Week 2 HW: DNA Read, Write, & Edit
DNA Design Challenge
1. Choose Your Protein
I chose Reflectin A1, the structural protein found in squid skin that is responsible for their ability to dynamically change color and iridescence. Squid iridophores contain layers of reflectin that can shift what wavelengths of light they reflect by changing the protein’s conformation — essentially biological tunable thin-film optics. I’m interested in this because it represents a living material that produces a dynamic optical output as a visual response through intrinsic material properties rather than added pigments. Researchers have fabricated reflectin-based thin films that change color in response to chemical signals, which sits right at the intersection of my interests in materials, sensing, and interaction.
Protein sequence from UniProt (D3UA43) / NCBI (ACZ57764.1):
>ACZ57764.1 reflectin-like protein A1 [Doryteuthis pealeii]
MNRYLNRQRLYNMYRNKYRGVMEPMSRMTMDFQGRYMDSQGRMVDPRYYDYYGRMHDHDRYYGRSMFNQG
HSMDSQRYGGWMDNPERYMDMSGYQMDMQGRWMDAQGRFNNPFGQMWHGRQGHYPGYMSSHSMYGRNMYN
PYHSHYASRHFDSPERWMDMSGYQMDMQGRWMDNYGRYVNPFNHHMYGRNMCYPYGNHYYNRHMEHPERY
MDMSGYQMDMQGRWMDTHGRHCNPFGQMWHNRHGYYPGHPHGRNMFQPERWMDMSGYQMDMQGRWMDNYG
RYVNPFSHNYGRHMNYPGGHYNYHHGRYMNHPERHMDMSSYQMDMHGRWMDNQGRYIDNFDRNYYDYHMY2. Reverse Translate
Using NCBI, I traced the protein back to its original DNA coding sequence: FJ824804.1 — the mRNA for Reflectin-like protein A1 from Doryteuthis pealeii.
>FJ824804.1 Loligo pealei reflectin-like protein A1 mRNA, complete cds
ATGAATCGATATCTGAATCGACAGCGCCTGTACAACATGTACAGAAACAAGTACCGAGGTGTGATGGAAC
CGATGTCCAGAATGACCATGGACTTCCAAGGAAGATACATGGACTCCCAGGGTAGAATGGTCGACCCCAG
ATACTACGACTACTACGGAAGAATGCACGACCATGACCGATACTACGGAAGGTCCATGTTCAACCAGGGA
CACAGCATGGACAGTCAACGCTACGGCGGCTGGATGGACAACCCCGAGAGGTACATGGACATGTCTGGCT
ACCAGATGGACATGCAGGGACGCTGGATGGACGCCCAGGGGCGATTCAACAACCCGTTCGGTCAGATGTG
GCACGGCAGGCAAGGCCATTACCCTGGTTACATGTCATCTCACTCCATGTATGGTAGAAATATGTATAAC
CCCTACCACAGCCATTACGCCAGCCGGCATTTCGATTCCCCCGAGAGATGGATGGACATGTCCGGCTATC
AGATGGACATGCAAGGACGCTGGATGGATAACTACGGCCGTTACGTGAACCCGTTCAACCACCACATGTA
TGGCAGAAACATGTGTTATCCTTACGGCAACCATTACTACAATCGGCACATGGAGCACCCCGAGAGATAC
ATGGACATGTCCGGCTATCAGATGGACATGCAGGGACGCTGGATGGACACACACGGACGCCACTGCAACC
CGTTCGGTCAGATGTGGCACAACAGGCACGGTTACTATCCAGGACACCCACATGGTCGCAACATGTTCCA
GCCCGAAAGATGGATGGATATGTCCGGCTATCAGATGGACATGCAAGGACGCTGGATGGATAACTATGGC
CGTTATGTGAACCCGTTCAGTCATAACTACGGCAGGCATATGAACTACCCTGGAGGTCACTACAACTACC
ACCACGGCCGCTACATGAATCACCCCGAGAGACACATGGACATGTCCAGCTATCAGATGGACATGCACGG
ACGCTGGATGGACAACCAGGGCCGTTATATTGACAATTTCGATAGAAATTATTACGACTATCACATGTAT
TAAThe sequence starts with ATG (start codon) and ends with TAA (stop codon). It’s 1,053 base pairs long, which checks out: 350 amino acids × 3 bases per codon = 1,050, plus 3 for the stop codon.
