Labs
Lab writeups:
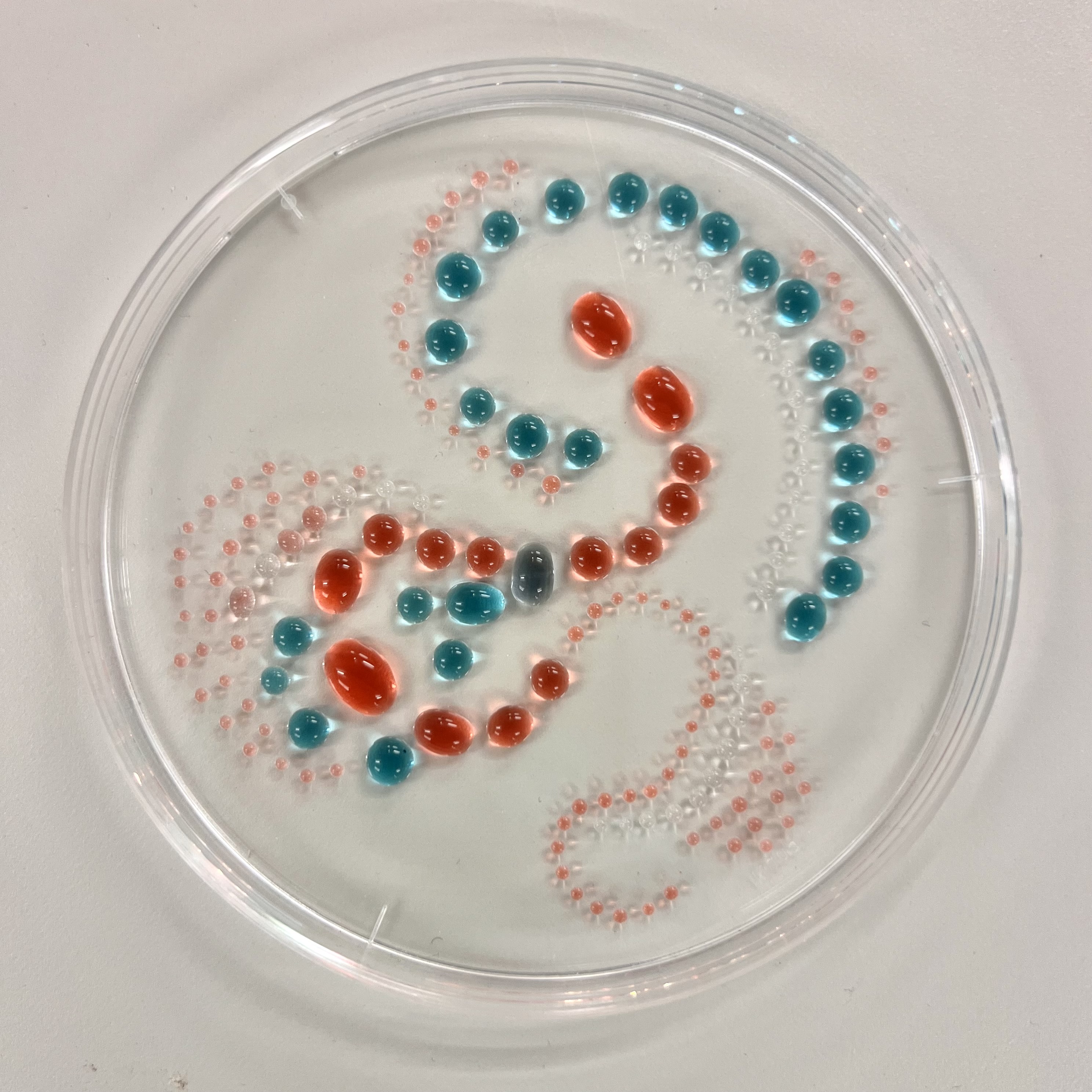
- Pepetting water droplet Art
- Restriction Digests and Gel Electrophoresis

- Lab Automation & Liquid Handling Robots
- PCR Mutagenesis, Gibson Assembly & Bacterial Transformation
- Intracellular Artificial Neural Networks in HEK293 Cells








Coming from a design background, I find it really interesting that DNA digestion patterns could be treated as a visual medium. I played around with Ronan’s gel art website trying different combination of digest results and I found some band patterns that generated a robot design. After I landed on the design, I used benchling to perform the virtual digest with the corresponding enzymes.

 Ronan’s website allowed me to scroll through different enzyme combinations and randomize to see what patterns came up, which was much faster for exploring visual patterns.
Ronan’s website allowed me to scroll through different enzyme combinations and randomize to see what patterns came up, which was much faster for exploring visual patterns.
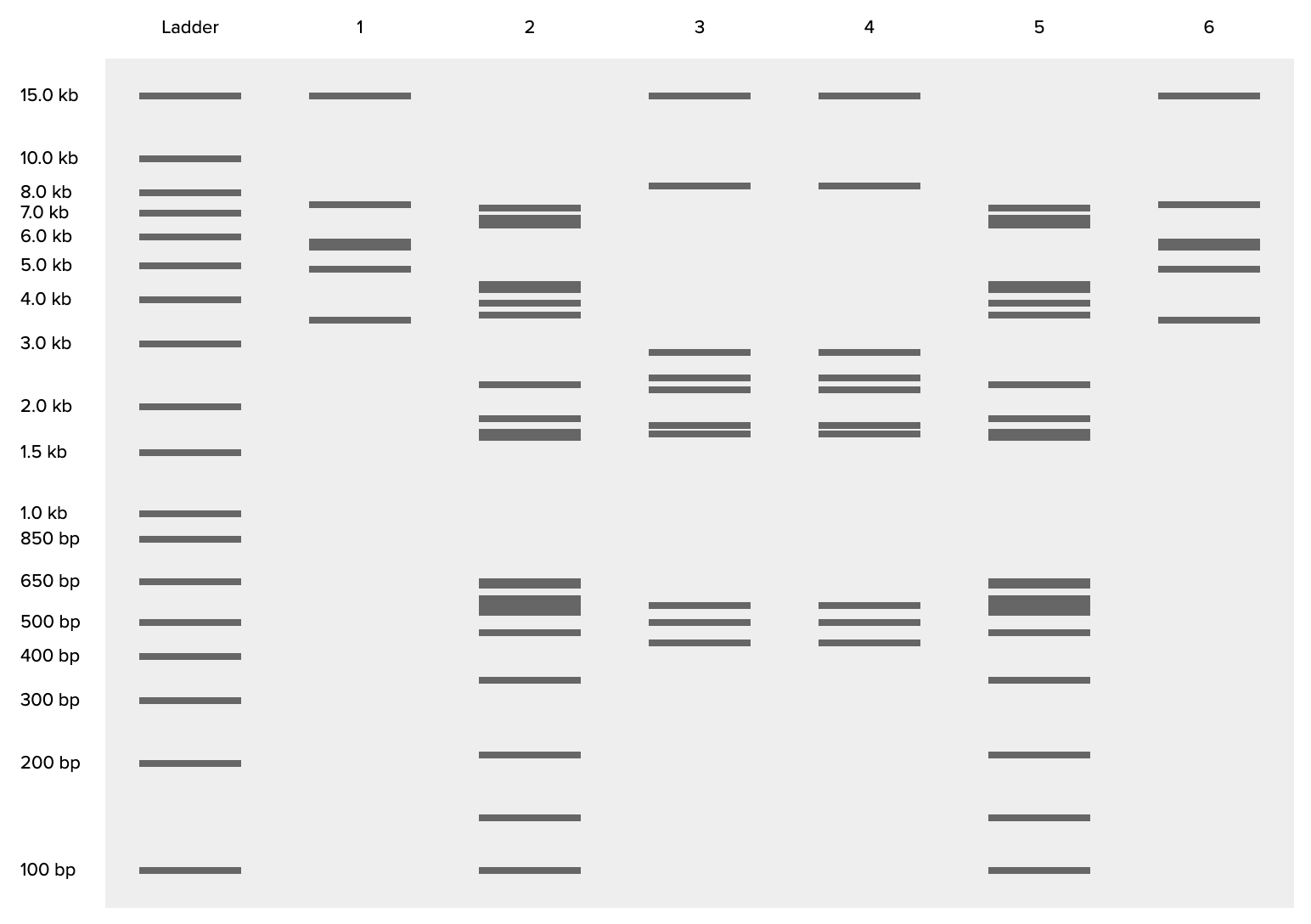

 Benchling virtual digest of my final 6-lane design, showing predicted band sizes for each enzyme combination against the 1 Kb Plus Thermo ladder.
Benchling virtual digest of my final 6-lane design, showing predicted band sizes for each enzyme combination against the 1 Kb Plus Thermo ladder.
My final design uses 6 lanes cut with different restriction enzyme combinations, mirrored across the center:


Each digest reaction was prepared in a PCR tube with the following components:
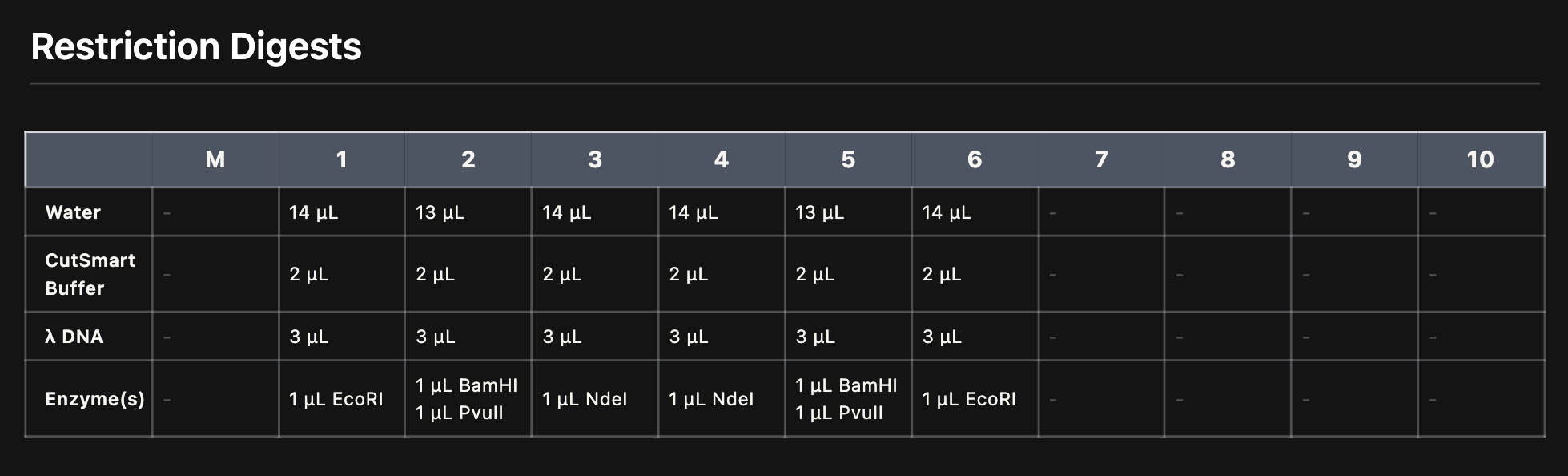
 *Reaction setup for each lane. Each tube contained water, CutSmart buffer, Lambda DNA, and the specific enzyme(s). *
*Reaction setup for each lane. Each tube contained water, CutSmart buffer, Lambda DNA, and the specific enzyme(s). *
The reactions were incubated at 37°C for 30 minutes to allow the enzymes to fully cut the DNA.
Step 1: Prepare the TAE buffer and agarose gel
We used 1× TAE buffer to mix with agarose powder. The flask was microwaved until the cloudy color became completely clear as the solution just bagan to have some bubbles.
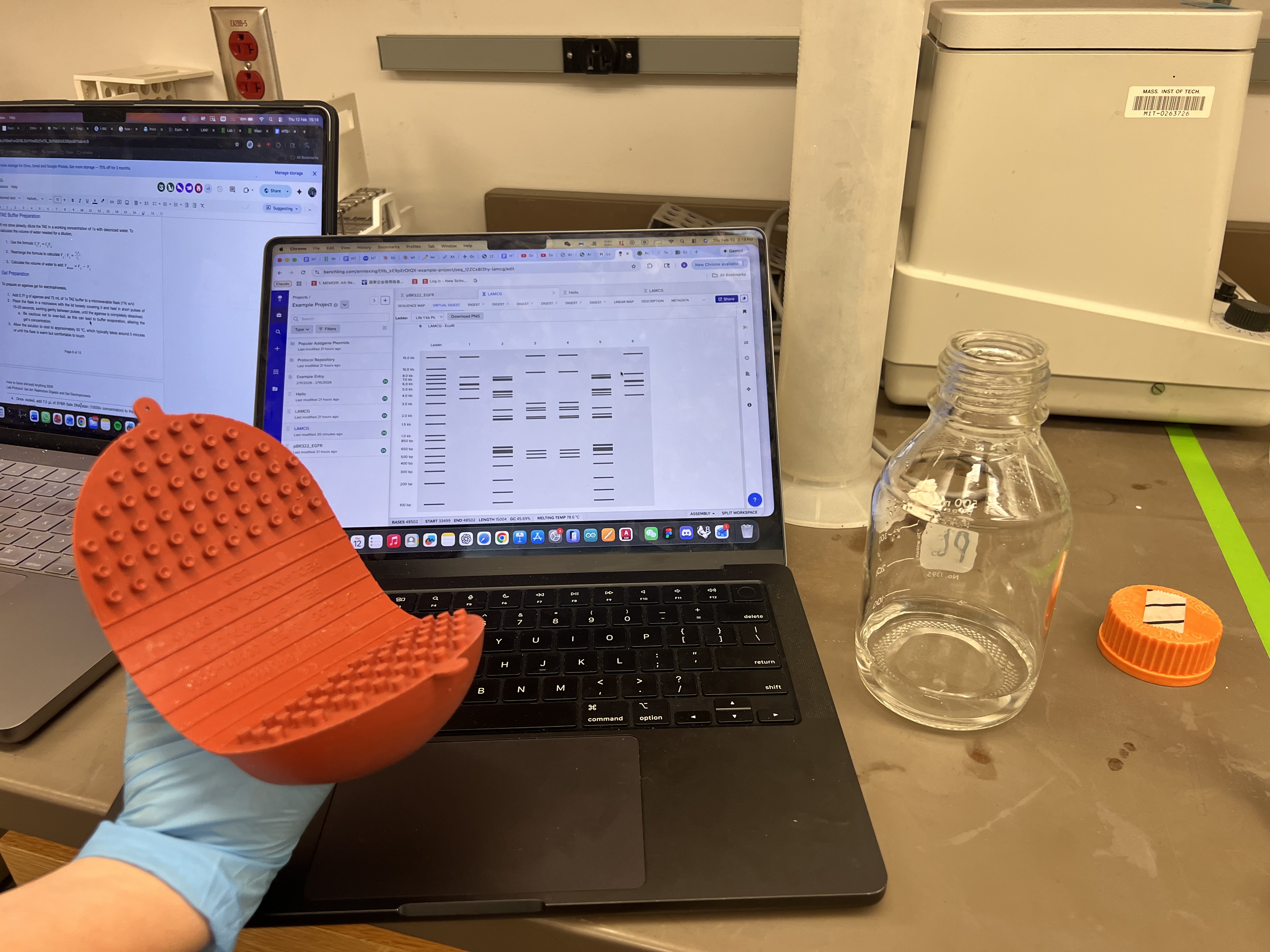

