Week 12 Lab: Bioproduction of Beta-Carotene and Lycopene
Bioproduction of Beta-Carotene and Lycopene
Overview | Objective
In this two-day lab, you will work with genetically modified E. coli to produce beta-carotene and lycopene, key plant pigments and antioxidants found in carrots and tomatoes. Using the plasmids pAC-LYC and pAC-BETA, which encode the pathways for lycopene and beta-carotene production, your goal will be to optimize the production of these two pigments.
This lab explores bioproduction, using biological systems—such as microorganisms (e.g., bacteria, fungi, algae) or plant and animal cells—to produce valuable compounds or materials. Bioproduction plays a critical role in various fields, including industrial biotechnology, pharmaceuticals, agriculture, and food production, enabling the creation of proteins, enzymes, antibiotics, biofuels, and more.
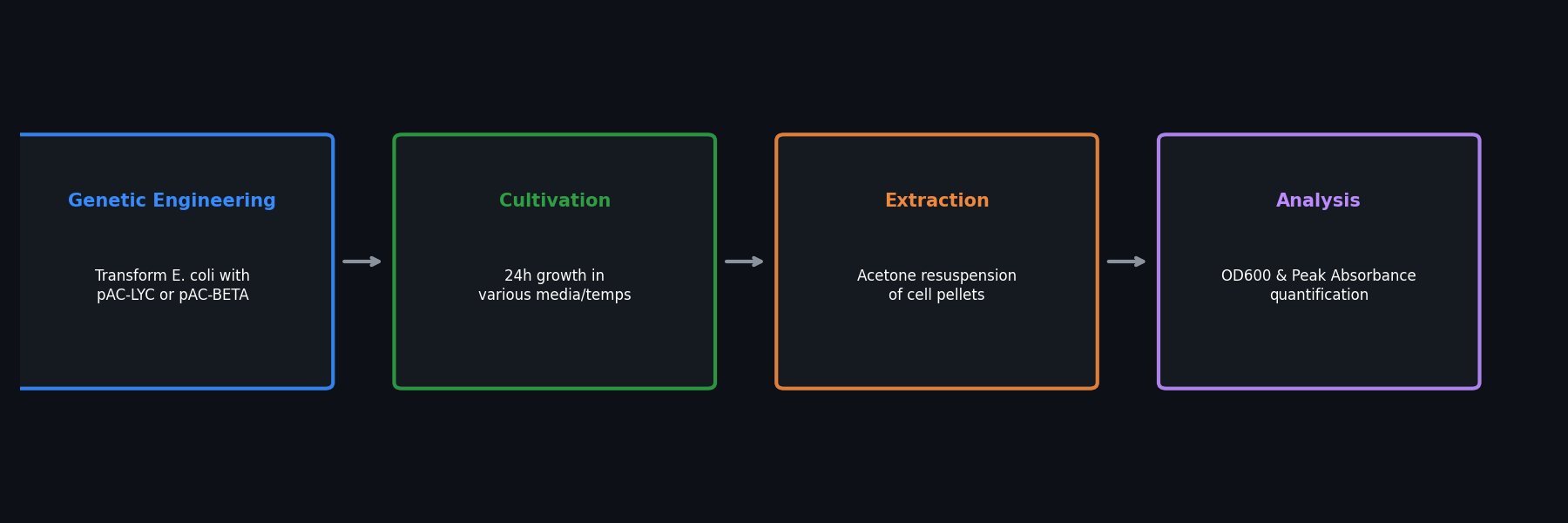
 Fig 1. Overview of the bioproduction workflow, from genetic engineering to pigment extraction.
Fig 1. Overview of the bioproduction workflow, from genetic engineering to pigment extraction.
Overview | Concepts Learned & Skills Gained
A major challenge in bioproduction is the metabolic competition between the organism’s natural drive to reproduce and the production of the target compound. In this lab, you will explore how to fine-tune this balance to maximize pigment production, gaining hands-on experience in optimizing bioproduction systems.
You’ll investigate this by modifying culture conditions:
- Temperature (30°C vs 37°C)
- Growth Media Composition (LB vs 2YT, with and without fructose)
By examining how different environmental factors influence bacterial growth and synthesis, you will gain practical insights into metabolic engineering, advancing the potential for scalable bioproduction of essential compounds.