3. Codon Optimization
I optimized for E. coli because that’s the organism we’re using in the lab to express proteins. The original DNA sequence comes from squid, which uses a different set of preferred codons than E. coli. Even though multiple codons can encode the same amino acid, cells have different amounts of the tRNA molecules that read each codon. E. coli has abundant tRNA for its commonly-used codons and very little for rare ones. If I put the original squid DNA into E. coli, the ribosome would stall at codons that E. coli rarely uses, resulting in slow or unsuccessful protein production. Codon optimization swaps those squid codons for ones E. coli translates quickly, so it can produce it much more efficiently.
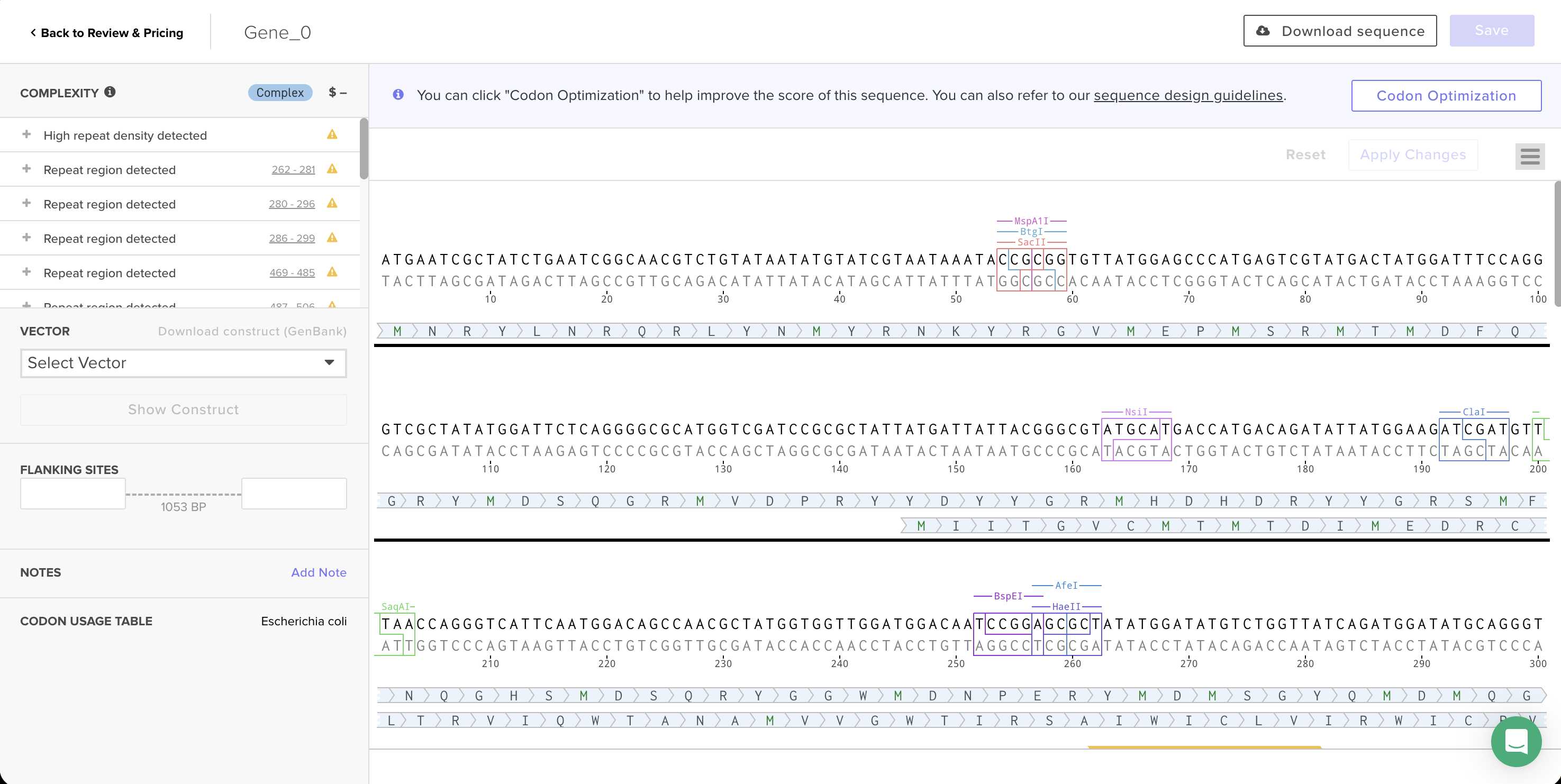
 Codon-optimized sequence in Twist’s editor, with Codon Usage Table set to Escherichia coli.