 Waiting for the solution to cool down
Waiting for the solution to cool down
Step 2: Add DNA stain and pour the gel
Once the agarose cooled slightly, we added a small amount of SYBR Safe DNA stain which made the solution tinted pink.
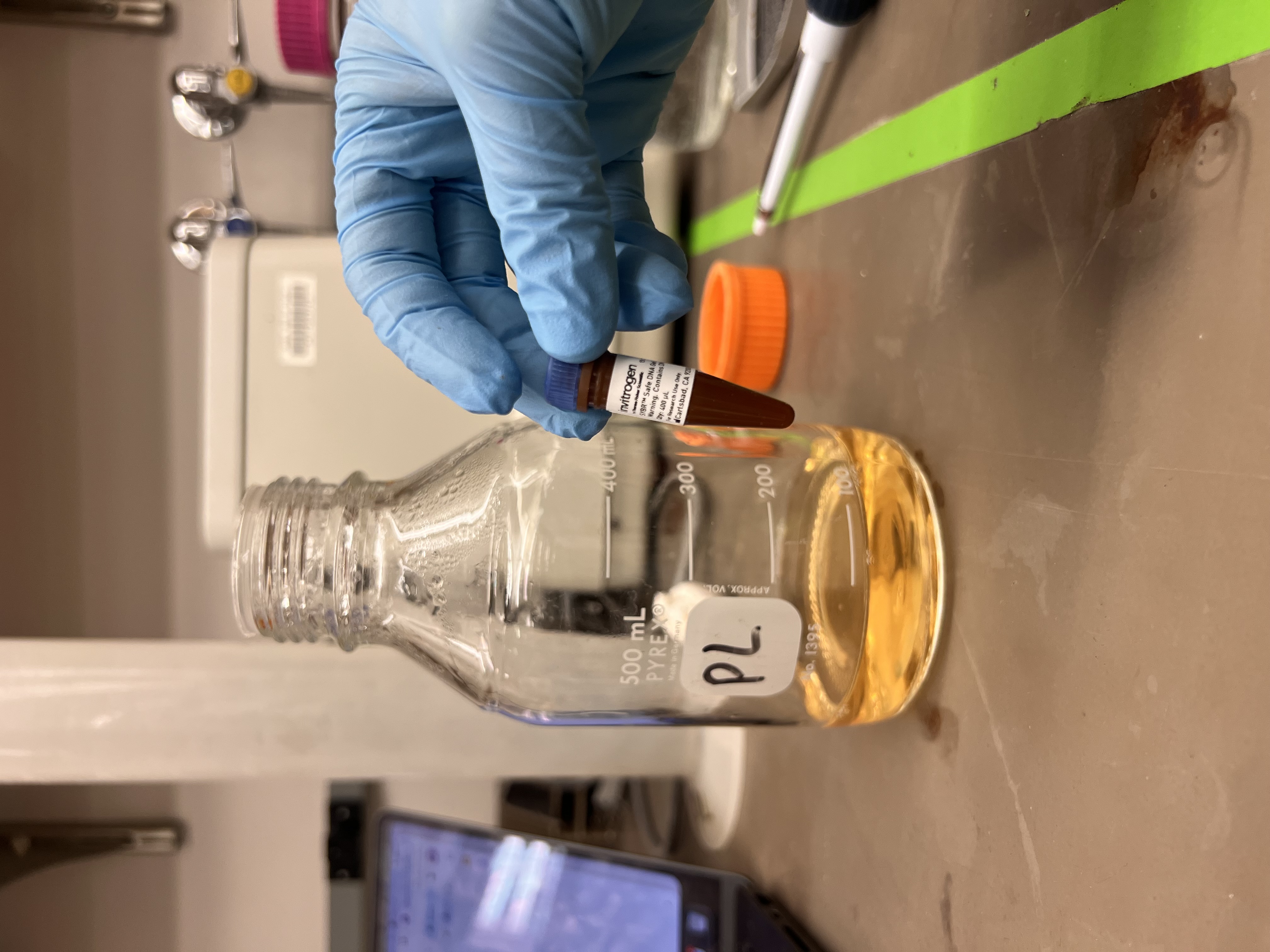

 The SYBR Safe stain next to the flask of melted agarose.
The SYBR Safe stain next to the flask of melted agarose.
We then poured the gel into the casting tray with the comb in place and waited for it to solidify.
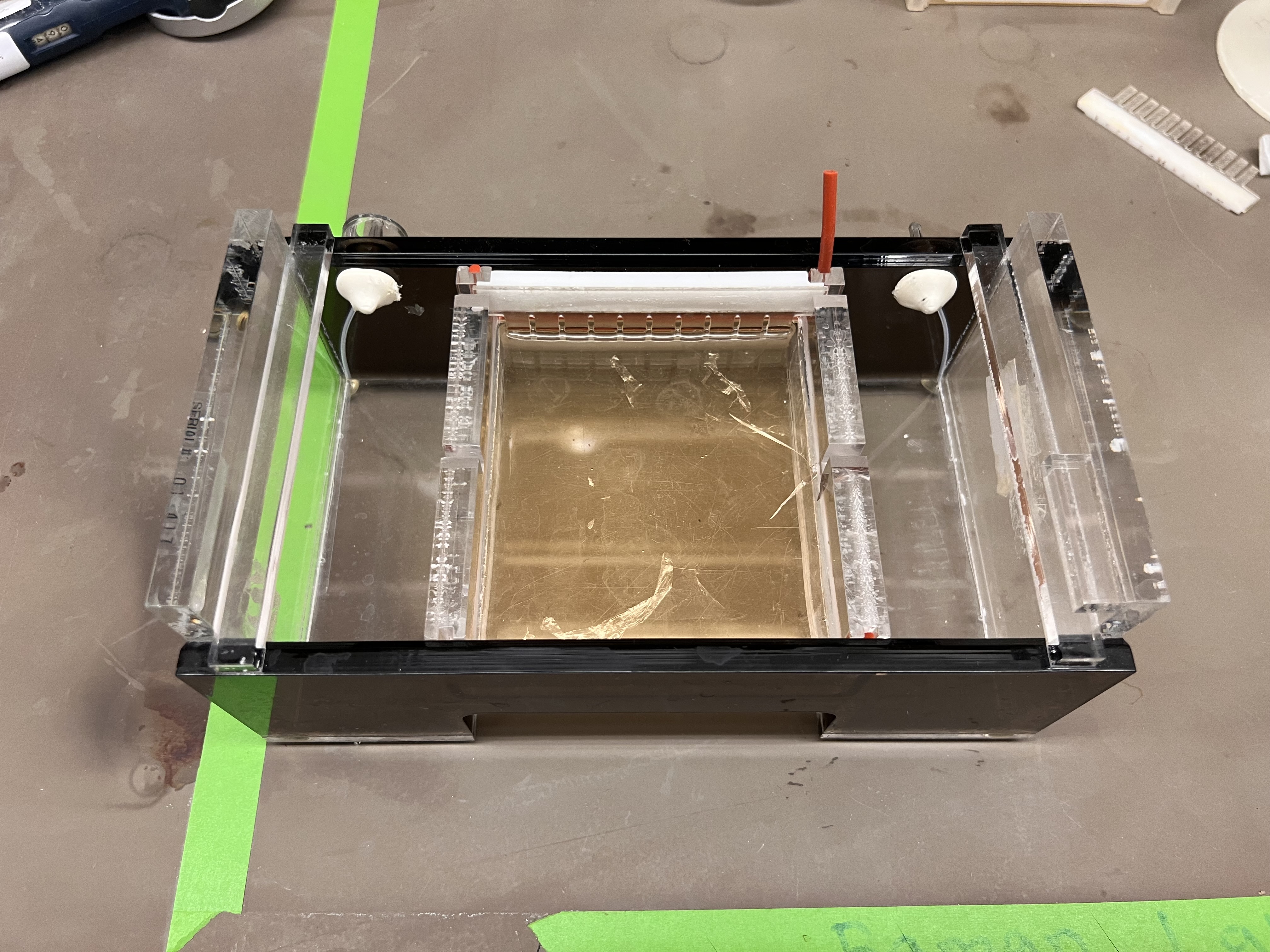

