 3. The four enzymatic steps of Gibson Assembly: Exonuclease chew-back, Annealing, Polymerase fill-in, and Ligase sealing.
Fig 3. The four enzymatic steps of Gibson Assembly: Exonuclease chew-back, Annealing, Polymerase fill-in, and Ligase sealing.
(3) Transformation
Once the plasmid is assembled, it must be introduced into E. coli cells. We use Heat Shock to create temporary pores in the cell membrane, allowing the DNA to enter by diffusion.
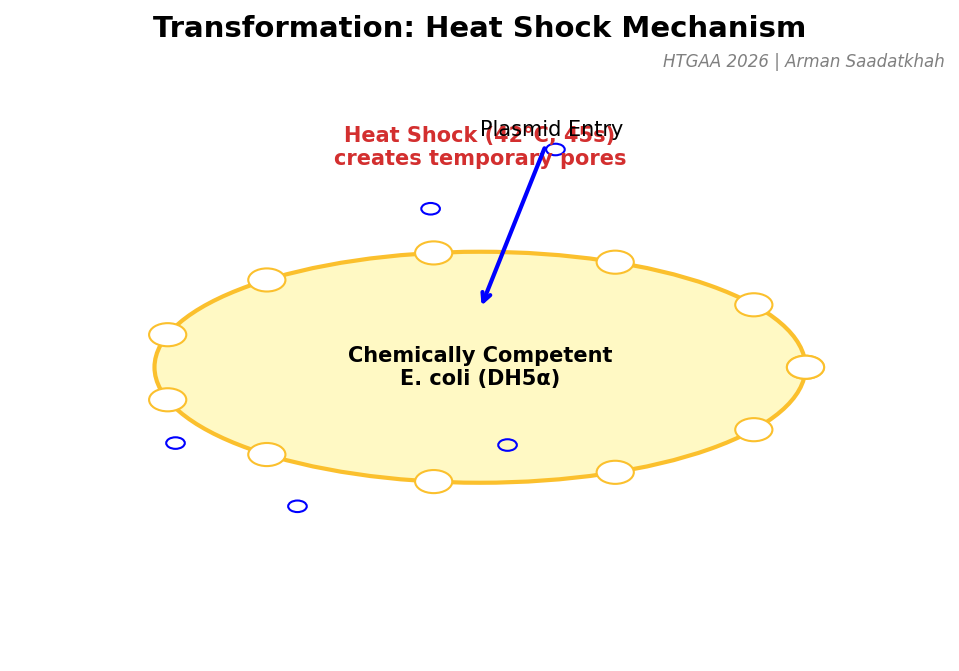
 Fig 4. Mechanism of Heat Shock transformation in chemically competent DH5α cells.
Fig 4. Mechanism of Heat Shock transformation in chemically competent DH5α cells.
Protocol | Part 1: PCR & Purification
PCR Setup
We set up two reactions to amplify the backbone and the mutant inserts.
- Backbone Fragment: ~2.5 kb (Primers: Backbone Fwd/Rev)
- Insert Fragment: ~0.5 kb (Primers: Color Fwd/Rev)
DpnI Digest & Purification
After PCR, we treat the samples with DpnI to digest the methylated template DNA (mUAV), ensuring only our newly synthesized mutant fragments are used in the assembly. We then purify the DNA using a silica-based column.
Protocol | Part 2: Assembly & Transformation
Gibson Assembly
We mix our purified Backbone and Insert fragments in a 1:2 molar ratio with the Gibson Assembly Master Mix and incubate at 50°C for 30 minutes.
Transformation
The assembled plasmids are transformed into DH5α competent cells.
- Thaw cells on ice for 10 minutes.
- Add 4 uL of assembly product.
- Heat Shock: 42°C for exactly 45 seconds.
- Outgrowth: Add SOC media and incubate at 37°C for 1 hour.
- Plating: Plate onto LB + Chloramphenicol agar.
Final Results | Example
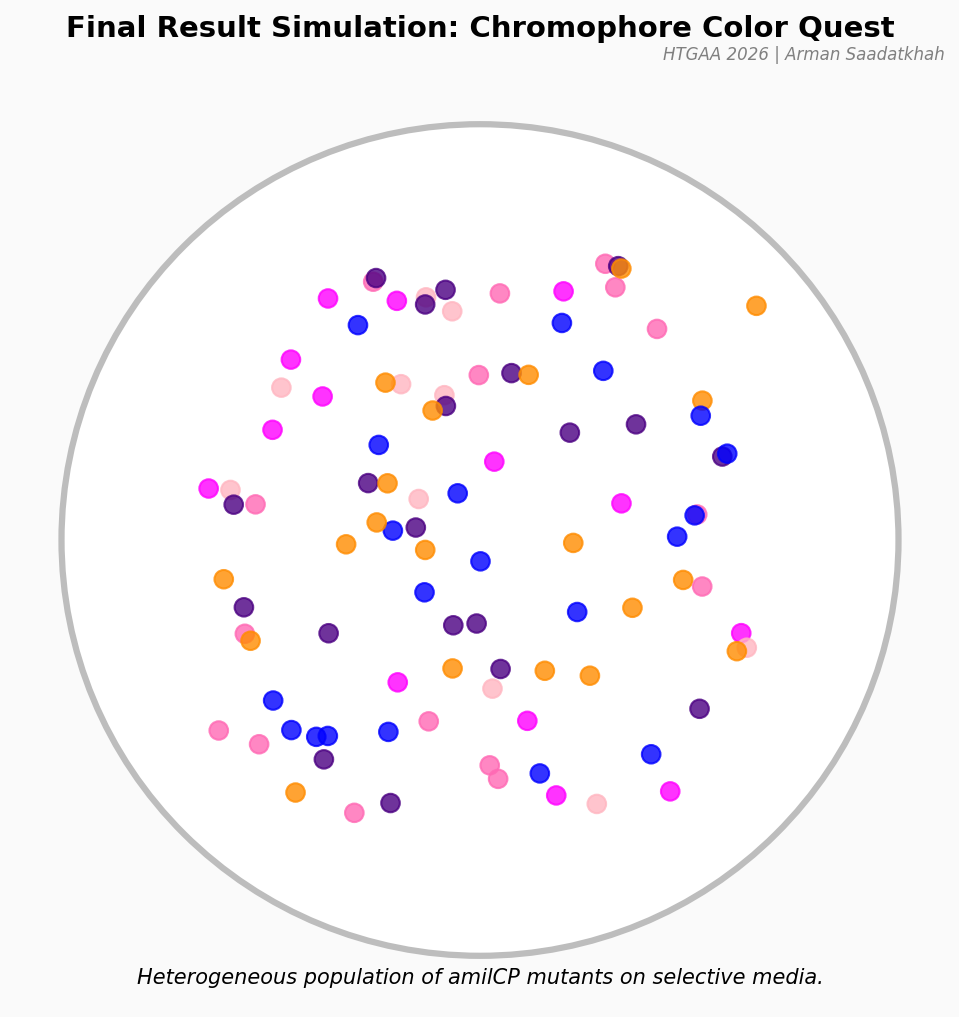
 Fig 5. Predicted result showing a variety of colorful colonies representing different amilCP mutations.
Fig 5. Predicted result showing a variety of colorful colonies representing different amilCP mutations.
After 72 hours of incubation, you should see a vibrant variety of purple, orange, pink, and blue colonies!
HTGAA 2026 | Arman Saadatkhah | Reference: Liljeruhm et al. (2018)