Group Final Project: MS2 Phage Engineering
Group Final Project: Battling Antibiotic Resistance with Engineered Phages
Overview | Phage Therapy
Phage therapy is the therapeutic use of bacteriophages to treat bacterial infections. Phages are viruses that infect bacteria with extreme specificity, often targeting only a single strain. This specificity allows phage therapy to kill harmful pathogens while sparing beneficial bacteria, offering a potential solution to the global crisis of antibiotic resistance.
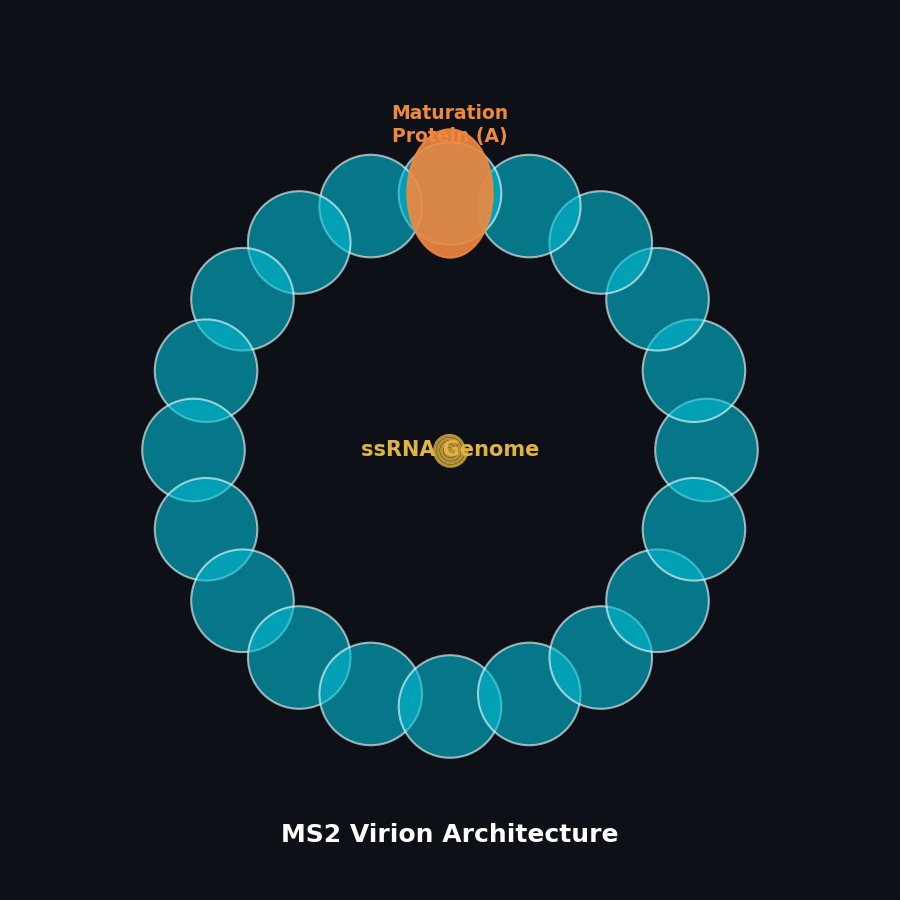

 Fig 1. Architecture of the MS2 virion, featuring a single-stranded RNA genome and maturation protein.
Fig 1. Architecture of the MS2 virion, featuring a single-stranded RNA genome and maturation protein.
The Arms Race
For billions of years, a co-evolutionary arms race has existed between phages and bacteria. Bacteria develop defense mechanisms, and phages evolve to overcome them. Our goal in this project is to use synthetic biology and protein engineering to give phages a “head start” in this race.
The Group Project Objective
We aim to engineer the MS2 bacteriophage to be more efficient at killing its host, Escherichia coli, particularly in overcoming host resistance mechanisms.
MS2 Biology and the Lysis Protein (L)
MS2 infects E. coli by attaching to the F-pilin on the host membrane. Once inside, it hijacks the host machinery to replicate. A critical component of its life cycle is the Lysis Protein (L), which triggers the breakdown of the bacterial cell wall to release new virions.