 The gel cast in the electrophoresis tray with the comb creating wells at the top.
The gel cast in the electrophoresis tray with the comb creating wells at the top.
Step 3: Prepare the digest reactions
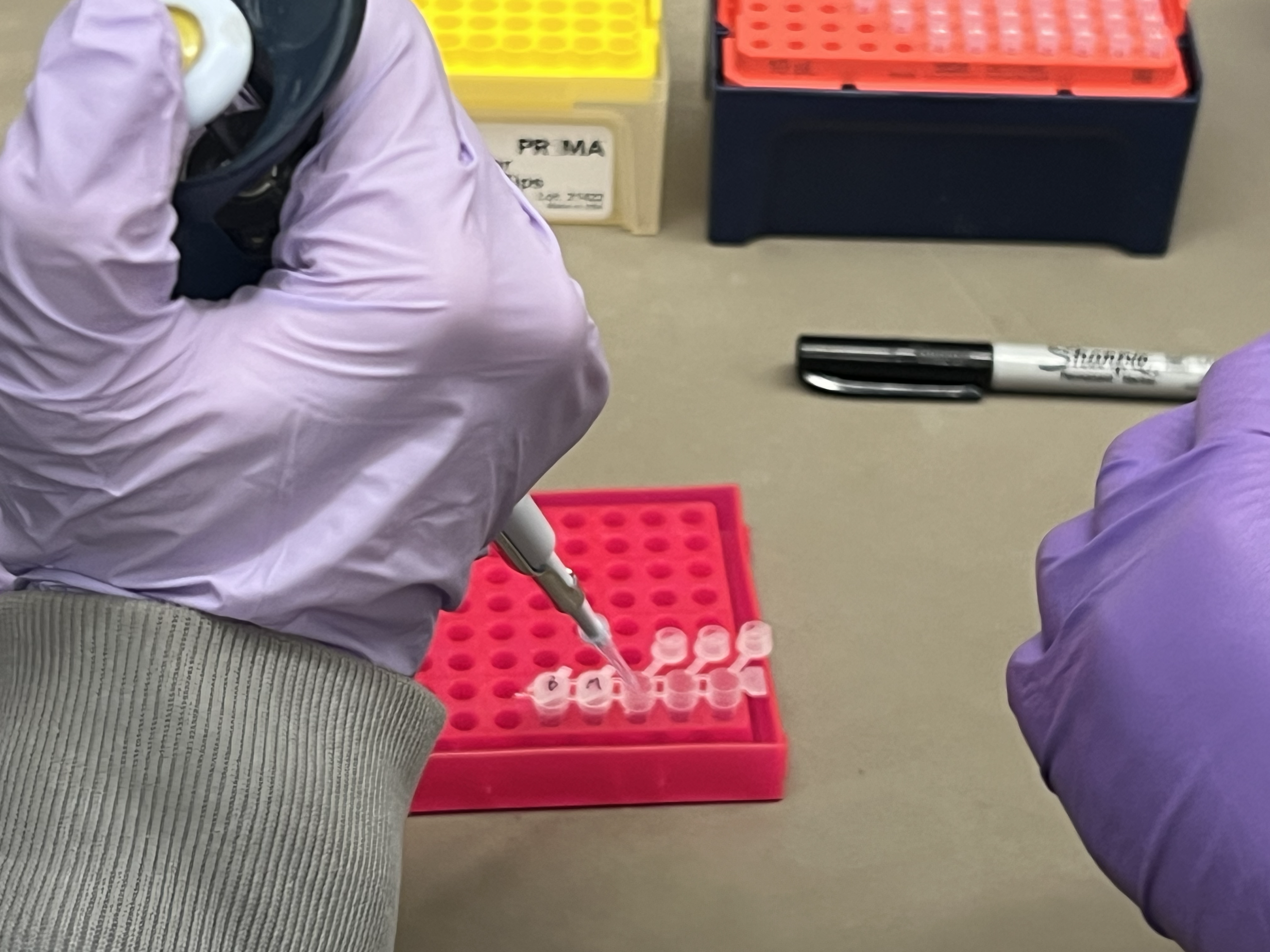
While waiting for the gel, we prepared the restriction digest reactions. Each tube was labeled 1-6 corresponding to our lane design. We pipetted water, CutSmart buffer, Lambda DNA, and the specific enzymes into each tube.
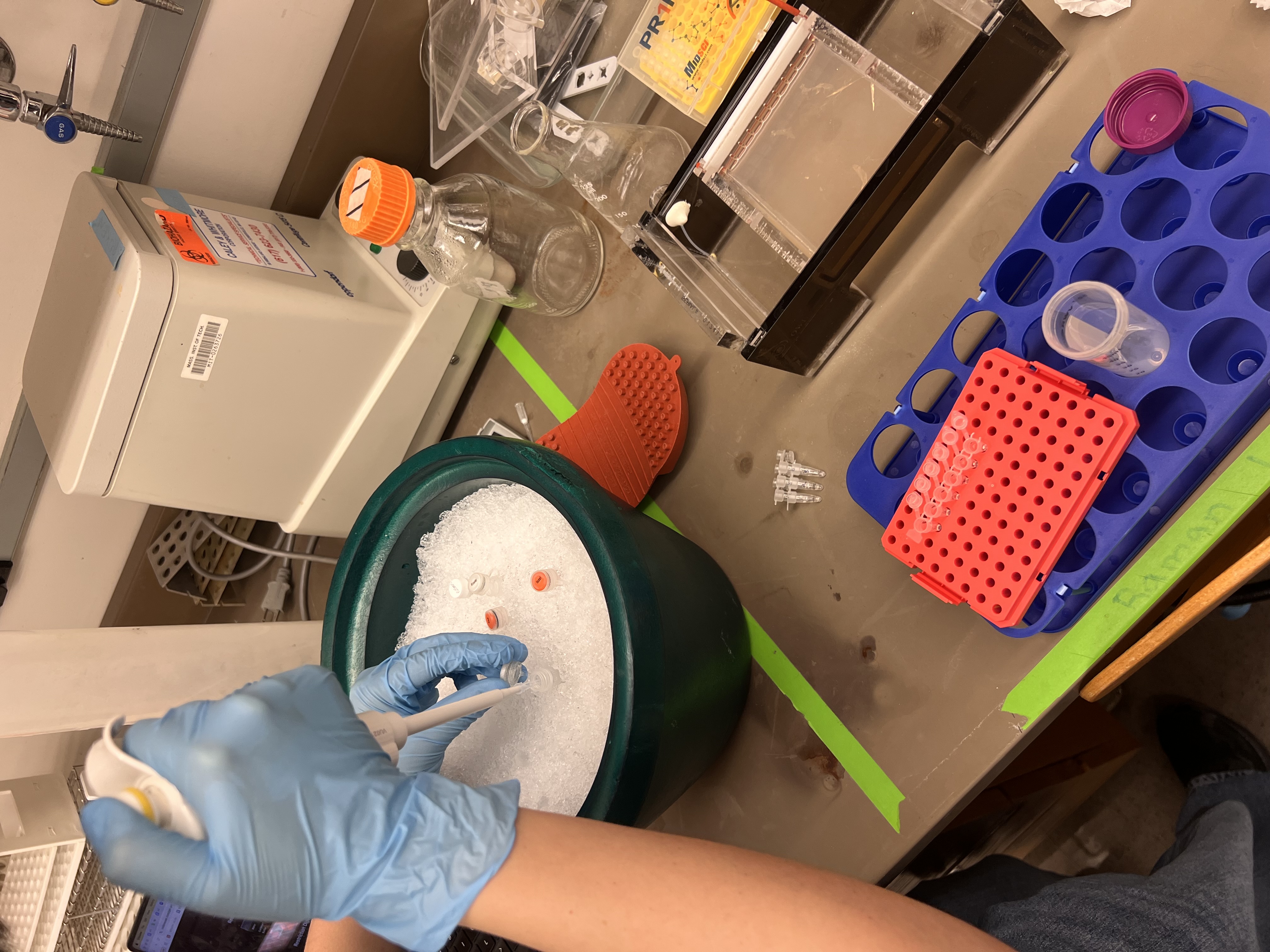
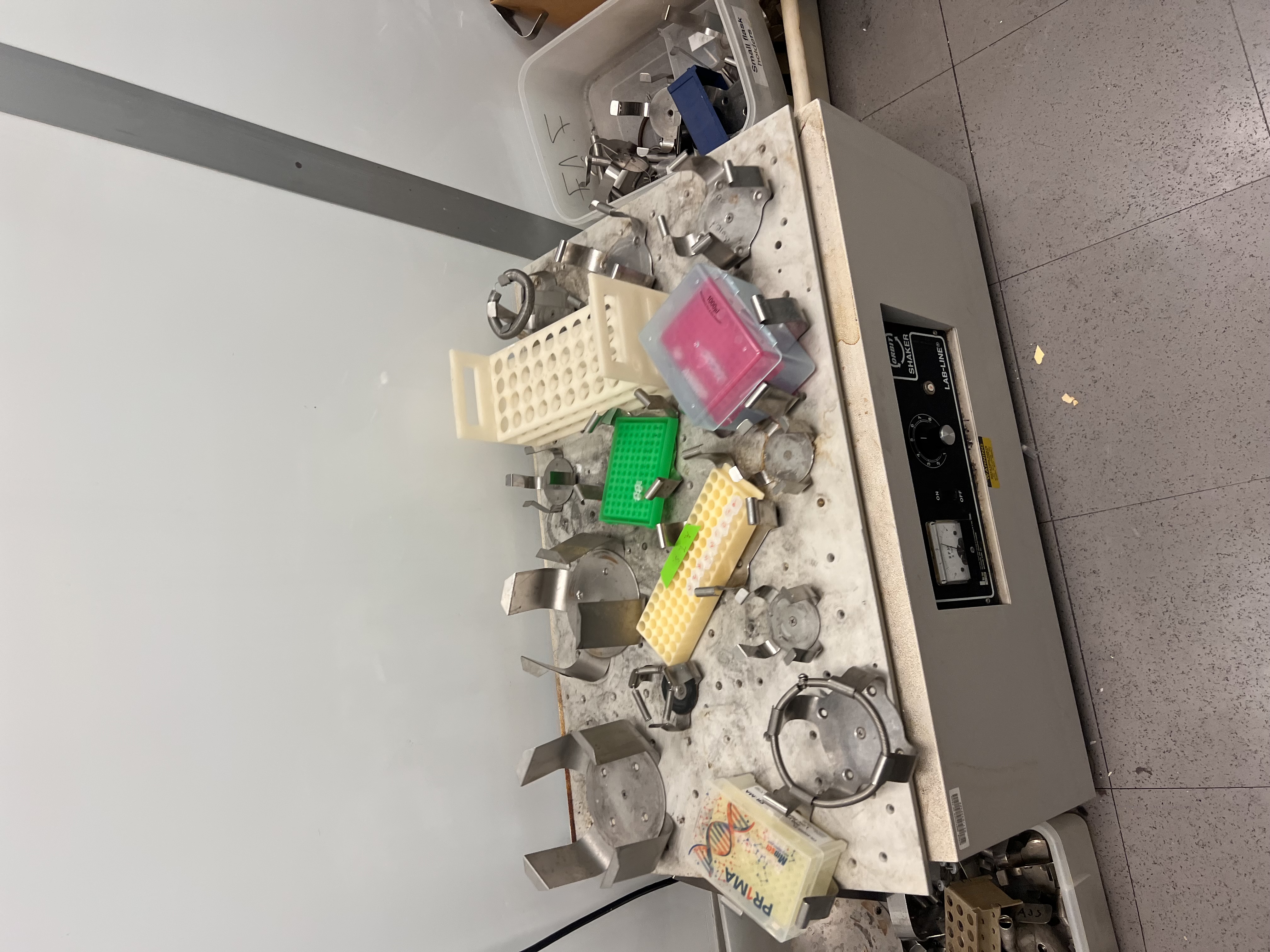
 Pipetting enzymes and Lambda DNA stored on ice.
Pipetting enzymes and Lambda DNA stored on ice.
This was the most challenging part because the micropipetting was tricky. It was hard to tell whether we had the right amount, since the volumes are so small they’re essentially invisible in the pipette tip.
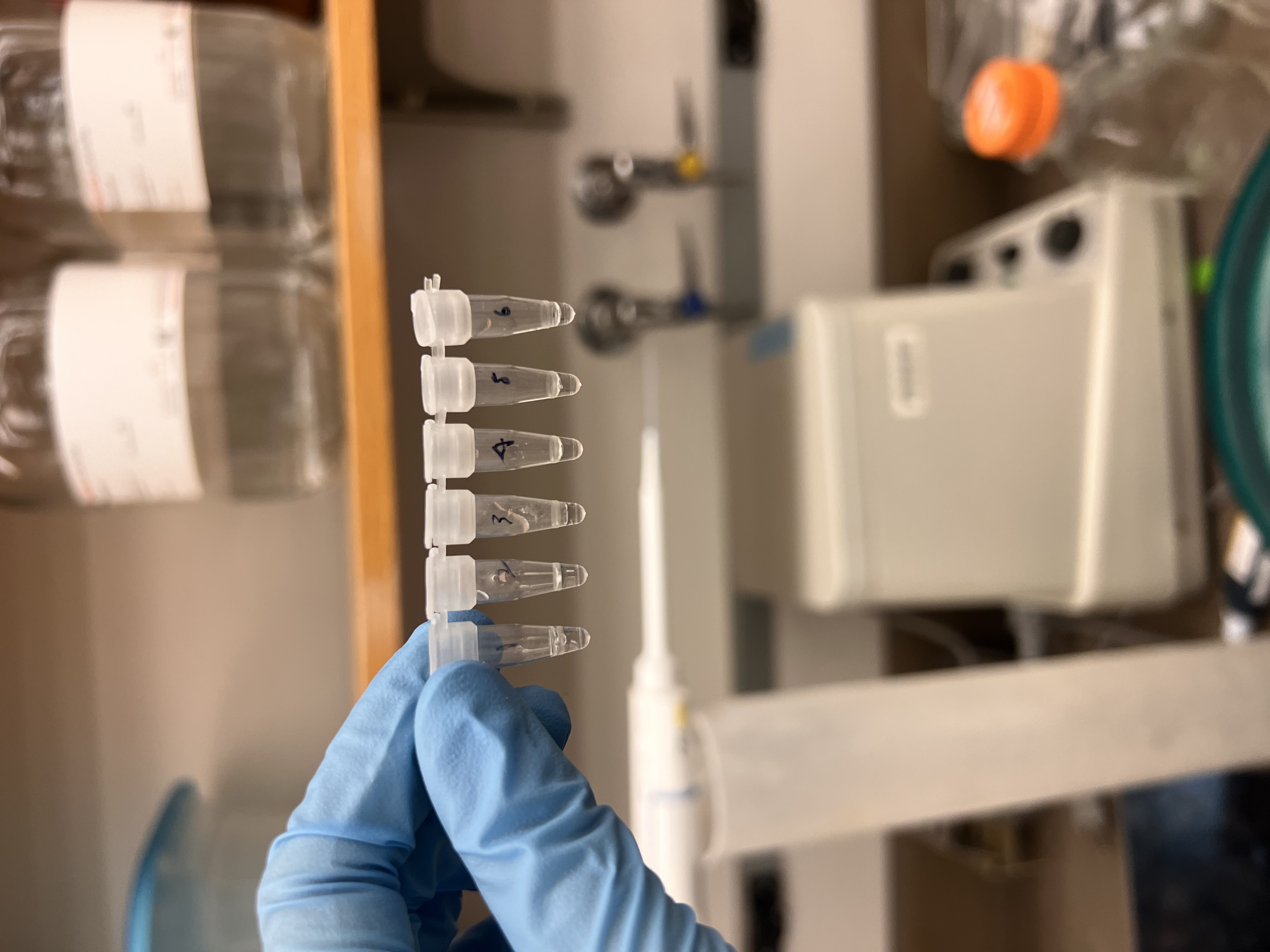