Codon-optimized sequence in Twist’s editor, with Codon Usage Table set to Escherichia coli.
ATGAATCGCTATCTGAATCGGCAACGTCTGTATAATATGTATCGTAATAAATACCGCGGTGTTATGGAGC
CCATGAGTCGTATGACTATGGATTTCCAGGGTCGCTATATGGATTCTCAGGGGCGCATGGTCGATCCGCG
CTATTATGATTATTACGGGCGTATGCATGACCATGACAGATATTATGGAAGATCGATGTTTAACCAGGGT
CATTCAATGGACAGCCAACGCTATGGTGGTTGGATGGACAATCCGGAGCGCTATATGGATATGTCTGGTT
ATCAGATGGATATGCAGGGTCGTTGGATGGACGCCCAGGGACGGTTCAATAACCCATTCGGACAGATGTG
GCATGGTCGTCAGGGCCATTACCCGGGTTATATGTCCAGCCACTCAATGTATGGTAGAAACATGTACAAC
CCGTATCATAGTCATTATGCTAGCCGCCATTTTGATAGCCCTGAACGGTGGATGGATATGAGTGGGTACC
AAATGGATATGCAAGGTAGATGGATGGATAATTATGGACGGTACGTAAATCCTTTTAACCACCACATGTA
CGGTCGGAATATGTGCTATCCGTACGGCAACCACTATTACAACCGGCACATGGAACATCCGGAACGTTAT
ATGGATATGTCTGGTTACCAAATGGATATGCAAGGGCGCTGGATGGATACACATGGTCGCCATTGTAATC
CGTTTGGCCAGATGTGGCATAACAGACACGGGTACTATCCGGGTCATCCGCATGGCCGAAATATGTTTCA
GCCAGAACGTTGGATGGATATGAGTGGCTATCAAATGGATATGCAGGGGCGGTGGATGGACAATTACGGC
AGATACGTTAACCCATTCAGCCATAATTATGGTCGCCATATGAATTACCCGGGGGGTCATTATAACTATC
ACCATGGCCGGTATATGAACCACCCCGAAAGACACATGGACATGTCCTCATATCAGATGGATATGCATGGC
AGATGGATGGATAATCAAGGACGTTACATTGATAATTTTGATCGTAACTATTATGACTATCACATGTATT
AASynthesis complexity: Twist flagged this sequence as “Complex” with 17 warnings — high repeat density and multiple repeat regions throughout the sequence. Reflectin’s amino acid sequence is highly repetitive (the motif YMDMSGYQMDMQGRWMD appears several times), and the underlying protein repetition still creates patterns that are hard to synthesize even after optimization. This is the same challenge the TAs described in recitation about spider silk proteins.