Pre-Lab | Reading
Referring to the pathway, lycopene is the red pigment that gives tomatoes their red color. This pigment is also made by microbes. In fact, transferring a 3-enzyme pathway to E. coli can convert farnesyl diphosphate (FPP) to lycopene.
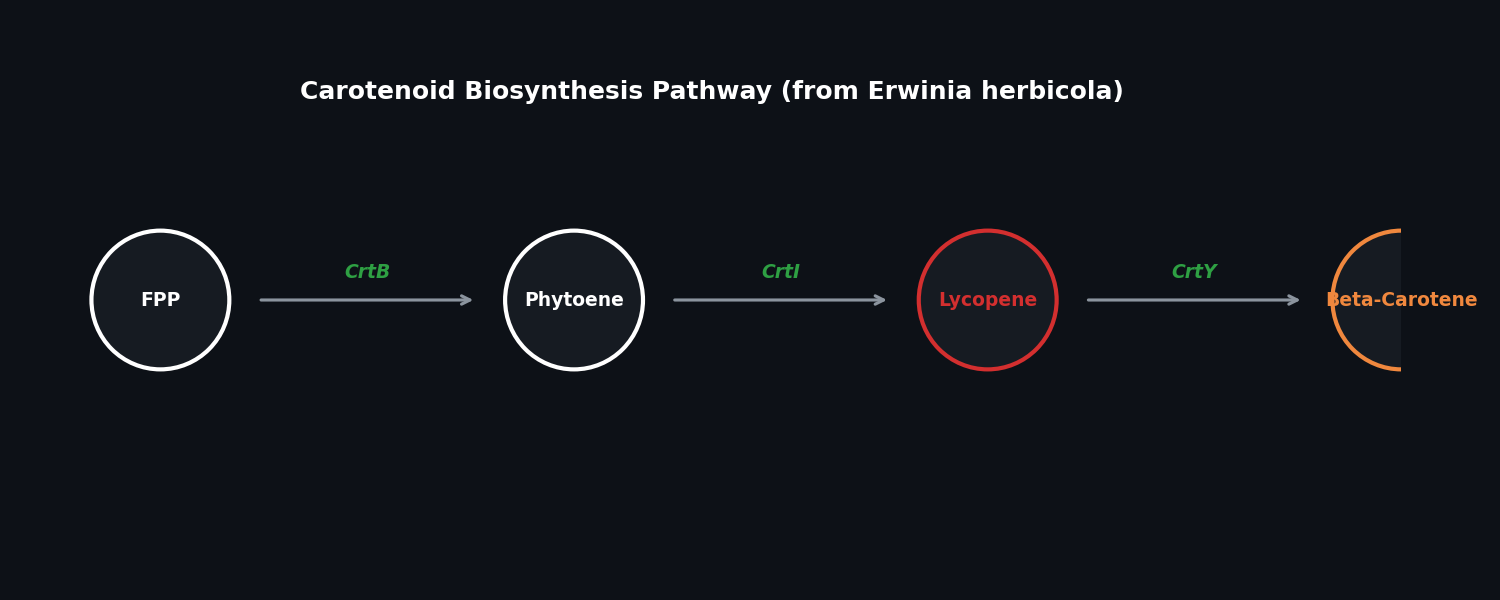
 Fig 2. The biosynthetic pathway for Lycopene and Beta-Carotene.
Fig 2. The biosynthetic pathway for Lycopene and Beta-Carotene.
Plasmids
- pAC-LYC: Contains three genes from Erwinia herbicola: CrtE, CrtI, and CrtB. Produces lycopene.
- pAC-BETA: Produces beta-carotene through the addition of the Erwinia herbicola CrtY gene to the lycopene pathway.
- Resistance: Both plasmids include the gene for chloramphenicol resistance.
- Reference: Addgene #53270, #53272.
Optical Density (OD600)
We will be estimating cell growth by measuring the optical density of cells at a light wavelength of 600 nm (OD600). At 600 nm, dense cell suspensions scatter light, which correlates to approximate cell count. Always blank with the specified media first.
Pre-Lab | Safety
- Acetone: Review the Safety Data Sheet. Acetone is compatible with polypropylene (50 mL conical and 1.5 mL microcentrifuge tubes).