 Six labeled digest tubes ready for incubation.
Six labeled digest tubes ready for incubation.
Step 4: Add loading dye and load the gel
After incubation, we added loading dye to each reaction which would help us visually track how far the gel has run.
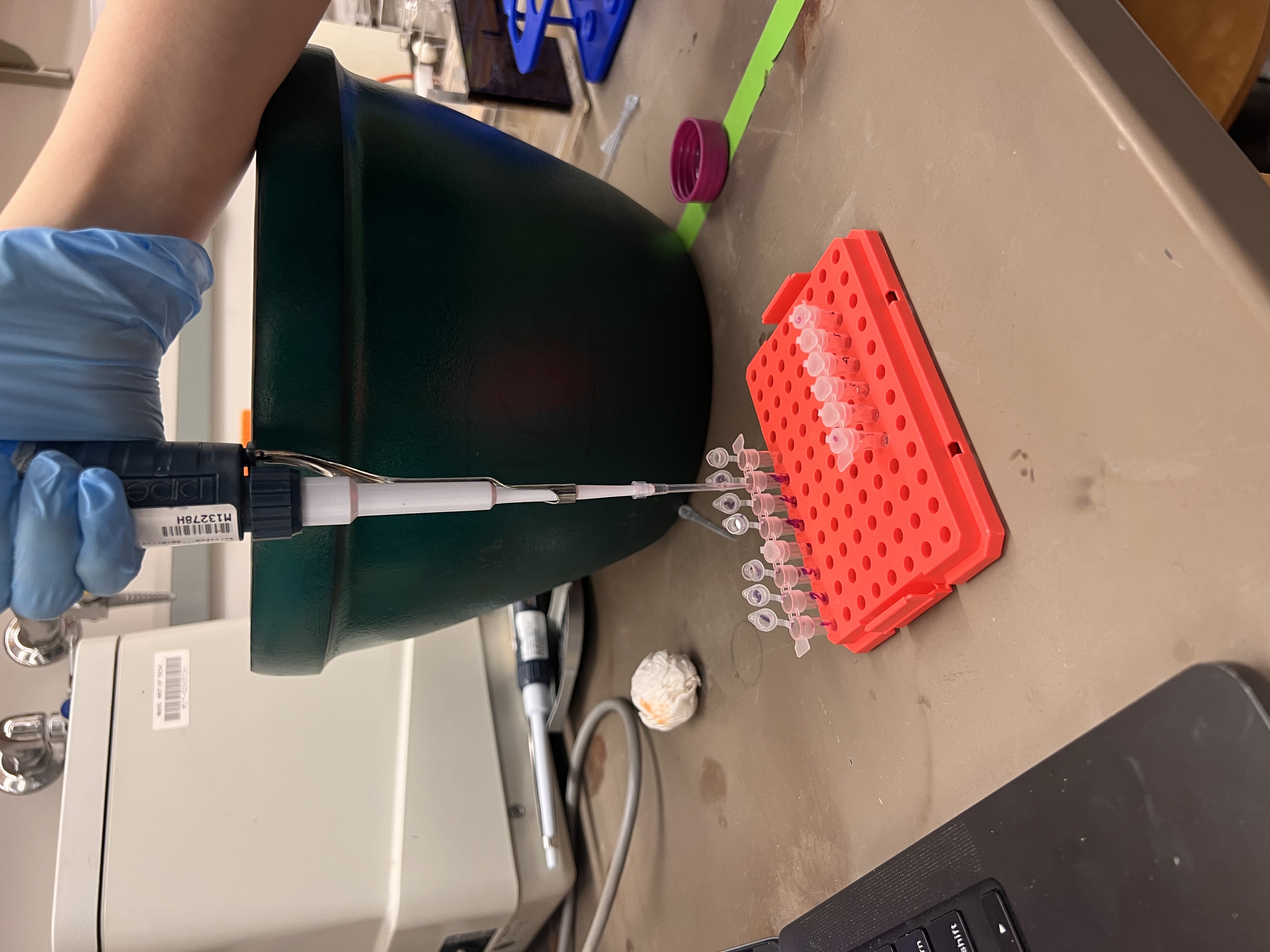

 Adding loading purple dye to the digest reactions before loading into the gel.
Adding loading purple dye to the digest reactions before loading into the gel.
We then filled casting wells with 1x TAE and loaded 20 uL of each samples
Step 5: Run the gel
Once loaded, we connected the electrodes and started running the gel. We noticed bubbles in the TAE buffer when the elctricity is on.
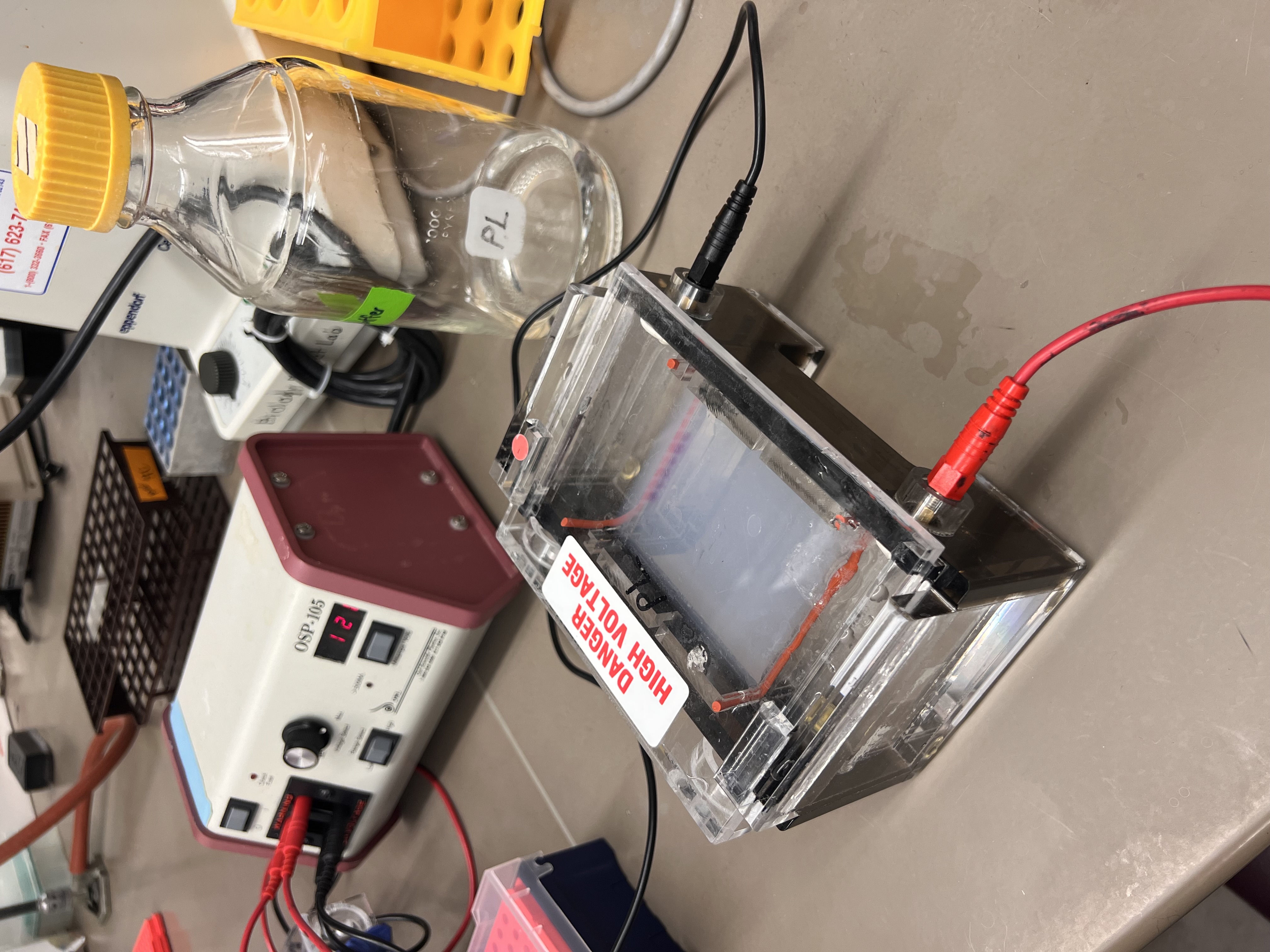

 The electrophoresis setup running.
The electrophoresis setup running.
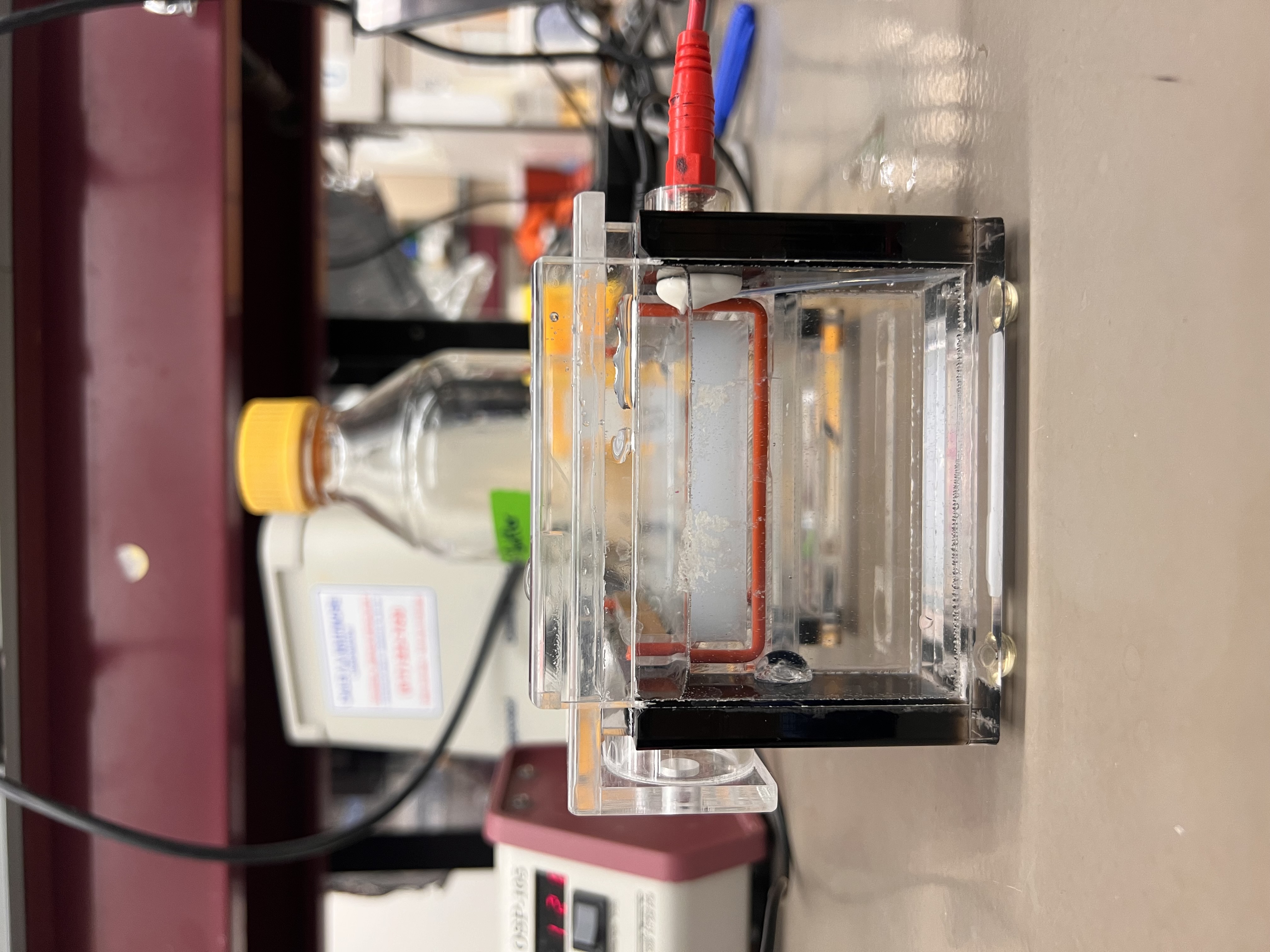