4. Producing the Protein
To produce reflectin from this DNA sequence, I would use a cell-dependent method by putting the DNA into living E. coli and letting them produce it:
- Expression cassette: The coding sequence needs to be wrapped with a promoter (“start transcribing here”), a ribosome binding site (so the ribosome knows where to land on the mRNA), and a terminator (“stop transcribing”).
- Insert into a plasmid: The expression cassette gets placed into a circular plasmid, which also carries an origin of replication (so the bacteria copy it when they divide) and an antibiotic resistance gene (to filter).
- Transform into E. coli: Using heat shock — the bacteria are chilled, the plasmid DNA is added, then a brief heat pulse causes temporary openings in the cell membrane allowing plasmids to enter. Cooled again to close the membrane.
- Grow: The bacteria are plated on media with antibiotics so only the ones carrying the plasmid (and its resistance gene) survive.
- Transcription and translation: The E. coli’s own RNA polymerase reads the DNA and creates mRNA (transcription). Then the cell’s ribosomes read the mRNA codons and build the reflectin protein chain using tRNA (translation).
- Protein: The bacteria are broken open to release the protein inside.
Prepare a Twist DNA Synthesis Order
1. Build DNA Insert Sequence
I built an expression cassette in Benchling by stitching together all the regulatory parts needed for E. coli to read and produce my reflectin protein. The parts, in order:
| Part | Description |
|---|---|
| Promoter (BBa_J23106) | Constitutive promoter — always on |
| RBS (BBa_B0034) | Ribosome binding site — tells ribosome where to land |
| Start Codon (ATG) | Begin translation |
| Reflectin A1 (codon-optimized) | My coding sequence from previous section |
| His Tag (7×His) | For protein purification |
| Stop Codon (TAA) | End translation |
| Terminator (BBa_B0015) | Stop transcription |
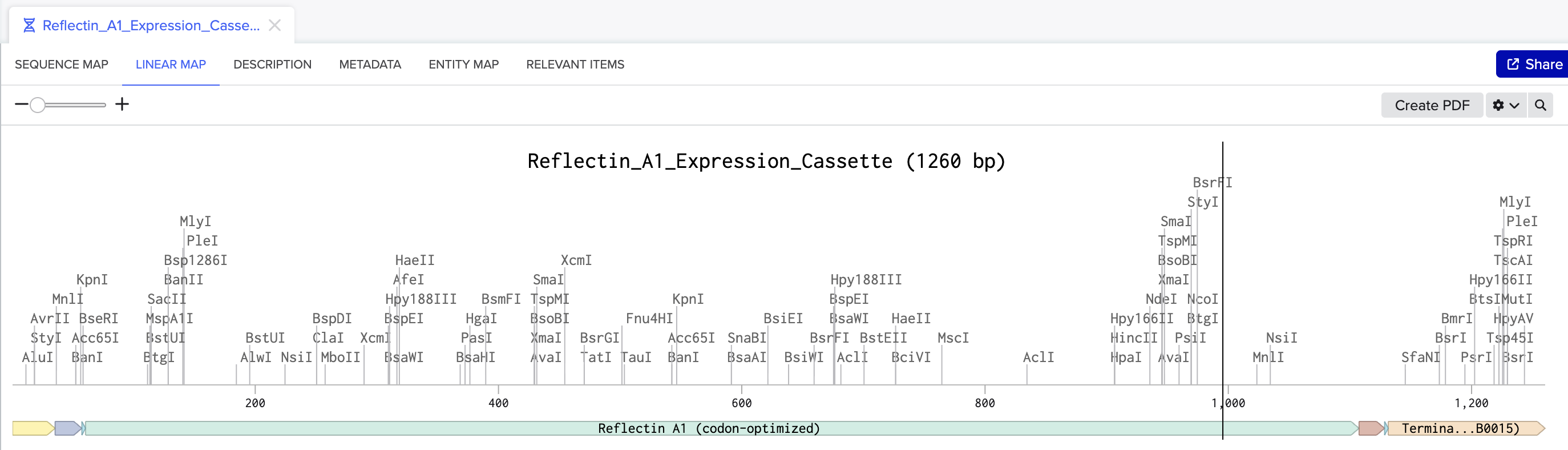
 Linear map of the Reflectin A1 expression cassette in Benchling (1,260 bp), showing all annotated parts.