Protocol | Part 1: Overnight Cultures
Time Estimate: 30 Minutes setup, 24 Hour Incubation
Media, Equipment and Consumables
- LB and 2YT with chloramphenicol
- Fructose (for supplementation)
- Pipette set, serological pipettes, and tips
- Culture tubes
- Incubation room (30°C and 37°C)
Experimental Matrix (16 Conditions)
You will set up 16 unique conditions (plus duplicates and media controls, total 34 cultures).
| Condition | Plasmid | Temp | Medium |
|---|---|---|---|
| 1, 2 | pAC-LYC | 30°C, 37°C | LB |
| 3, 4 | pAC-LYC | 30°C, 37°C | LB + Fructose |
| 5, 6 | pAC-LYC | 30°C, 37°C | 2YT |
| 7, 8 | pAC-LYC | 30°C, 37°C | 2YT + Fructose |
| 9, 10 | pAC-BETA | 30°C, 37°C | LB |
| 11, 12 | pAC-BETA | 30°C, 37°C | LB + Fructose |
| 13, 14 | pAC-BETA | 30°C, 37°C | 2YT |
| 15, 16 | pAC-BETA | 30°C, 37°C | 2YT + Fructose |
Procedure
- Prepare 3 mL of the specified media (supplemented with antibiotic).
- Inoculate with 1 µL of E. coli starter culture containing the specified plasmid.
- Grow for 24 hours in the circular roller drum at the appropriate temperature.
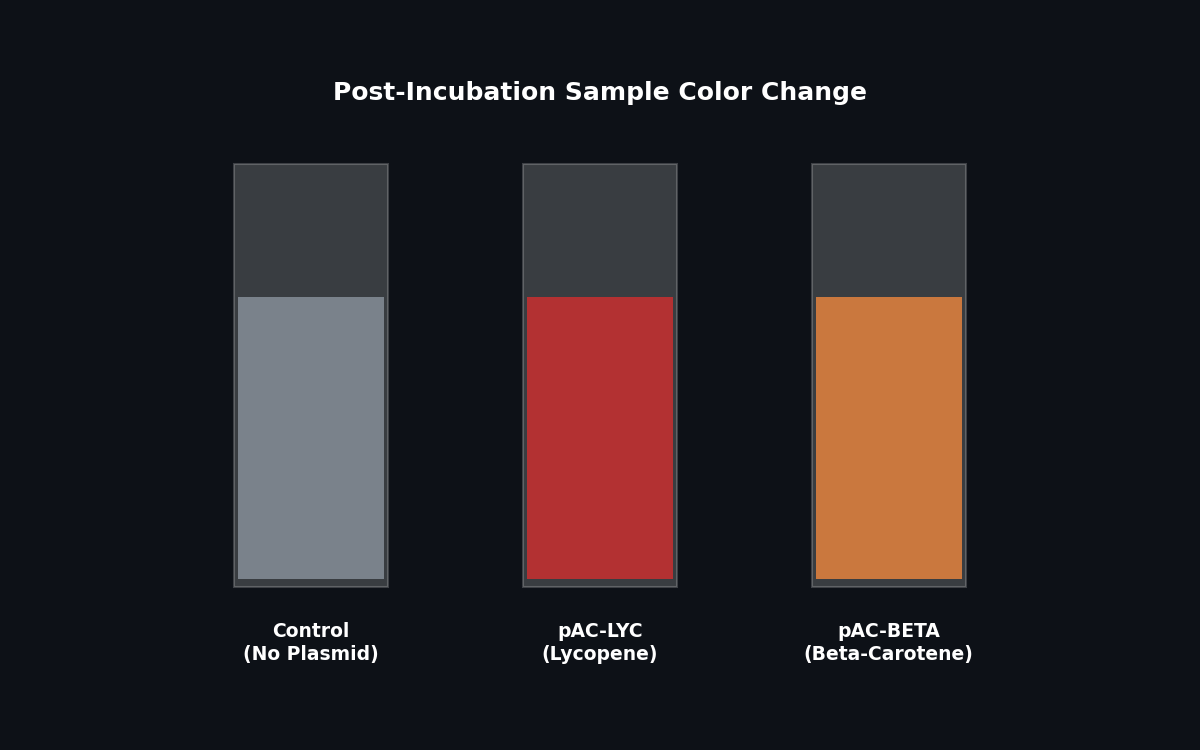
 Fig 3. Expected color change post-incubation indicating pigment production.
Fig 3. Expected color change post-incubation indicating pigment production.
Protocol | Part 2: Analyze OD600 and Peak Absorbance
Time Estimate: 180 Minutes
OD600 Measurement
- Open the OD600 program on the spectrophotometer and blank with the respective media.