 Small bubbles rising to surface
Small bubbles rising to surface
We ran it for about 45 minutes to allow the DNA to travel more than halfway throught the gel.
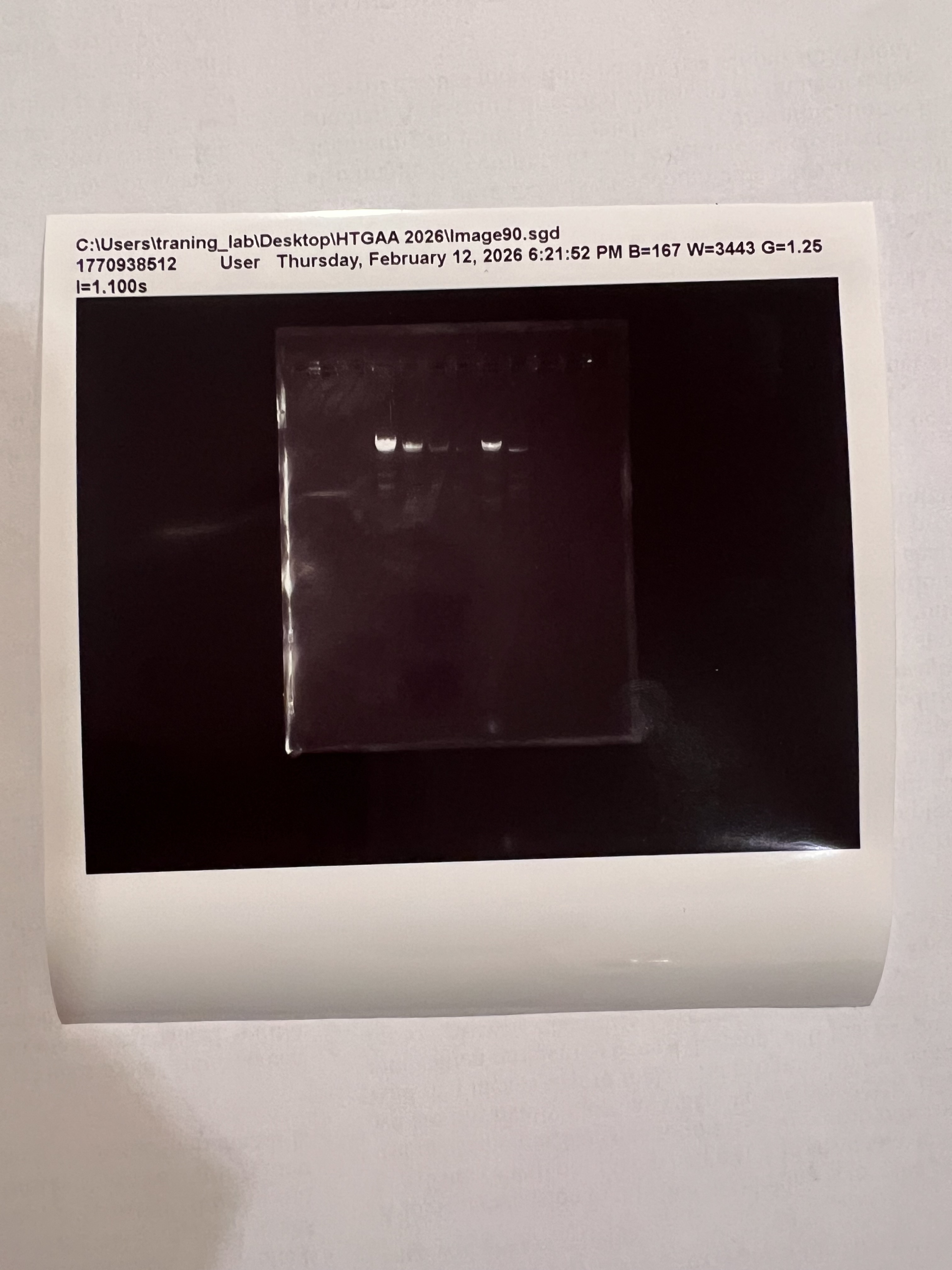
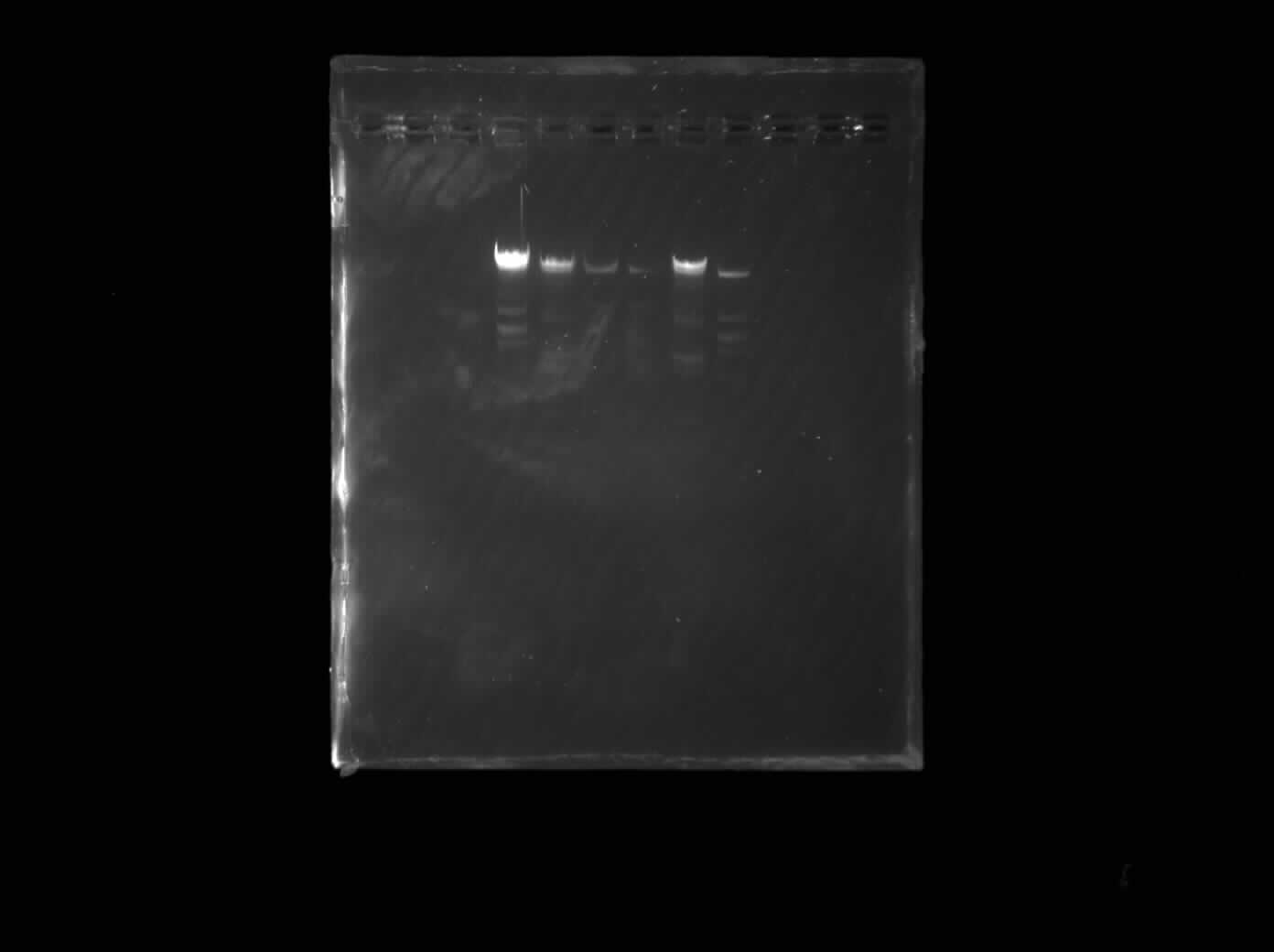
 Printed gel image from the lab imager
Printed gel image from the lab imager
The symmetric design concept came through but the overall image was fainter and the lower section was not visible. Possible reasons might be too little DNA samples and shorter runtimes.
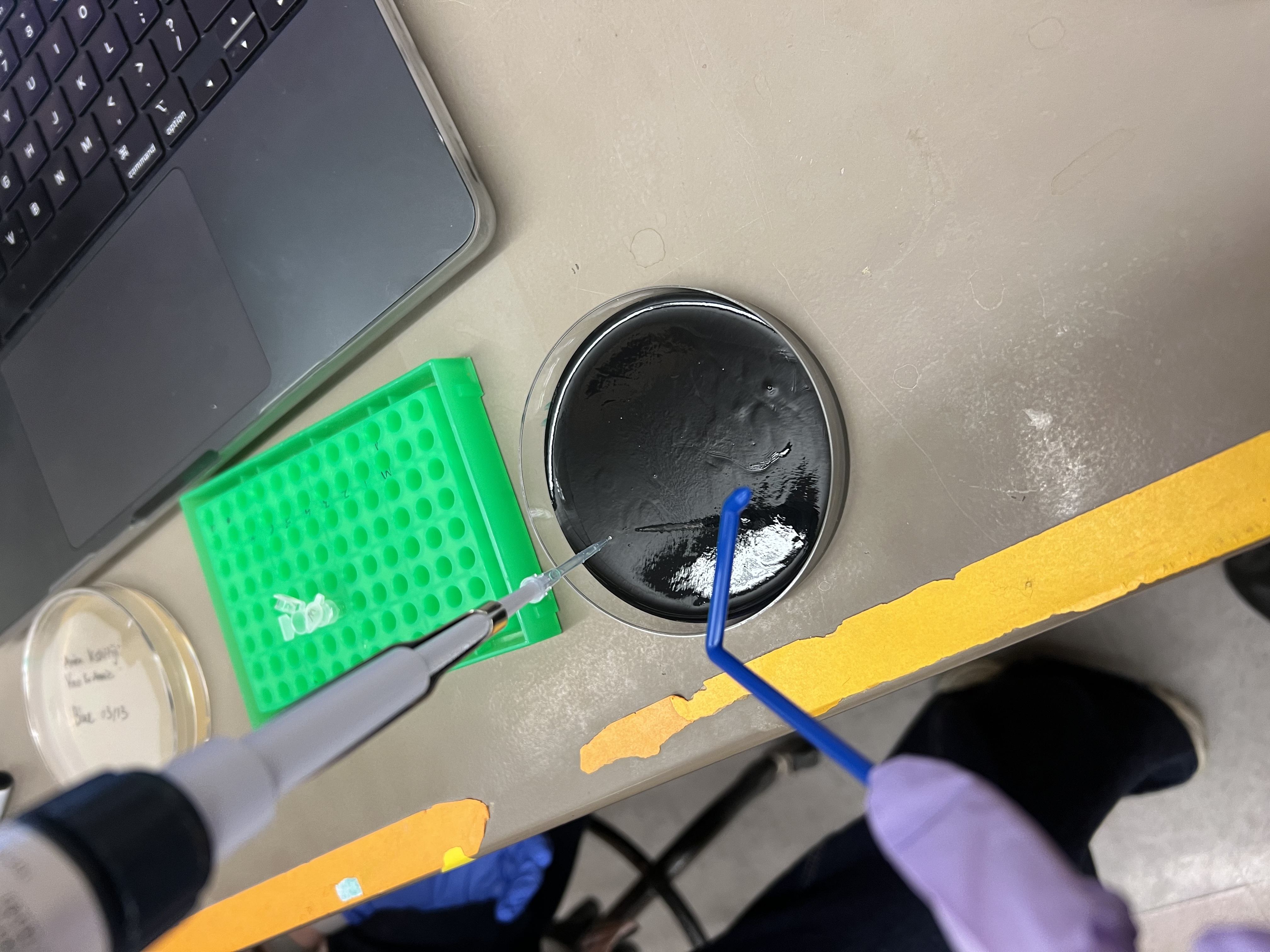
I designed a mandala-inspired symmetrical pattern using the Opentrons Art Interface, drawing on the visual language of radial symmetry. The script was exported from the GUI and ran on the Opentrons software
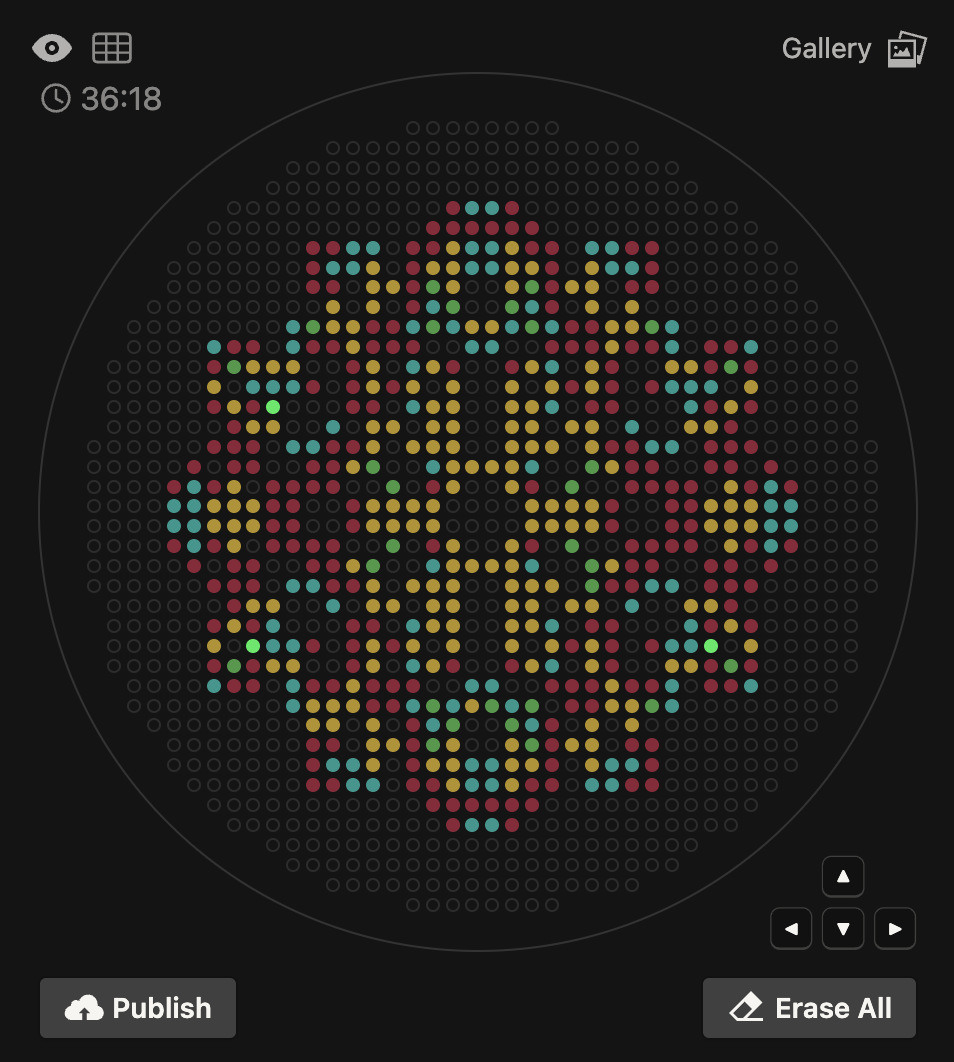

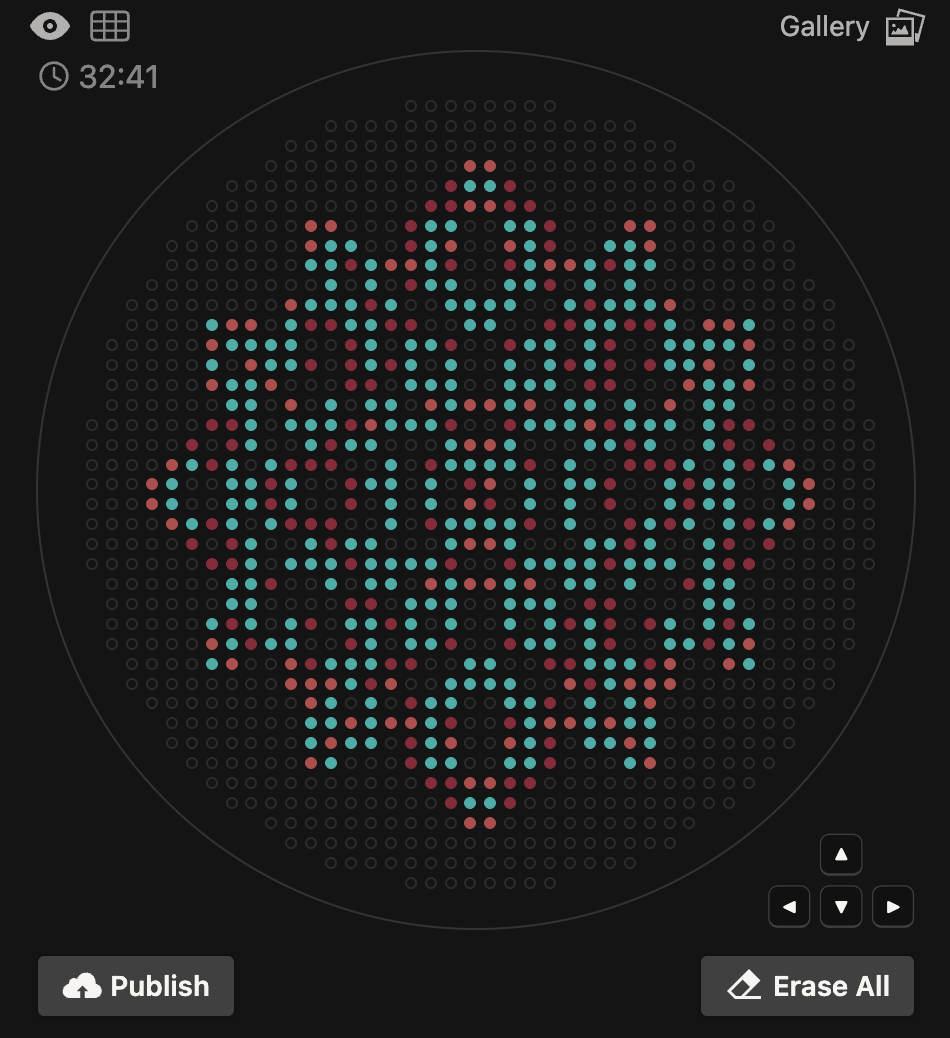

The OT-2 dispensed 0.75 µL drops of each fluorescent strain onto the charcoal agar plate.
After overnight incubation, the plate was imaged under UV light.