Linear map of the Reflectin A1 expression cassette in Benchling (1,260 bp), showing all annotated parts.
Benchling sequence link | Download FASTA | Download GenBank
2. Upload to Twist & Choose Vector
I exported the expression cassette as a FASTA file and uploaded it to Twist as a Clonal Gene using the nucleotide sequence import option. I selected pTwist Amp High Copy as the backbone vector, which provides ampicillin resistance and a high-copy origin of replication. Twist inserts the expression cassette into this backbone to complete the circular plasmid.
I downloaded the full construct as a GenBank file and imported it back into Benchling:
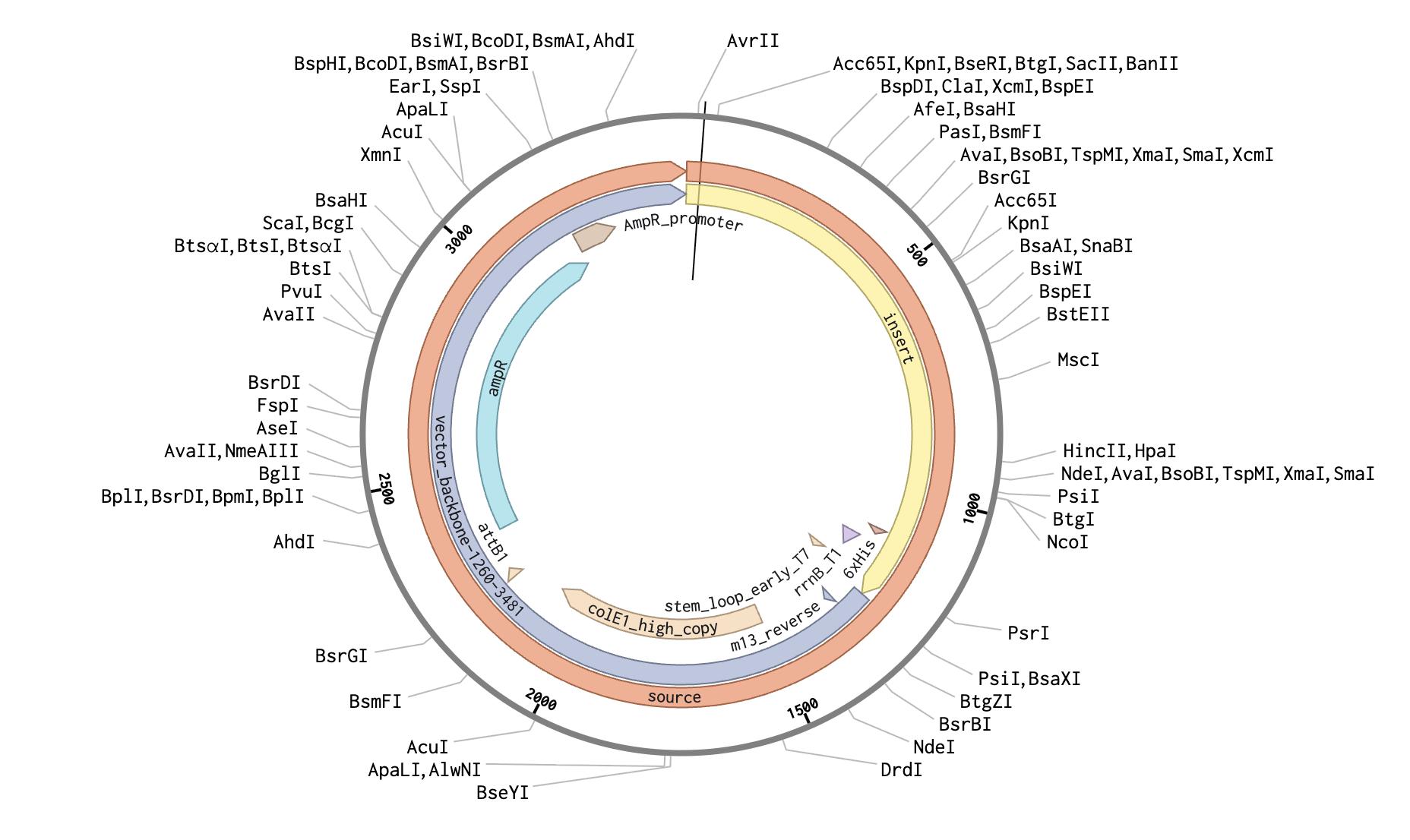
 The complete plasmid in Benchling - Reflectin A1 expression cassette inserted into the pTwist Amp High Copy backbone.