- Measure using 800 µL of each culture in a cuvette.
- Record values in an external table.
Pigment Extraction (Peak Absorbance)
- Vortex the culture tube to ensure bacteria are suspended.
- Transfer 1000 µL to a 1.5 mL microcentrifuge tube.
- Centrifuge at 14,000 rpm for 1 minute.
- Remove and discard the supernatant.
- Repeat steps 2-4 two more times (total 3 mL concentrated into one pellet). Photograph the pellets!
- Add 700 µL of acetone to the pellet and pipette up and down until resuspended. Acetone disrupts cell structure and solubilizes the carotenoids.
- Centrifuge at 14,000 rpm for 1 minute.
- Transfer 600 µL of the pigmented supernatant to a fresh tube.
- Dilute with 600 µL of water (prevents acetone from corroding polystyrene cuvettes).
- Measure absorbance on the spectrophotometer:
- Lycopene: Measure at 474 nm.
- Beta-Carotene: Measure at 456 nm.
- Bleach samples and clean cuvettes after use.
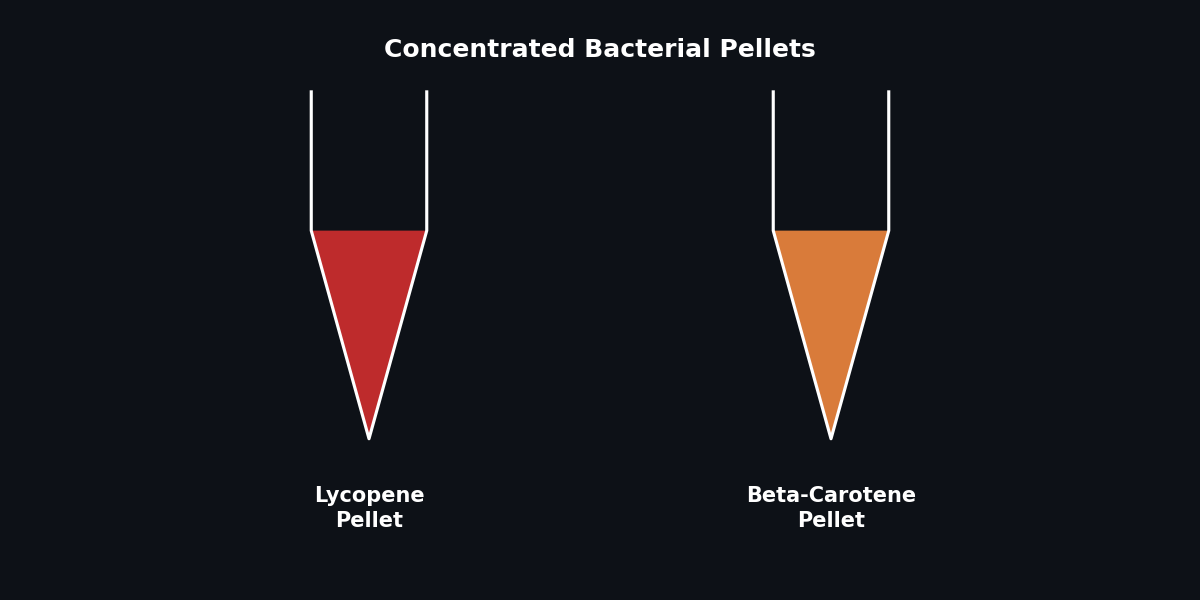
 Fig 4. Visualizing carotenoid accumulation in bacterial pellets.
Fig 4. Visualizing carotenoid accumulation in bacterial pellets.
Protocol | Part 3: Analysis
Time Estimate: 1 Hour
Compare relative pigment production per cell. Normalize each sample’s absorption peak measurement by its OD600 value.
- Which culture conditions led to the highest production?
- Plot your results using Excel or Python and include in your writeup.
Key Formula: Absorbance = 2 – log10(%Transmittance) (Beer-Lambert Law).