Using PCR-based mutagenesis, Gibson Assembly, and bacterial transformation, we intended to introduce targeted mutations into the amilCP gene to produce magenta and blue color variants in E. coli.
We ran two separate PCR reactions from the mUAV plasmid template using Phusion HF PCR Master Mix:
Backbone Fragment (Backbone Fwd + Backbone Rev primers): amplifies the origin of replication, chloramphenicol resistance gene, promoter, and RBS.
Color Insert Fragment (Color Fwd + Color Rev primers): amplifies the region from 24 bp before the chromophore to the terminator. The Forward primer contains a mutation at the CP site to introduce the desired color mutation(we chose magenta and blue variants).
Each reaction was run in a 25 µL volume with 0.8 µL template (38.5 ng/µL), 2.5 µL of each primer (5 µM stock), and 12.5 µL Phusion HF PCR Mix.



Thermocycler conditions:
Backbone PCR: 98°C 30s → 26 cycles of (98°C 10s / 57°C 25s / 72°C 1.5 min) → 72°C 5 min → 12°C hold
Color PCR: 98°C 15s → 26 cycles of (98°C 10s / 53°C 20s / 72°C 15s) → 72°C 5 min → 12°C hold


PCR products were purified using the Zymo DNA Clean & Concentrator kit. Each 20µL sample was mixed with 100 µL DNA Binding Bufferinto micro centrifuge tube and loaded onto a ZymoSpin column, and centrifuged. After two washes with 200 µL DNA Wash Buffer, DNA was eluted in 6 µL nuclease-free water.
Purified samples were run on through gel electrophoresis to confirm correct fragment sizes. 2 µL of each sample was mixed with 18 µL of water and loaded into the gel wells along with 20µL of pre-diluted ladder and original mUAV plasmid as references.




Purified backbone and color fragments were assembled using Gibson Assembly Master Mix. Reactions were set up in 10 µL total volume and incubated at 50°C for 15 minutes. After incubation, 100 µL of nuclease-free water was added to dilute the reaction before transformation.
Gibson Assembly works by: (1) exonuclease chewing back 5’ ends to expose complementary single-stranded overhangs, (2) annealing of complementary overhangs between backbone and insert, (3) polymerase filling in gaps, and (4) ligase sealing nicks to form a complete circular plasmid.
Gibson Assembly products were transformed into DH5α chemically competent E. coli cells using heat shock:
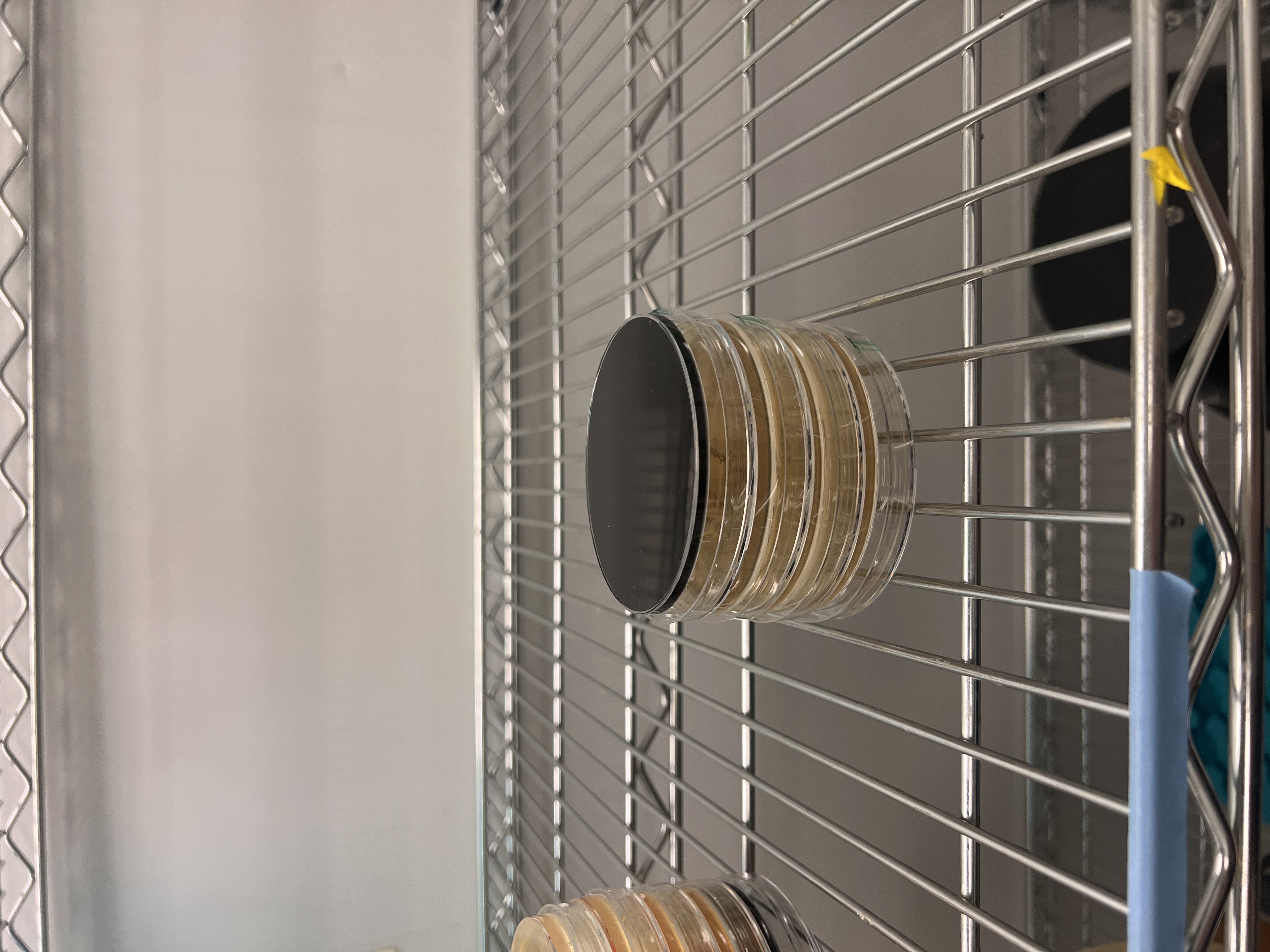
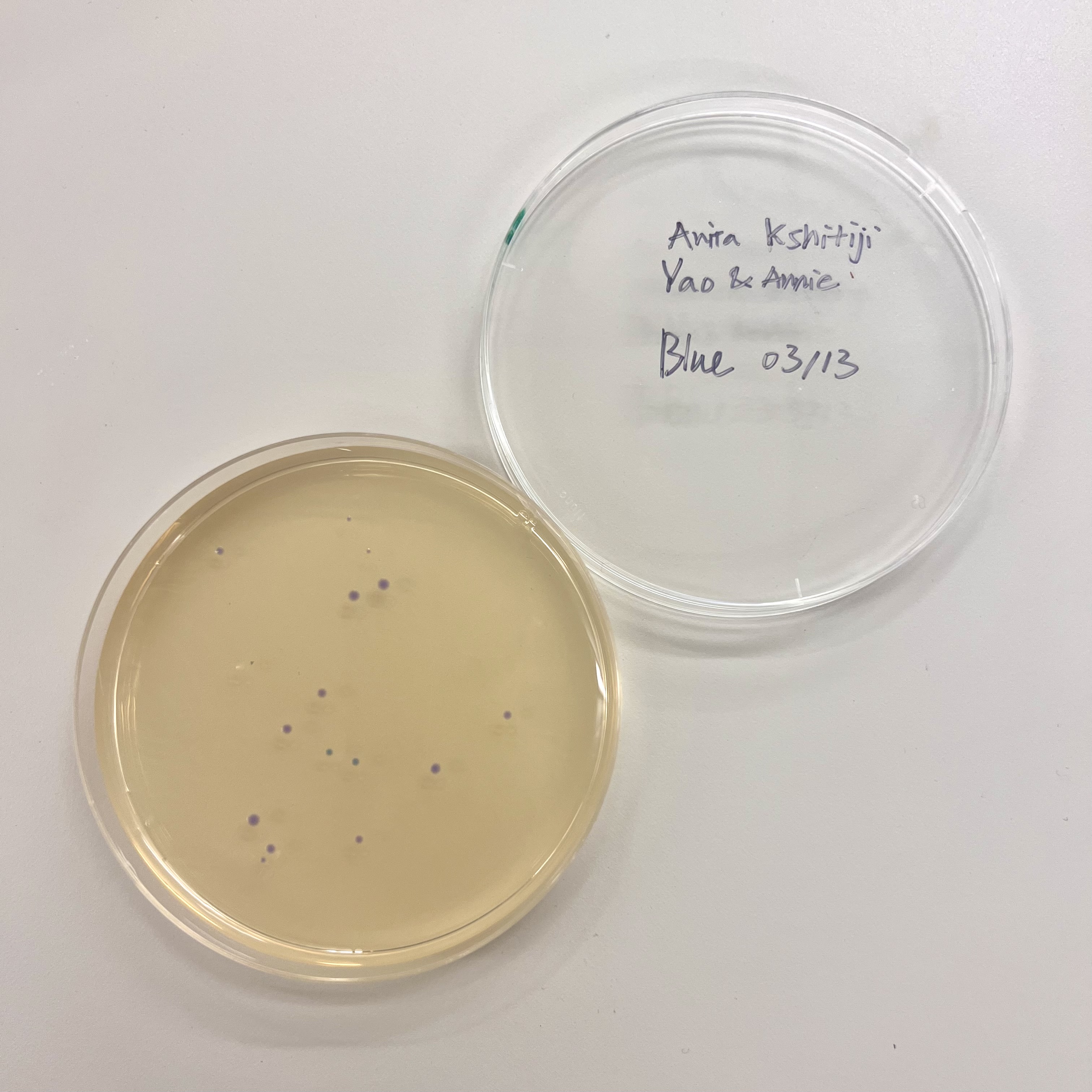
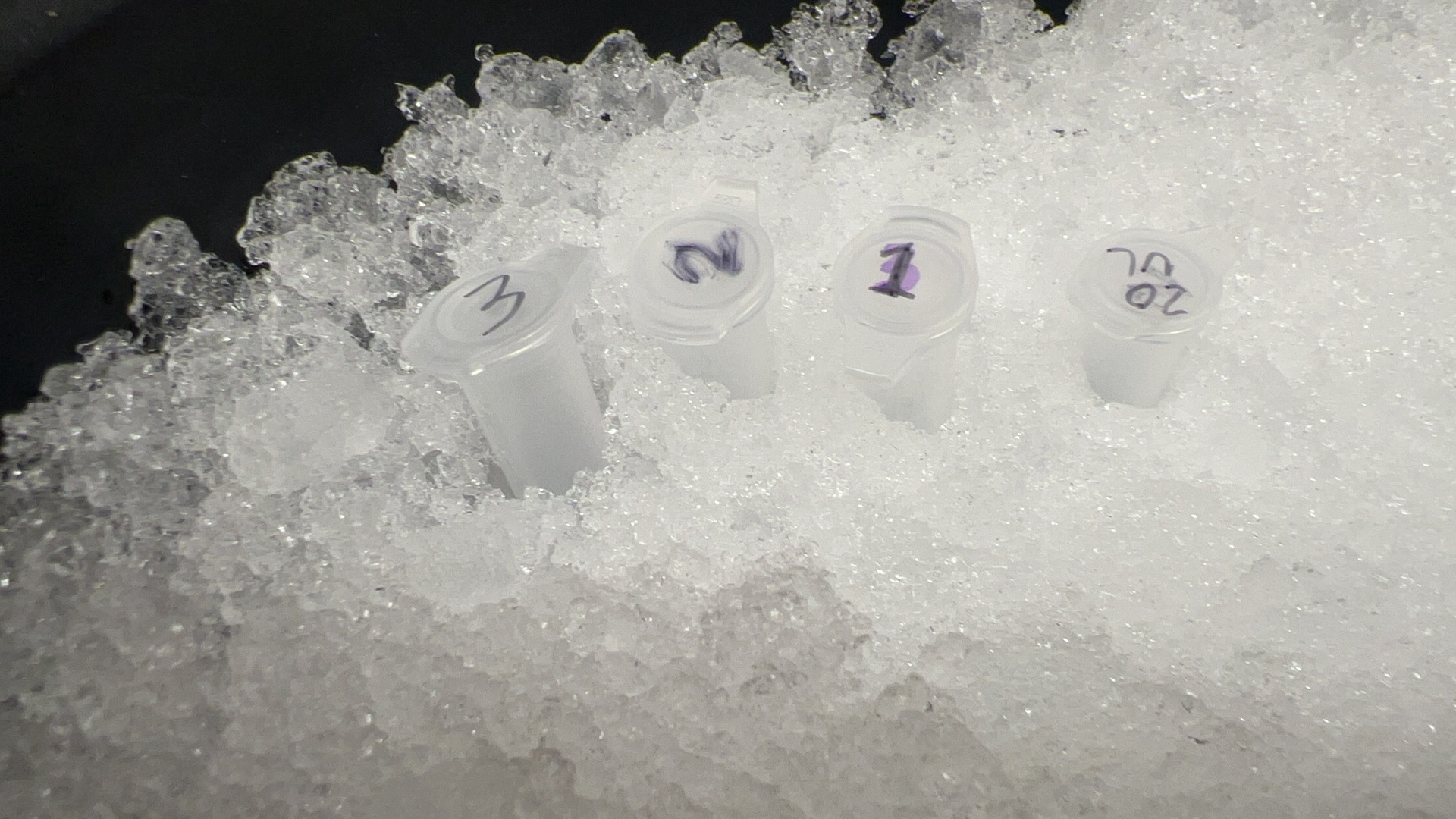
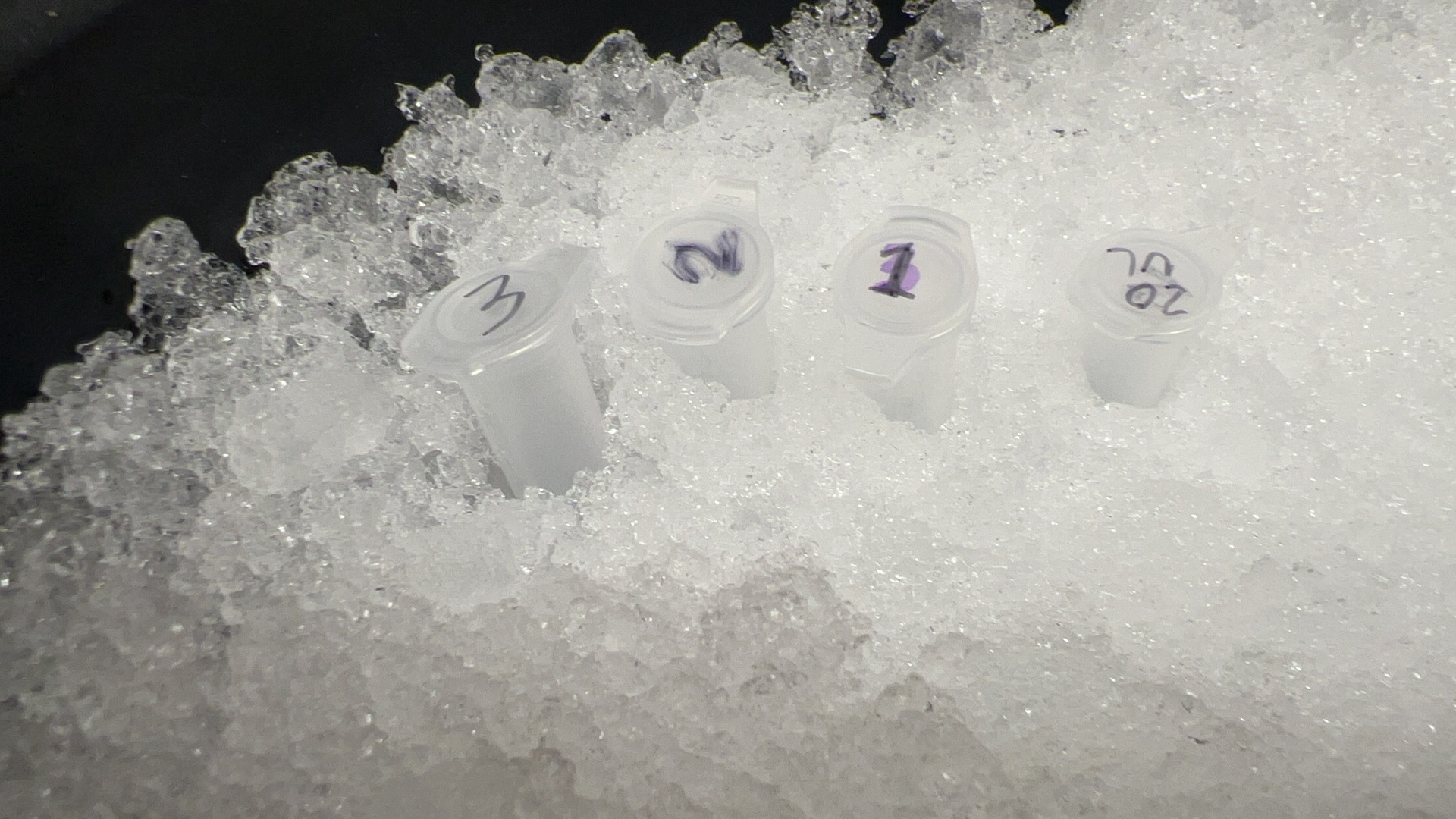