The complete plasmid in Benchling - Reflectin A1 expression cassette inserted into the pTwist Amp High Copy backbone.
DNA Read/Write/Edit
1. DNA Read
What DNA would I want to sequence and why?
I want to sequence DNA that is used for digital data storage — DNA molecules encoding binary information like images, text, or other digital files. The concept is taking digital data (0s and 1s), mapping it onto DNA bases (A, T, G, C), synthesizing those sequences, and storing them physically. DNA is incredibly dense as a storage medium and it lasts without power. George Church’s lab demonstrated this in 2012 by encoding an entire book into DNA.
I am interested in the reading-back problem. Storing data in DNA is only useful if we can reliably retrieve it later, which means sequencing the DNA and decoding it back into the original binary. With the right sequencing technology for retrieval, DNA data storage could become practical for things like archiving cultural heritage, scientific datasets, or even personal data.
What technology would I use?
I would use nanopore sequencing, such as Oxford Nanopore’s MinION device. For DNA data storage, portability matters so that we could read data anywhere outside of just biofacilities.
This is a third-generation sequencing technology: it reads single molecules of DNA directly, without needing to copy the DNA first that rely on PCR amplification which can introduce copying errors.
Input and preparation: The input is the DNA sample containing the encoded data.
- Library preparation: Attach adapter sequences to the ends of the DNA molecules. They are short DNA sequences with a motor protein attached that guides the strand into the nanopore.
- No PCR amplification needed: Nanopore sequencing reads native single molecules, so you skip the amplification step that other methods require. This is faster and avoids errors introduced by copying.
- Load onto the flow cell: Pipette the prepared library onto the MinION flow cell, which contains an array of nanopore proteins embedded in a membrane.
Base calling: Each nanopore is a tiny protein channel in an electrically charged membrane. A voltage drives ionic current through the pore, and when a DNA strand gets fed through by a motor protein, each base (A, T, G, C) blocks the current slightly differently because of its unique size and charge. The device measures these current fluctuations in real time, and a machine learning algorithm figures out the sequence from the pattern of electrical shadow each base casts as it squeezes through a tiny hole.
Output: The outputs are long reads of continuous sequences that can be tens of thousands to millions of bases long and they show up in real time, so we can start seeing results within minutes. For DNA data storage, these reads would be decoded back into binary using whatever encoding scheme was used to write the data.
2. DNA Write
What DNA would I want to synthesize and why?
I want to synthesize a mechanosensitive genetic circuit, a system where living cells can detect physical pressure and produce a visible response. The core component would be MscL (Mechanosensitive Channel of Large Conductance), a protein found in E. coli that acts as a pressure-activated gate in the cell membrane. When the cell experiences mechanical force, MscL opens and allows ions to flow through, which can be wired to trigger gene expression.
The full circuit would involve multiple genes working together: MscL as the sensor, a signal transduction pathway that converts the mechanical input into a transcriptional response, and a fluorescent reporter (like sfGFP) as the visible output. When the cells experience pressure, they glow. What excites me about this is that it’s essentially a biological touch sensor but with living cells instead of electronics. Embedding these cells in a material creates a surface that visually responds to touch.
