Week 11 Lab: Introduction to Cloud Laboratories
Cloud Laboratories: Collective Art and Cell-Free Optimization
Overview | Introduction
Cloud laboratories are making science accessible, affordable, and reproducible. This lab showcases how cloud labs enable human creativity at scale and provide a platform for global collaboration. Our goal is to design a scientifically rigorous cell-free fluorescent protein optimization experiment together.
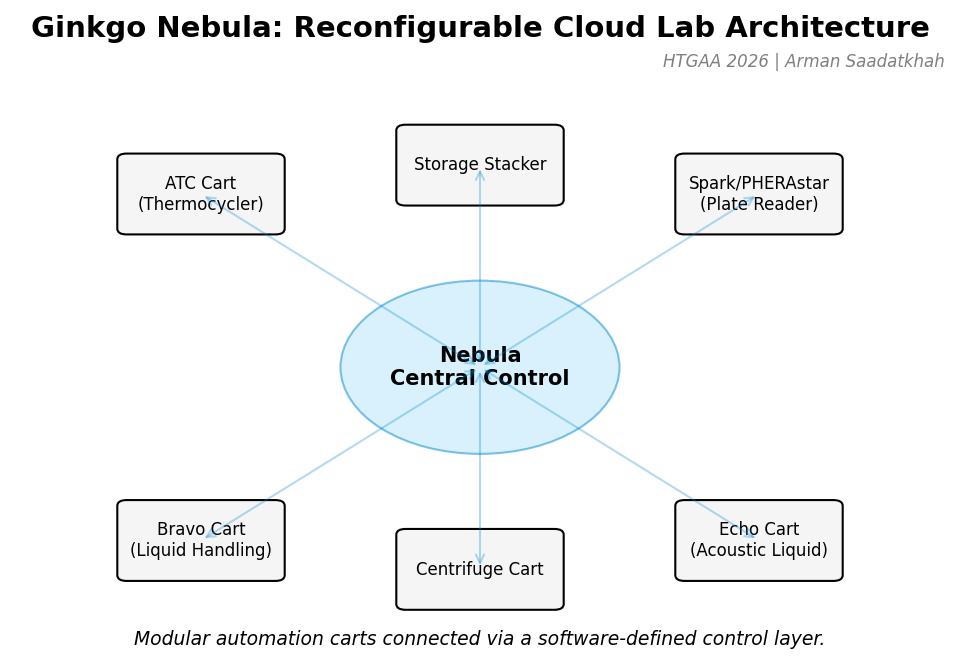
 Fig 1. Ginkgo Nebula architecture featuring modular automation carts connected via a software-defined control layer.
Fig 1. Ginkgo Nebula architecture featuring modular automation carts connected via a software-defined control layer.
1. The 1,536 Pixel Artwork Canvas
The community bioart project involved contributing pixels to a global artwork experiment. This demonstrates the power of automation (Opentrons and Echo systems) to handle high-density layouts that would be impossible manually.

 Fig 2. Simulation of the collective 1,536-pixel bioart canvas using various fluorescent proteins.
Fig 2. Simulation of the collective 1,536-pixel bioart canvas using various fluorescent proteins.
My Contribution:
- Pixel Location: I contributed to the “DNA” pattern on the bottom right plate.
- Reflection: I liked the seamless integration of individual designs into a unified biological canvas. For next year, real-time visualization of the design progress would be a great addition.
2. Cell-Free Protein Synthesis Reagents
A cell-free reaction requires a complex mixture of “hardware” (lysate) and “fuel” (master mix).
Master Mix Comparison
There are two main strategies for energy provision:
- 1-hour PEP-NTP: Optimized for speed; uses high-energy phosphate donors (PEP) for rapid protein synthesis.
- 20-hour NMP-Ribose-Glucose: Optimized for sustained production; uses secondary metabolism to regenerate ATP over longer periods.
