Week 5 HW: Protein Design II
Part 1: Generate Binders with PepMLM
I retrieved the human SOD1 sequence from UniProt (P00441) in FASTA format.
Original canonical sequence:
I introduced the A4V mutation by changing Alanine (A) to Valine (V) at position 4 of the mature protein sequence (index 4, 0-based).
Mutant SOD1-A4V sequence:
The change is visible at positions 4-5: the original MATKA...
becomes MATKV...
Using the PepMLM-650M Colab, I ran the model with the following parameters:
| Parameter | Value |
|---|---|
| Target sequence | SOD1-A4V (mutant) |
| Peptide length | 12 |
| Top-K | 3 |
| Number of binders | 4 |
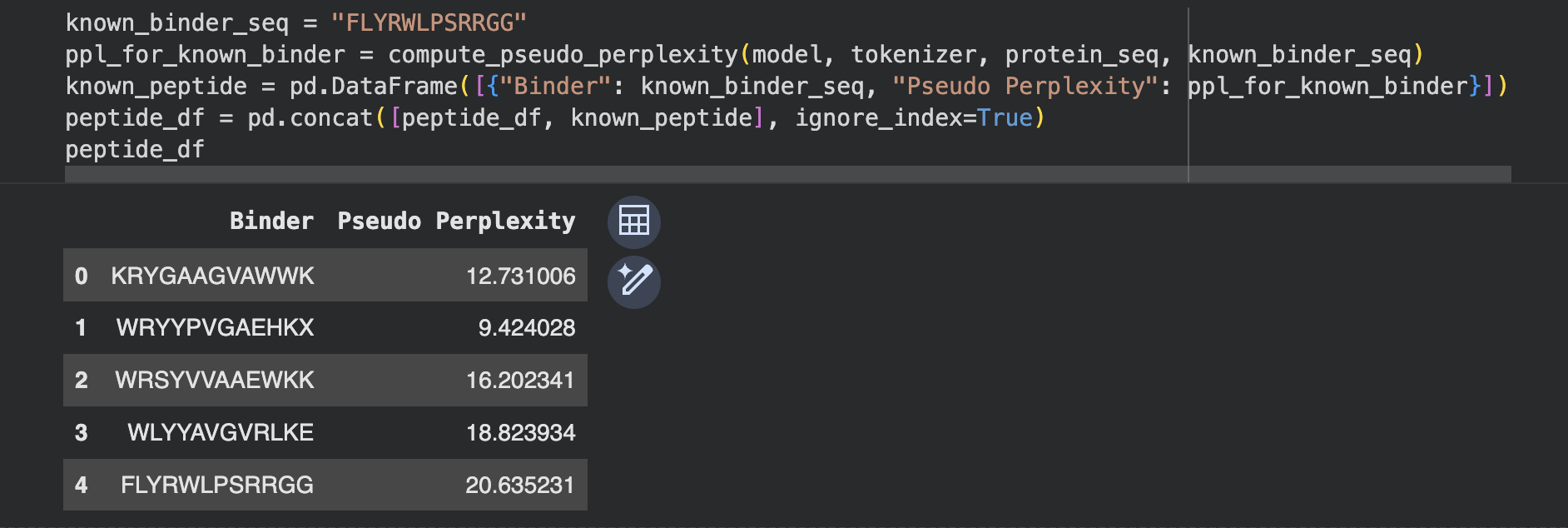
I generated 4 peptides conditioned on the SOD1-A4V sequence,
and added the known SOD1-binding peptide FLYRWLPSRRGG as a
positive control for comparison.
Perplexity indicates PepMLM’s confidence that a peptide will bind the target — lower scores indicate higher confidence.