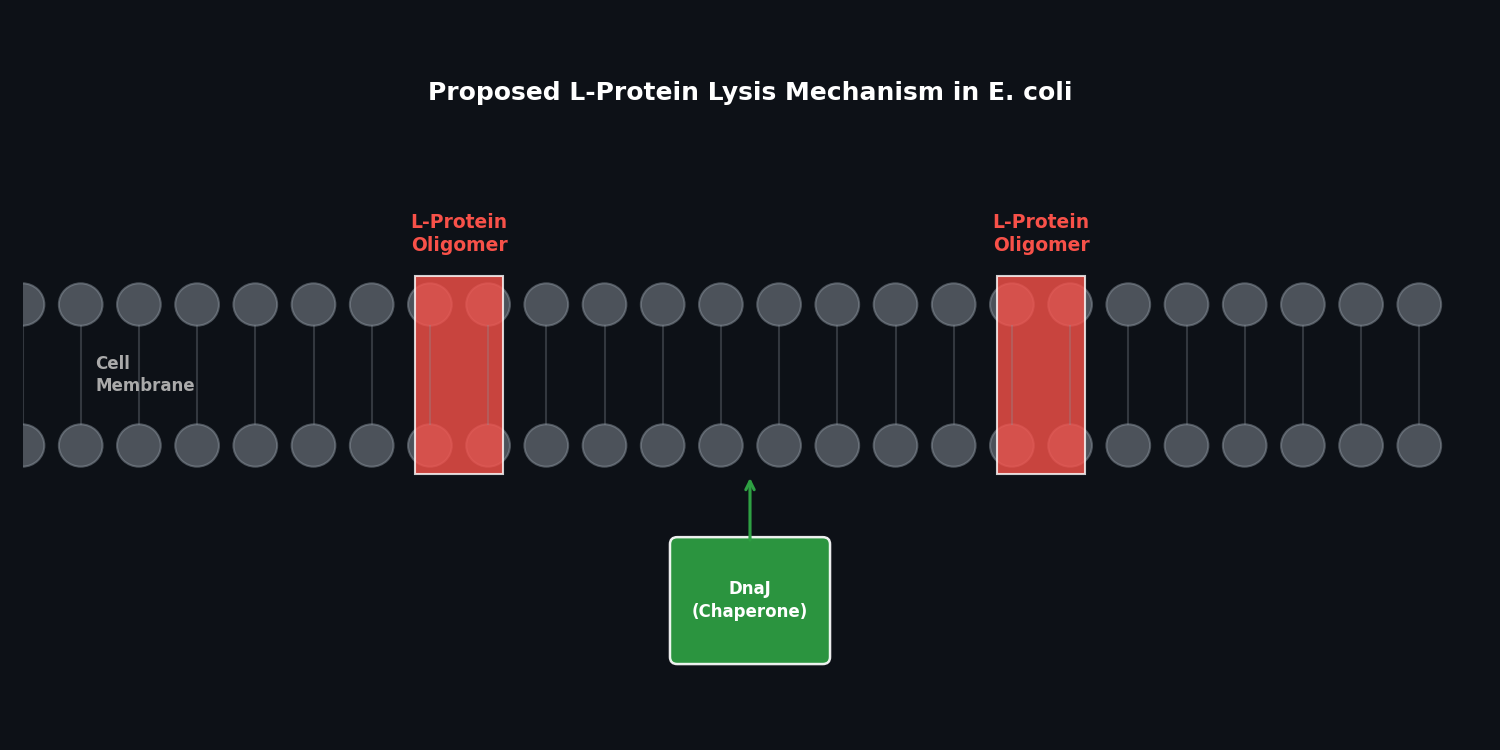
 Fig 2. Proposed mechanism of L-protein mediated lysis and its dependency on host chaperones like DnaJ.
Fig 2. Proposed mechanism of L-protein mediated lysis and its dependency on host chaperones like DnaJ.
The DnaJ Bottleneck
E. coli can develop resistance by mutating host chaperones like DnaJ, which the L protein requires for proper folding. If DnaJ is mutated, the L protein fails, and the infection cycle stops.
Our Strategy: Engineering the L Protein
We are searching for mutations in the L gene that achieve:
- Chaperone Independence: Allowing the L protein to function even if DnaJ is mutated.
- Increased Efficiency: Faster or more potent lysis to reduce the window for host adaptation.
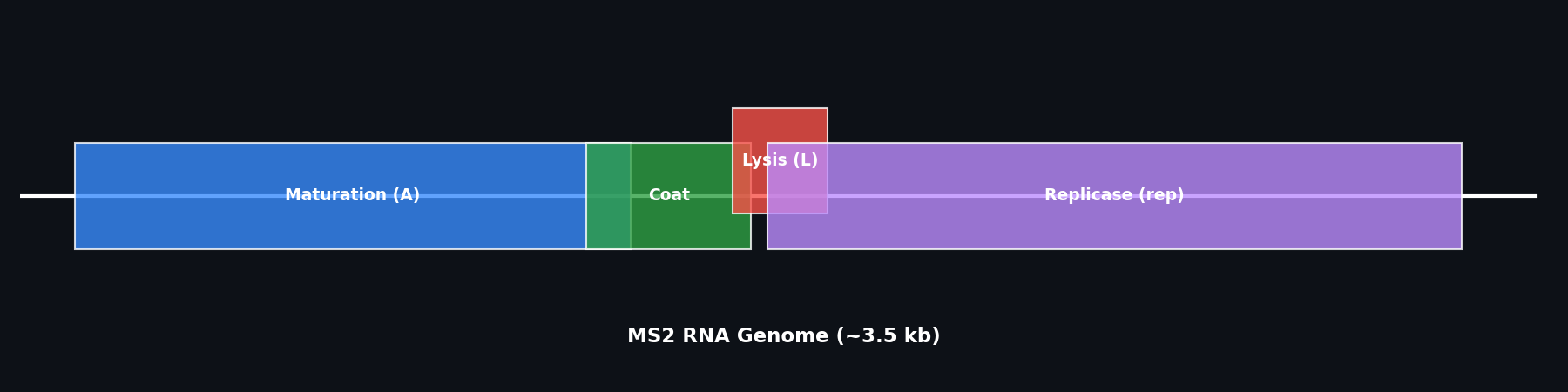
 Fig 3. Genetic map of the MS2 RNA genome, highlighting the overlap between the Lysis (L) gene and other viral proteins.