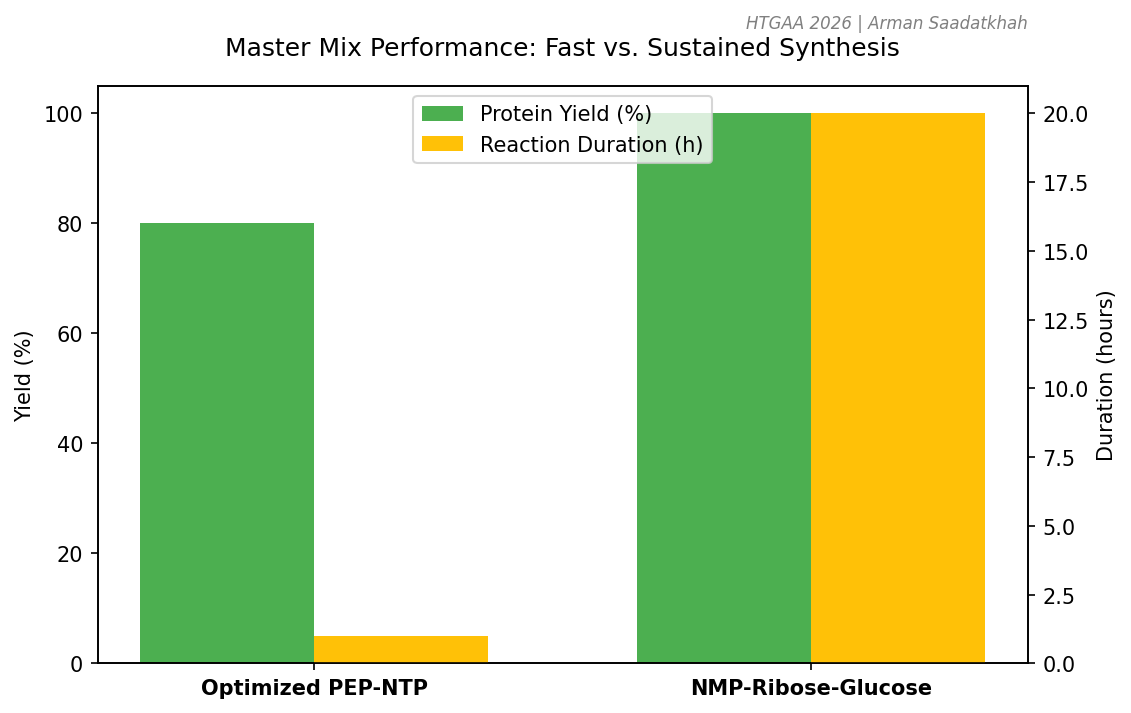
 Fig 3. Performance trade-offs between rapid (PEP-NTP) and sustained (NMP-based) energy systems.
Fig 3. Performance trade-offs between rapid (PEP-NTP) and sustained (NMP-based) energy systems.
Component Roles:
- BL21 (DE3) Star Lysate: The molecular machinery (ribosomes, tRNA synthetases, T7 RNA Polymerase).
- HEPES-KOH / Potassium Glutamate: Buffers and salts to maintain optimal pH and ionic strength.
- Ribose/Glucose/NMPs: Building blocks and energy precursors for sustaining the reaction.
- Guanine Bonus: Transcription can occur even without GMP if Guanine is present because the lysate contains salvage pathway enzymes (e.g., phosphoribosyltransferases) that can convert Guanine into GMP/GTP.
3. Global Experiment: Fluorescent Protein Properties
We used 6 different proteins for our collaborative painting, each with unique biophysical characteristics.
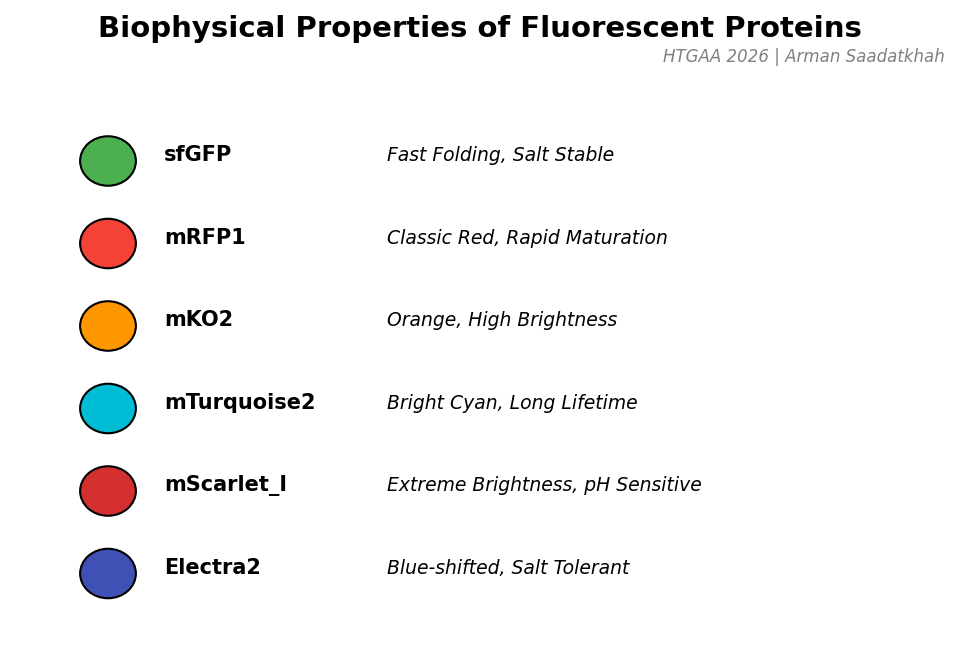

 Fig 4. Key functional properties of the fluorescent proteins used in the collective optimization experiment.
Fig 4. Key functional properties of the fluorescent proteins used in the collective optimization experiment.
Optimization Hypothesis:
- Target Protein:
mScarlet_I - Identified Property: High pH sensitivity (fluorescence drops in acidic conditions).
- Hypothesis: By increasing the
HEPESbuffer concentration in the master mix from 500mM to 750mM, we can maintain a more stable neutral pH as metabolic byproducts accumulate, thereby maximizing fluorescence over the 36-hour incubation.
4. Generic Cloud Lab Operations (JSON)
Automation in cloud labs like Ginkgo Nebula is driven by software-defined protocols. Below is a sample configuration for a spark_read operation:
HTGAA 2026 | Arman Saadatkhah | Reference: Ginkgo Bioworks Nebula Platform