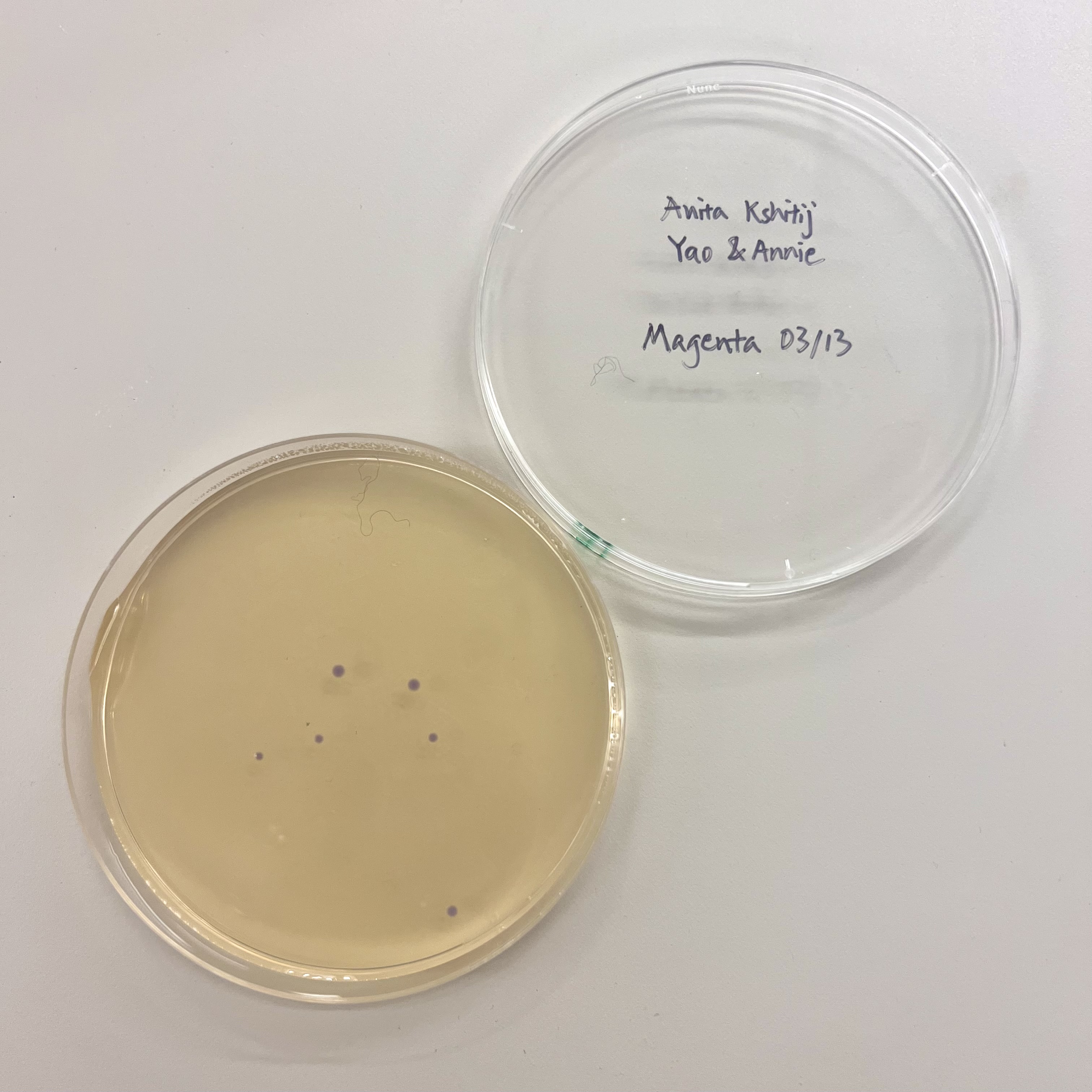

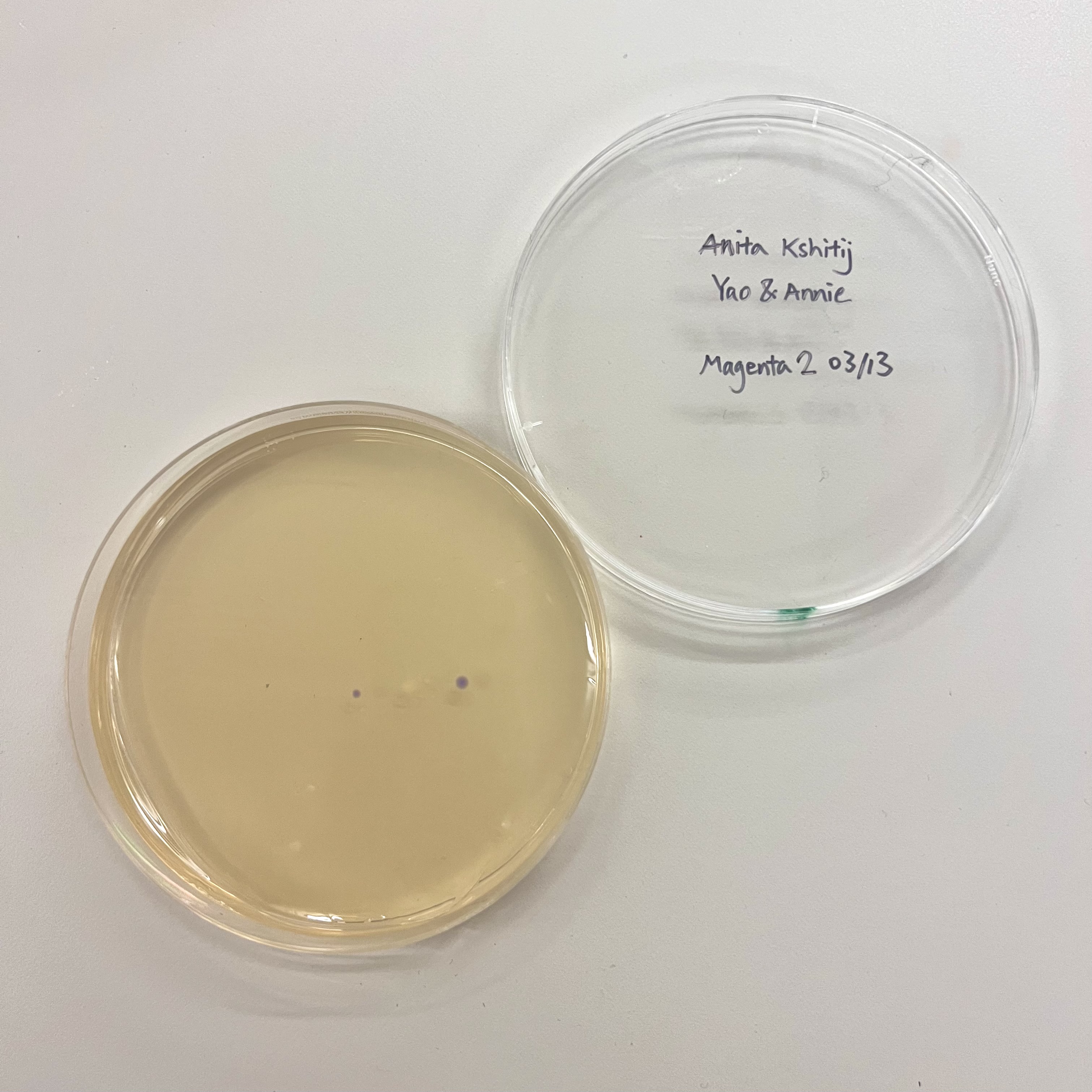


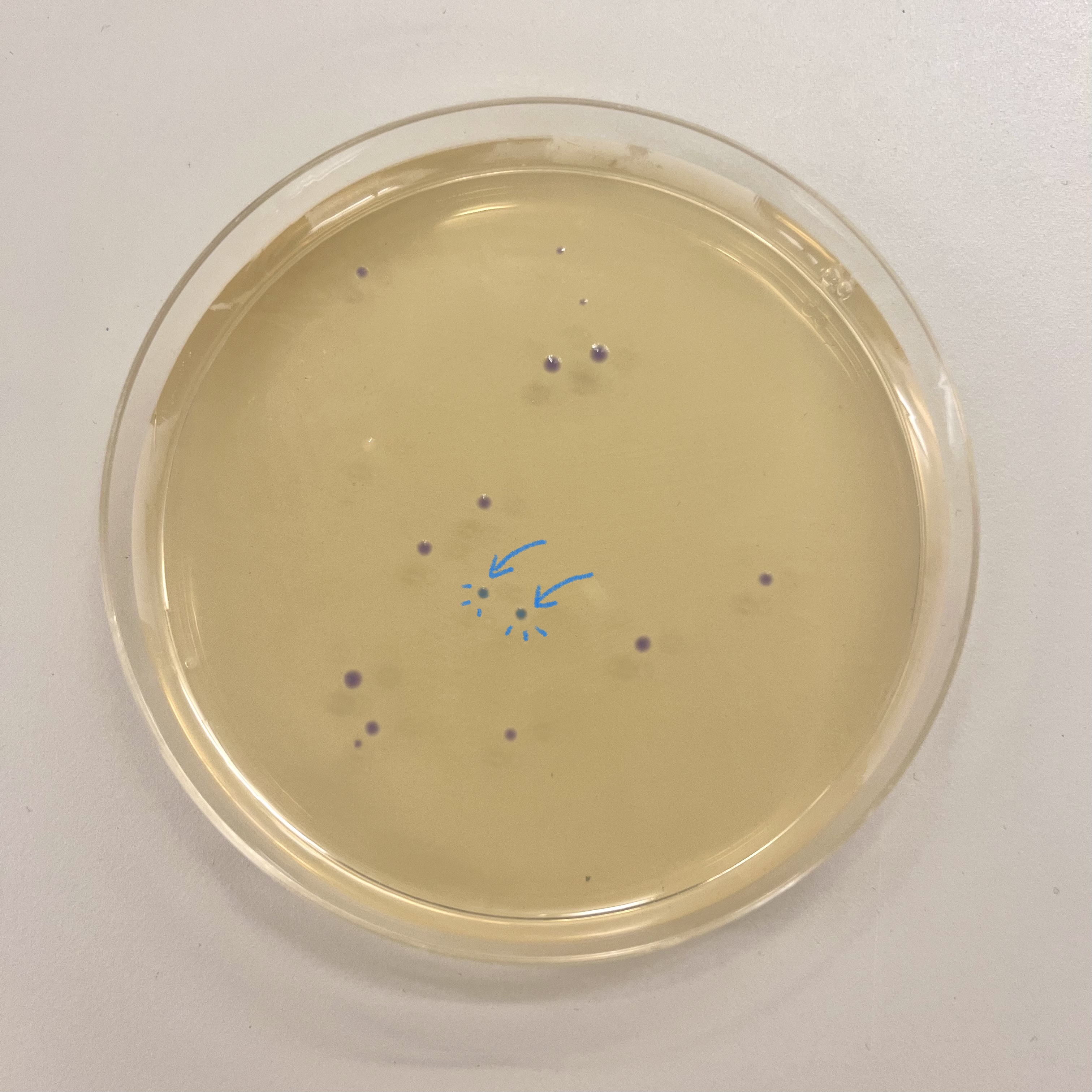
 All plates had very low colony count overall. The blue plate had 2 visibly blue-colored colonies, small partial success. Both magenta plates showed no magenta coloration, so the colonies that did grow were likely untransformed or reverted to original amilCP color of purple.
All plates had very low colony count overall. The blue plate had 2 visibly blue-colored colonies, small partial success. Both magenta plates showed no magenta coloration, so the colonies that did grow were likely untransformed or reverted to original amilCP color of purple.
Our group wanted to design a dual region neuromorphic circuit using all three ERNs. The circuit takes two inputs (X1 and X2) and produces mNeonGreen as the fluorescent output, with a bias providing a baseline level of output fluorescent mRNA.
The plasmid composition for each transfection group was as follows:
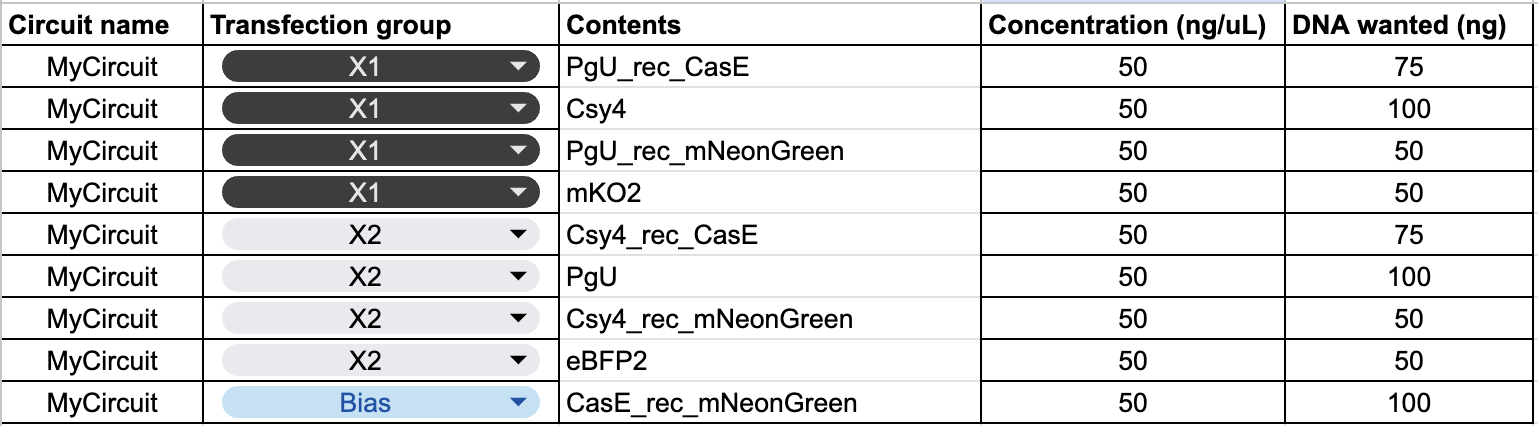

The circuit has three mNeonGreen mRNA sources with different survival conditions:
The behavior predicted under different input conditions:
X1 HIGH, X2 LOW: Csy4(from X1) destroys X2’s Csy4_rec_mNeonGreen. PgU is low, so PgU_rec_mNeonGreen from X1 survives. Result: high output.
X2 HIGH, X1 LOW: PgU(from X2) destroys X1’s PgU_rec_mNeonGreen. Csy4 is low, so Csy4_rec_mNeonGreen from X2 survives. Result: high output.
BOTH HIGH: Both direct mNeonGreen sources are destroyed. However, both ERNs also suppress each other’s CasE mRNAs, so CasE levels drop and the bias mRNA survives. Result: moderate output from bias only(will depend on bias level).
