Week 9 Lab: Cell-Free Systems
Cell-Free Transcription-Translation (TX-TL) Systems
Overview | What is Cell-Free?
A cell-free system allows biological reactions to occur outside of living cells. By extracting and using cellular components like ribosomes, RNA polymerase, amino acids, and ATP, this method enables reactions in a controlled, simplified environment.
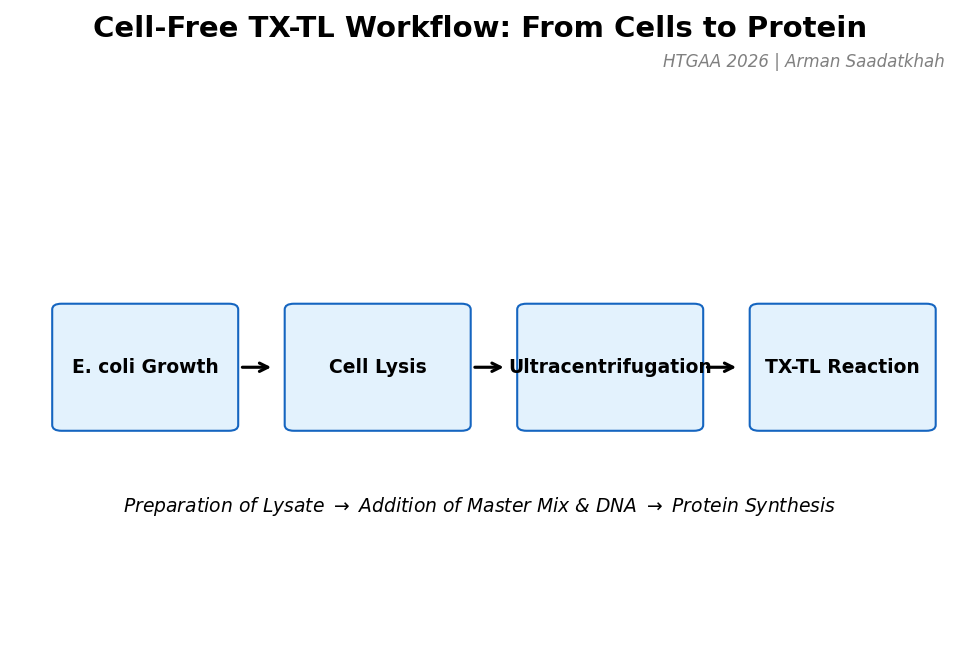
 Fig 1. The general workflow for preparing and running a cell-free TX-TL reaction.
Fig 1. The general workflow for preparing and running a cell-free TX-TL reaction.
Applications
- Synthetic Biology: Testing circuits without cellular constraints.
- Protein Engineering: Rapid production of toxic or difficult proteins.
- Biosensing: Creating portable, paper-based diagnostic tools.
Technical Overview | TX-TL Production
A. Cell Extract Preparation
The process begins with E. coli growth, followed by washing and cell disruption (sonication or freeze-thaw). Ultracentrifugation at 30,000g separates the necessary machinery (ribosomes, factors) from debris. A strict cold chain is maintained to prevent enzymatic degradation.
B. Master Mix Components
The master mix provides the chemical environment and energy required for synthesis.
| Component | Function |
|---|---|
| HEPES (500 mM) | pH buffering for optimal enzyme activity. |
| ATP, GTP, CTP, UTP | Nucleotides for transcription and energy (ATP/GTP). |
| E. coli tRNA | Essential for amino acid delivery during translation. |
| 3-PGA or PEP | Energy regeneration sources to maintain ATP levels. |
| Mg/K-Glutamate | Essential ionic cofactors for enzymatic machinery. |
| Murine RNase Inhibitor | Protects mRNA templates from degradation. |
System Comparison | PURE vs. Lysate
There are two primary types of cell-free systems: the PURE System (defined, purified components) and Whole Cell Extract (crude lysate).