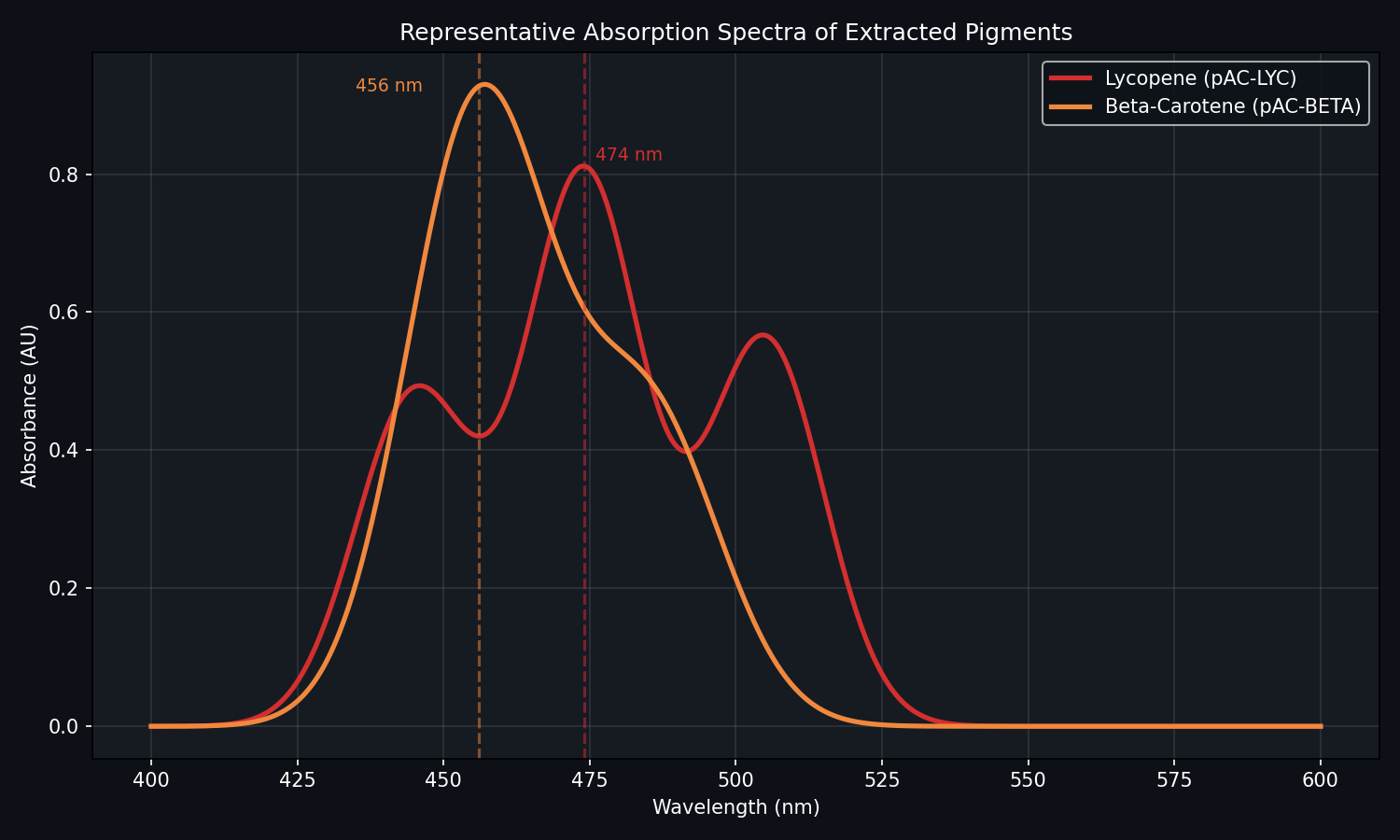
 Fig 5. Example absorption spectrum. Note how pAC-BETA often performs better at 37°C.
Fig 5. Example absorption spectrum. Note how pAC-BETA often performs better at 37°C.
Post-Lab Questions | Mandatory
- Which genes when transferred into E. coli will induce the production of lycopene and beta-carotene, respectively?
- Lycopene: CrtE, CrtI, and CrtB.
- Beta-Carotene: CrtE, CrtI, CrtB, and CrtY.
- Why do the plasmids need to contain an antibiotic resistance gene?
- To provide a selective advantage to the transformed E. coli. Only bacteria containing the plasmid will survive on media supplemented with chloramphenicol, ensuring the population maintains the engineered pathway.
- What outcomes might we expect when we vary the media, presence of fructose, and temperature?
- Higher temperatures (37°C) typically increase growth rates but may lead to protein misfolding or metabolic stress. Nutrient-rich media (2YT) supports higher biomass than LB. Fructose provides an alternative carbon source that can bypass certain regulatory bottlenecks in the MEP pathway, potentially increasing pigment titer.