BOTH LOW: Little DNA overall, low expression across all components, and no ERN suppression means some CasE still gets made and cleaves the bias. Result: lowest output.
The circuit is therefore behaving as a Dual Region (Top-Left, Bottom-Right) circuit. Output is high when the two inputs are mismatched (one high, one low) and low when they are matched in either direction.
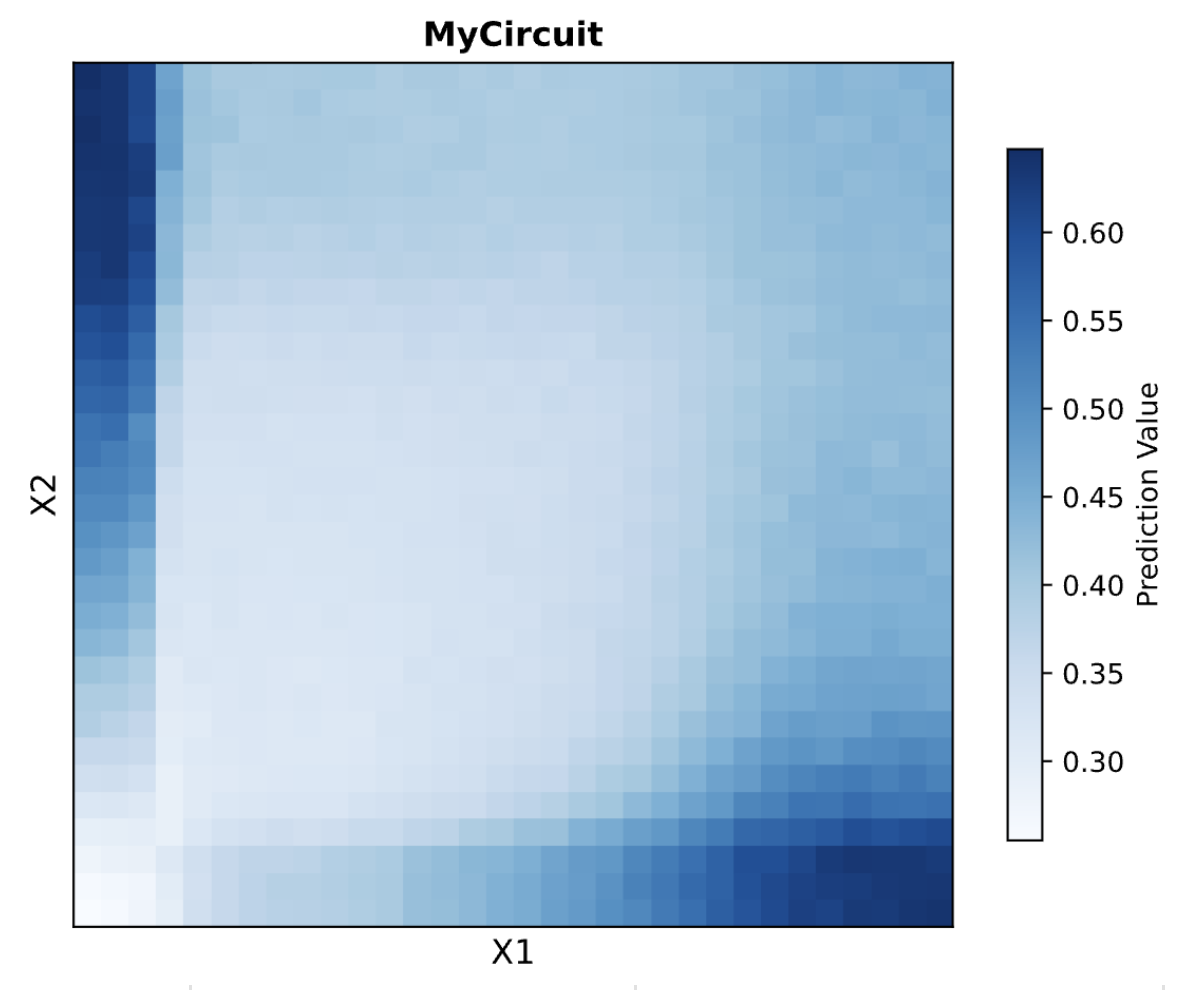

The prediction heatmap shows Dual Region behavior. High output appears in the top-left (X1 low, X2 high) and bottom-right (X1 high, X2 low) corners. The bottom-left (both low) is the minimum, and the top-right (both high) is moderate rather than high, reflecting the partial bias contribution when both ERNs cancel each other.


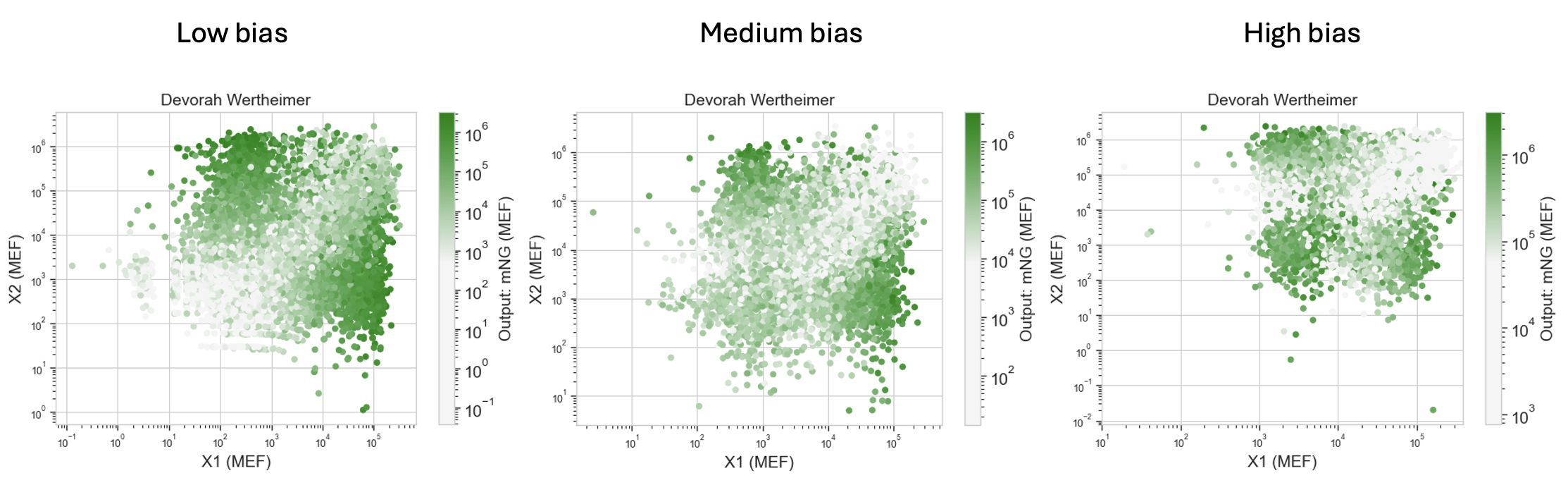
Each dot in the class scatterplots represents a single HEK293 cell. X-axis is X1 input (via mKO2), Y-axis is X2 input (via eBFP2), and dot color is mNeonGreen output intensity. The three panels show results at low, medium, and high bias DNA concentrations.
Low bias: Bright cells concentrate in the mismatched input regions (high X2 + low X1, or high X1 + low X2), consistent with the direct mNeonGreen mRNAs dominating. The pattern is clearly dual-region.
Medium bias: Output spreads more uniformly across the input space and dims overall. The bias is now competing with the direct mNeonGreen sources but not yet dominating, producing a flatter, less structured response.
High bias: The brightest cells shift toward the both-high region (top-right). With so much CasE_rec_mNeonGreen available, the dominant output source is now the bias when both ERNs are high and mutually suppress CasE production.