The DNA I’d need to synthesize would include: the MscL gene (codon-optimized for E. coli), a synthetic promoter responsive to the downstream signal, the sfGFP reporter gene, and the necessary regulatory parts (RBS, terminators).
What technology would I use?
I would use phosphoramidite chemical synthesis, which is the method companies like Twist Bioscience use. This is currently the standard for synthesizing custom DNA sequences.
Essential steps:
- Sequence design: Design the full multi-gene circuit digitally, codon-optimize for E. coli, and check for synthesis constraints (repeats, GC content, homopolymers).
- Oligonucleotide synthesis: Short single-stranded DNA fragments (~200 bases) are built one base at a time on a solid support. Each cycle adds one nucleotide through a chemical reaction, with about 99% efficiency per step.
- Assembly: The short oligos are assembled into longer sequences through overlapping ends.
- Cloning and verification: The assembled gene is inserted into a plasmid, transformed into E. coli, grown and then sequenced to verify.
Limitations:
- Length: Phosphoramidite chemistry has a practical limit of about 200 bases per oligo because the ~99% per-step yield compounds.
- Repetitive sequences: As I saw with reflectin, repetitive protein motifs create repetitive DNA that is hard to synthesize accurately.
- GC content: Sequences with very high or very low GC content are harder to synthesize because they affect the secondary structure of the DNA during the chemical process.
3. DNA Edit
What DNA would I want to edit and why?
I want to edit the MSTN gene (myostatin) in human muscle cells. Myostatin is a protein that signals muscles to stop growing once they reach a certain size. This makes sense developmentally, but as humans age, muscle mass naturally declines (sarcopenia), leading to frailty, falls, and loss of independence. By editing the myostatin gene, we could preserve muscle mass well into old age.
Myostatin knockouts have been demonstrated across multiple animal species. There are famously muscular “mighty mice” with the gene disabled, and Belgian Blue cattle naturally carry a myostatin mutation that gives them dramatically increased muscle mass. A few humans have been documented with natural myostatin mutations and they show unusually high muscle mass with no apparent negative health effects.
What technology would I use?
I would use CRISPR-Cas9, the most widely used gene editing system. It was adapted from a bacterial immune defense mechanism and has become the standard tool for making precise edits to DNA in living cells. It has two components: the Cas9 protein (molecular scissors that cut DNA) and a guide RNA (a short sequence that tells Cas9 exactly where to cut for gene editing, designed to match the target sequence).
Preparation and steps:
- Design a guide RNA (~20 nucleotides) matching a region of the MSTN gene, adjacent to a PAM sequence (short DNA motif that Cas9 requires to bind).
- Assemble the components: Cas9 protein + guide RNA into a ribonucleoprotein complex. RNP delivery is preferred for therapeutic use because it degrades quickly, reducing the risk of unintended edits.
- Deliver to muscle cells using a viral vector injected into muscle tissue, or lipid nanoparticles.
- Cas9 cuts the MSTN gene: Once inside the cell, the guide RNA directs Cas9 to the myostatin gene. Cas9 cuts both DNA strands at that spot.
- Cell repair disrupts the gene: The cell tries to repair the break using a process called non-homologous end joining (NHEJ), which often introduces small insertions or deletions at the cut site which disrupt the gene’s reading frame, effectively no longer producing functional myostatin.
Limitations:
- Off-target edits: The guide RNA might bind to similar sequences elsewhere in the genome and cause unintended cuts.
- Delivery: Getting CRISPR into enough muscle cells across the body is hard. Vectors have limited capacity and can trigger immune responses.
- Mosaic results: Not every cell gets edited, so you end up with a mix of edited and unedited cells. The effect depends on hitting enough of them.
- Irreversibility: Once knocked out, you can’t easily turn the gene back on if something goes wrong.
- Ethical and regulatory concerns: Editing healthy human genomes for enhancement (rather than treating disease) raises significant ethical questions.