- Generally describe what “OD600” measures and how it is interpreted here.
- OD600 measures the turbidity (light scattering) of a culture at 600 nm. It is used as a proxy for cell density. In this lab, it allows us to normalize pigment production to the number of cells, ensuring we compare efficiency rather than just total volume.
- What are other experimental setups where acetone could be used to separate cellular matter from a compound?
- Extracting chlorophyll from leaves, isolating lipids from tissues, or precipitating proteins during DNA extraction.
- Why engineer E. coli for this when Erwinia herbicola naturally produces them?
- E. coli is a well-characterized “workhorse” with faster growth cycles, established genetic tools, and a lack of native pigments that would interfere with quantification. It allows for higher-scale, more controllable bioproduction.
Post-Lab Questions | For Committed Listeners Only
The following questions are based on supplemental readings and metabolic pathway design.
Metabolic Pathway Deep-Dive
- Enzymes: What are the enzymes of the carotene pathway?
- CrtE (GGPP synthase), CrtB (Phytoene synthase), CrtI (Phytoene desaturase), and CrtY (Lycopene β-cyclase).
- Rate-Limiting Step: Which step takes the longest, and which enzyme is responsible for it?
- Within the carotenoid pathway, CrtB (Phytoene synthase) is the primary rate-limiting enzyme. Globally in E. coli, the DXS enzyme (precursor supply) is often the major bottleneck.
DNA Construct Design
- Organism Choice: Would you choose E. coli or S. cerevisiae for bioproduction? Why?
- E. coli is preferred for rapid prototyping (20 min doubling time). S. cerevisiae (yeast) is preferred for food-grade products (GRAS) and has a superior native MVA pathway for precursor supply.
- Expression Vector: Design an expression vector for E. coli. What parts are needed (RBS, terminators, operators, promoters)?
- Required parts: Promoter (transcription start), Operator (regulation), RBS (translation initiation), Gene of Interest, Terminator (transcription stop), Selectable Marker (antibiotic resistance), and ORI (Origin of Replication).
- Promoters:
- What is the function of a promoter? To recruit RNA Polymerase to the DNA.
- What is the difference between repressible and inducible promoters? Repressible are “on” until a signal turns them off; Inducible are “off” until a signal (e.g., IPTG) turns them on.
- Which promoter would you use for a carotene/lycopene gene? Why? An inducible promoter (like pLac). This allows the culture to reach high biomass before “switching on” production, minimizing metabolic burden.
- Origin of Replication (ORI):
- What is an ORI? The site where DNA replication begins.
- What are compatibility groups? Groups of plasmids that cannot coexist because they compete for the same replication machinery.
- What is the best ORI for your chosen promoter and gene? A medium-copy ORI like p15A is often used for carotenoids to balance high yield with cellular health.
Advanced Engineering (Global Listeners)
- Tuning: Elaborate on how RBS, terminators, and operators contribute to metabolic tuning.
- RBS strength determines the protein concentration; Terminators prevent transcriptional read-through; Operators allow for dynamic control based on environmental signals.
- Aptamers & Riboswitches: How can these be used for metabolic engineering in prokaryotes?
- They act as RNA-based sensors that can sense product levels and automatically down-regulate or up-regulate expression to maintain optimal metabolic flux.
- Assembly: What approach (e.g., Gibson, Golden Gate) would you use to join these parts?
- Gibson Assembly for large, scarless constructs; Golden Gate for high-throughput modular library testing.
- Dream Application: Elaborate on a biosynthetic pathway you would engineer in E. coli. What is its potential impact?
- Engineering E. coli to produce Astaxanthin (a potent antioxidant) for sustainable aquaculture, reducing the need for synthetic dyes in the salmon industry.
Yeast Engineering (S. cerevisiae)
- Integration: Create an integration cassette for homologous recombination.
- Structure: [5’ Homology Arm] - [Promoter] - [Gene] - [Terminator] - [Marker] - [3’ Homology Arm].
- Parts: List the eukaryotic-specific parts (e.g., Kozak sequence, terminators) needed.
- Kozak sequence (ACC ATGG), Eukaryotic Promoters (pGAL1), and Eukaryotic Terminators (CYC1).
- Site Selection: What chromosome site would you use for integration and why?
- The HO locus is ideal as it is a “safe harbor” where integration does not disrupt essential cellular functions.
HTGAA 2025 | Lab Protocol: Bioproduction of Beta-Carotene and Lycopene