Fig 3. Genetic map of the MS2 RNA genome, highlighting the overlap between the Lysis (L) gene and other viral proteins.
Research Workflow
This large-scale group effort proceeds through five distinct stages:
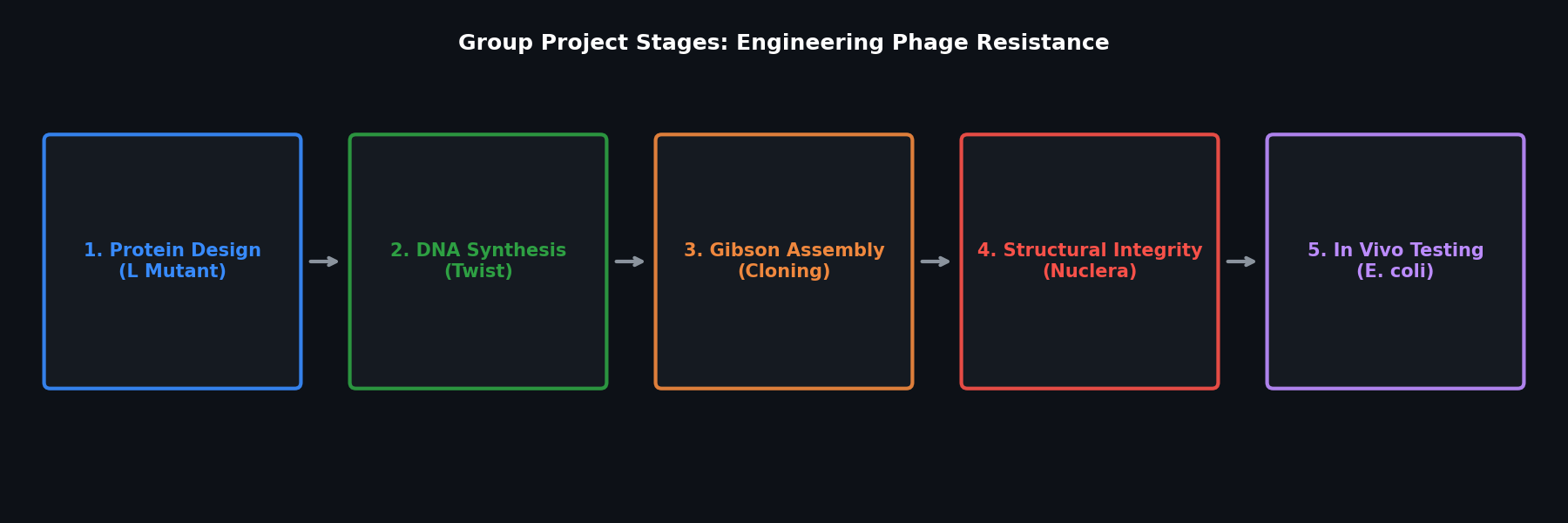
 Fig 4. The five stages of the HTGAA Group Final Project.
Fig 4. The five stages of the HTGAA Group Final Project.
- Stage 1: Computational Design: Use protein design tools to engineer novel L protein mutants.
- Stage 2: DNA Synthesis: Synthesize the mutant genes via Twist Bioscience.
- Stage 3: Cloning: Insert the mutant genes into plasmids using Gibson Assembly.
- Stage 4: Structural Validation: Test the structural integrity of the mutants using the Nuclera system.
- Stage 5: In Vivo Testing: Characterize the lysis efficiency of the mutants within E. coli cultures.
In-Depth Reading
- Identification of MS2 Lysis Protein Dependency on DnaJ
- Mutational Analysis of the MS2 Lysis Protein L
- Phage Therapy: Biological Mechanisms and Future Directions
HTGAA 2026 | Group Final Project | Engineering the Phage-Host Arms Race