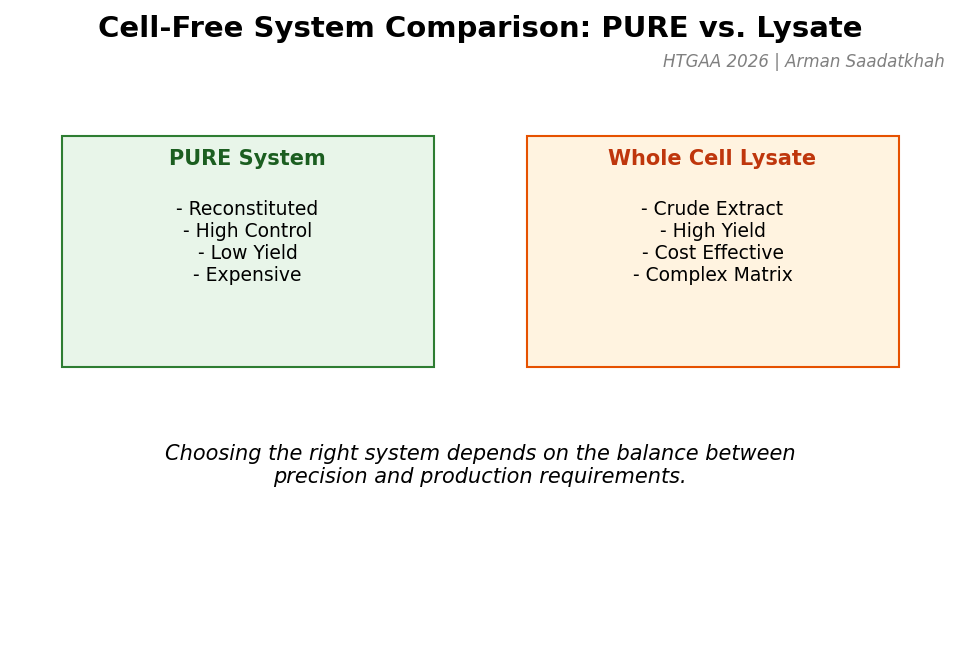

 Fig 2. Comparison between the PURE system and Whole Cell Lysate systems.
Fig 2. Comparison between the PURE system and Whole Cell Lysate systems.
Lab Exercise | amilGFP Induction Quest
The objective of this lab was to quantify protein production in a cell-free extract using different IPTG levels to induce the expression of amilGFP from a T7-IPTG-inducible plasmid.
Results & Analysis
Fluorescence was monitored over an 8-hour incubation at 30°C. We analyzed the final-point results using ImageJ Color Histogram Analysis.
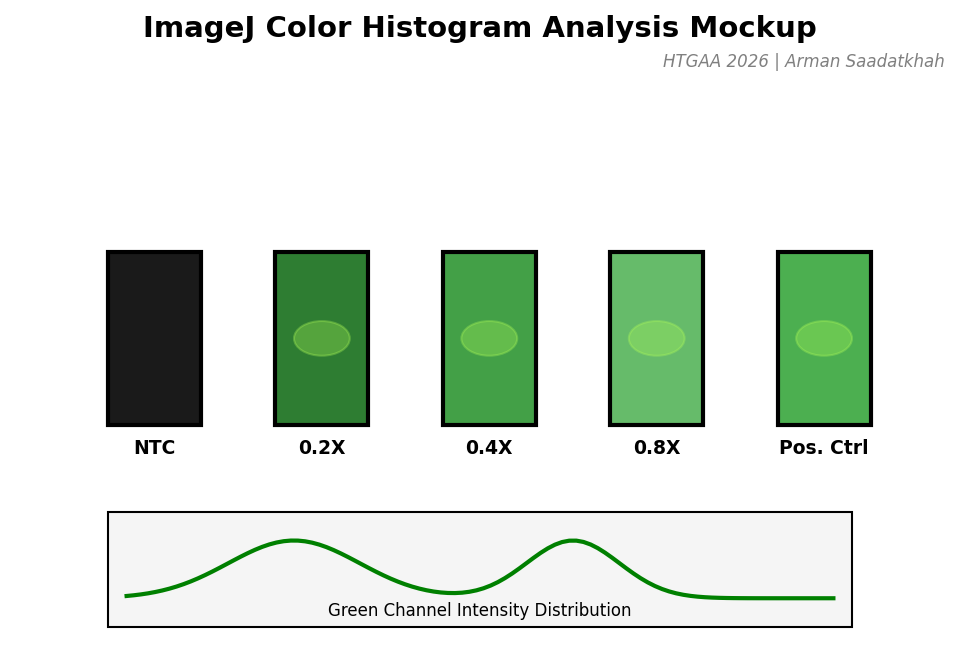
 Fig 3. Mockup of the color histogram analysis used to quantify GFP expression levels.
Fig 3. Mockup of the color histogram analysis used to quantify GFP expression levels.
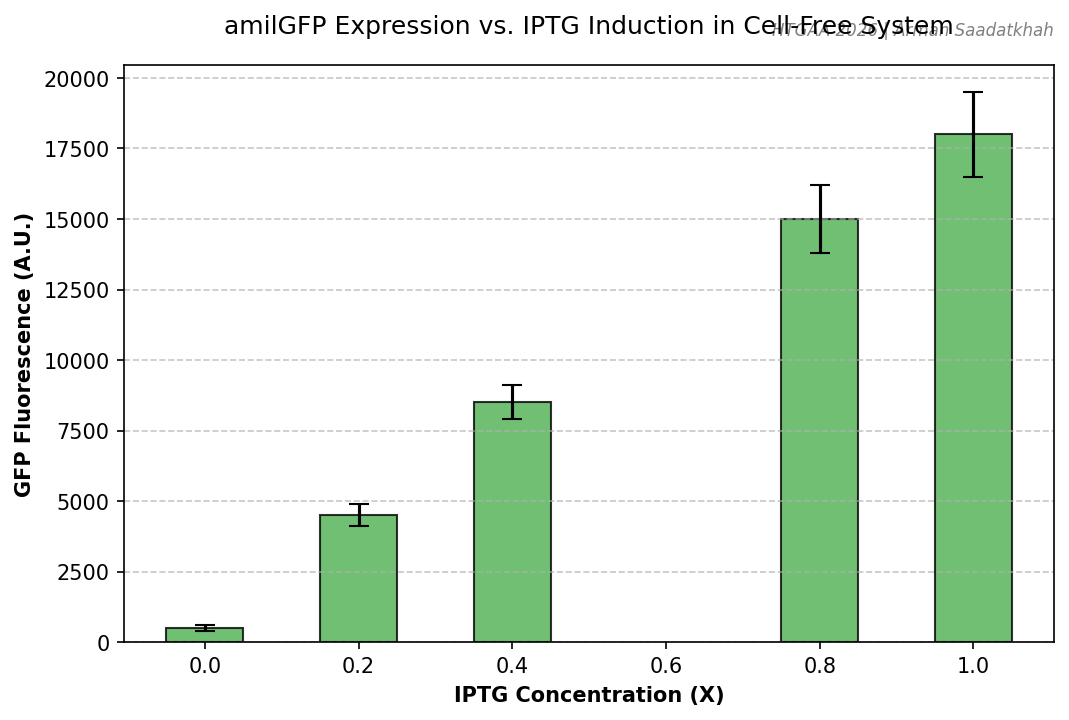
 Fig 4. Quantified GFP fluorescence across varying IPTG inducer concentrations.
Fig 4. Quantified GFP fluorescence across varying IPTG inducer concentrations.
Observations
- Fold Change: We observed a significant dose-response relationship between IPTG concentration and GFP yield.
- Background: The Non-Template Control (NTC) showed minimal background fluorescence, validating the specificity of the T7-IPTG system.
Homework Questions
1. Advantages of Cell-Free Protein Synthesis
Cell-free systems offer unparalleled flexibility because the reaction is directly accessible. You can add non-canonical amino acids, adjust magnesium concentrations mid-run, or introduce toxic components that would kill a living cell.
- Beneficial Case 1: Toxic Proteins. Expressing antimicrobial peptides that would lyse the production host.
- Beneficial Case 2: Rapid Prototyping. Testing 100+ genetic circuit variants in a single day without the time-consuming transformation/cloning cycle.
2. Main Components and Roles
- Extract: Contains the molecular hardware (ribosomes, tRNA synthetases, RNA polymerase).
- Master Mix: Provides the fuel (ATP/GTP) and chemical environment (buffers, ions).
- Template (DNA): The software/instructions for the specific protein to be synthesized.
3. Energy Regeneration
Energy regeneration is critical because the initial ATP/GTP supply is exhausted within minutes. We use secondary energy sources like 3-PGA or PEP which, through the action of metabolic enzymes in the lysate, phosphorylate ADP back into ATP, ensuring a continuous supply for several hours.
4. Prokaryotic vs. Eukaryotic Systems
- Prokaryotic (E. coli): High yield, fast, and simple. Ideal for producing amilGFP or other reporter proteins.
- Eukaryotic (Wheat Germ): Slower but capable of complex Post-Translational Modifications (PTMs). Ideal for producing human signaling proteins like Insulin or complex antibodies that require chaperones for correct folding.
5. Optimizing Membrane Protein Expression
I would design the experiment by adding nanodiscs or synthetic liposomes directly to the cell-free reaction.
- Challenge: Membrane proteins are hydrophobic and aggregate in aqueous buffers.
- Solution: The presence of a lipid bilayer allows the protein to co-translationally insert into a stable environment, mimicking its natural state.
6. Troubleshooting Low Yield
- Reason 1: Template Degradation. Strategy: Increase concentration of Murine RNase Inhibitor or use a circular plasmid instead of linear PCR product.
- Reason 2: Substrate Depletion. Strategy: Perform the reaction in a dialysis format to continuously supply small molecules (NTPs, amino acids) and remove byproducts.
- Reason 3: Incorrect Mg2+ Concentration. Strategy: Perform a magnesium titration (e.g., 4mM to 16mM) to find the specific optimum for the T7 polymerase and ribosome used.
HTGAA 2026 | Arman Saadatkhah | Reference: TX-TL Laboratory Manual