Week 6: Gibson Assembly

Using PCR-based mutagenesis, Gibson Assembly, and bacterial transformation, we intended to introduce targeted mutations into the amilCP gene to produce magenta and blue color variants in E. coli.
Part 1: PCR
We ran two separate PCR reactions from the mUAV plasmid template using Phusion HF PCR Master Mix:
Backbone Fragment (Backbone Fwd + Backbone Rev primers): amplifies the origin of replication, chloramphenicol resistance gene, promoter, and RBS.
Color Insert Fragment (Color Fwd + Color Rev primers): amplifies the region from 24 bp before the chromophore to the terminator. The Forward primer contains a mutation at the CP site to introduce the desired color mutation(we chose magenta and blue variants).
Each reaction was run in a 25 µL volume with 0.8 µL template (38.5 ng/µL), 2.5 µL of each primer (5 µM stock), and 12.5 µL Phusion HF PCR Mix.



Thermocycler conditions:
Backbone PCR: 98°C 30s → 26 cycles of (98°C 10s / 57°C 25s / 72°C 1.5 min) → 72°C 5 min → 12°C hold
Color PCR: 98°C 15s → 26 cycles of (98°C 10s / 53°C 20s / 72°C 15s) → 72°C 5 min → 12°C hold


Part 1a: DNA Purification
PCR products were purified using the Zymo DNA Clean & Concentrator kit. Each 20µL sample was mixed with 100 µL DNA Binding Bufferinto micro centrifuge tube and loaded onto a ZymoSpin column, and centrifuged. After two washes with 200 µL DNA Wash Buffer, DNA was eluted in 6 µL nuclease-free water.
Part 1b: Diagnostic Gel Electrophoresis
Purified samples were run on through gel electrophoresis to confirm correct fragment sizes. 2 µL of each sample was mixed with 18 µL of water and loaded into the gel wells along with 20µL of pre-diluted ladder and original mUAV plasmid as references.




Part 2a: Gibson Assembly
Purified backbone and color fragments were assembled using Gibson Assembly Master Mix. Reactions were set up in 10 µL total volume and incubated at 50°C for 15 minutes. After incubation, 100 µL of nuclease-free water was added to dilute the reaction before transformation.
Gibson Assembly works by: (1) exonuclease chewing back 5’ ends to expose complementary single-stranded overhangs, (2) annealing of complementary overhangs between backbone and insert, (3) polymerase filling in gaps, and (4) ligase sealing nicks to form a complete circular plasmid.
Part 2b: Transformation
Gibson Assembly products were transformed into DH5α chemically competent E. coli cells using heat shock:
- Thawed competent cells on ice for 10 minutes
- Added 4 µL of diluted Gibson Assembly product to 20 µL of competent cells
- Incubated on ice for 30 minutes
- Heat shocked at 42°C for exactly 45 seconds, then returned to ice for 5 minutes
- Added 100 µL SOC media and incubated with shaking for 60 minutes


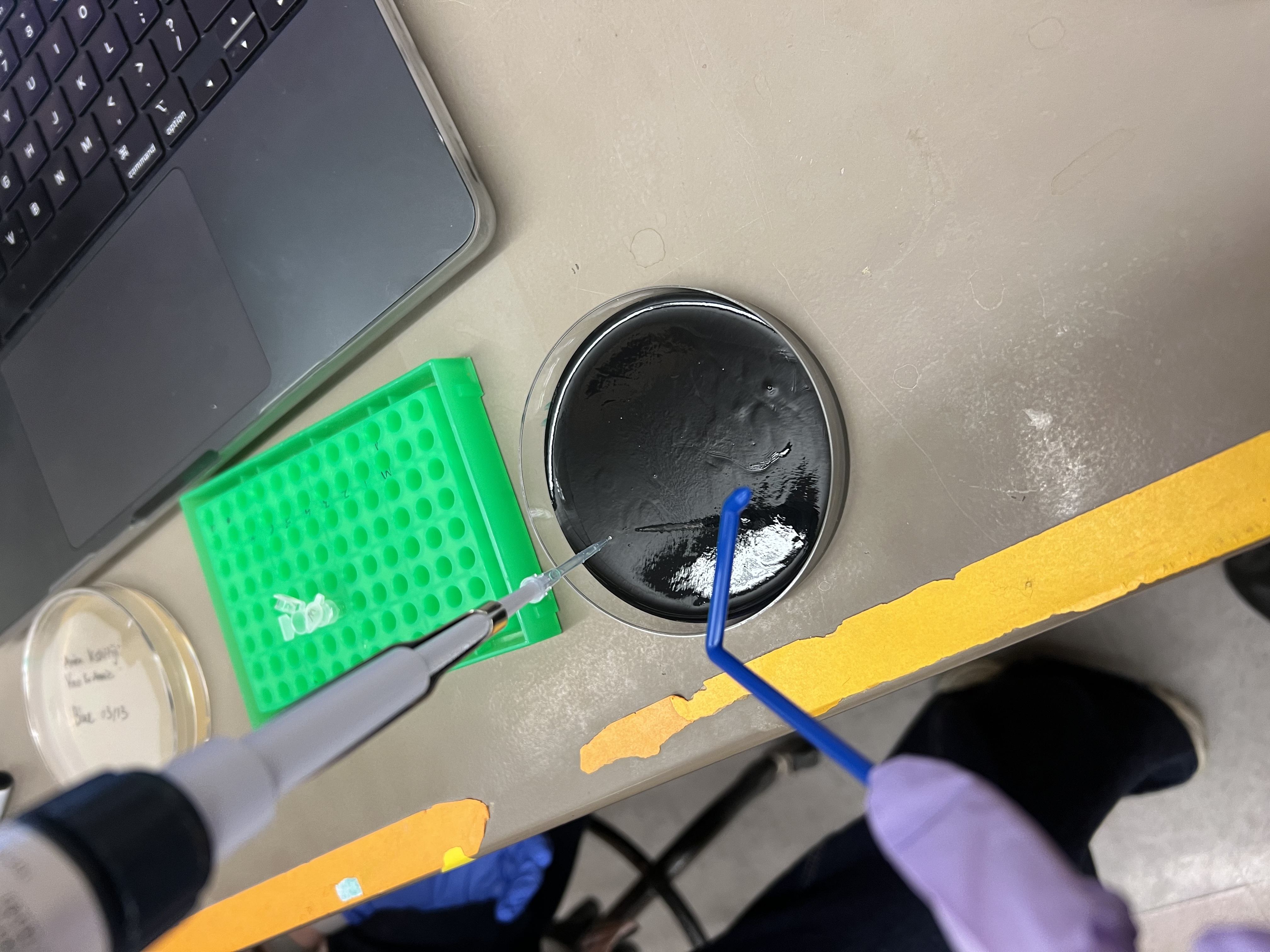
- Plated 100 µL onto agar plates


- Incubated plates at 37°C for 72 hours


Results
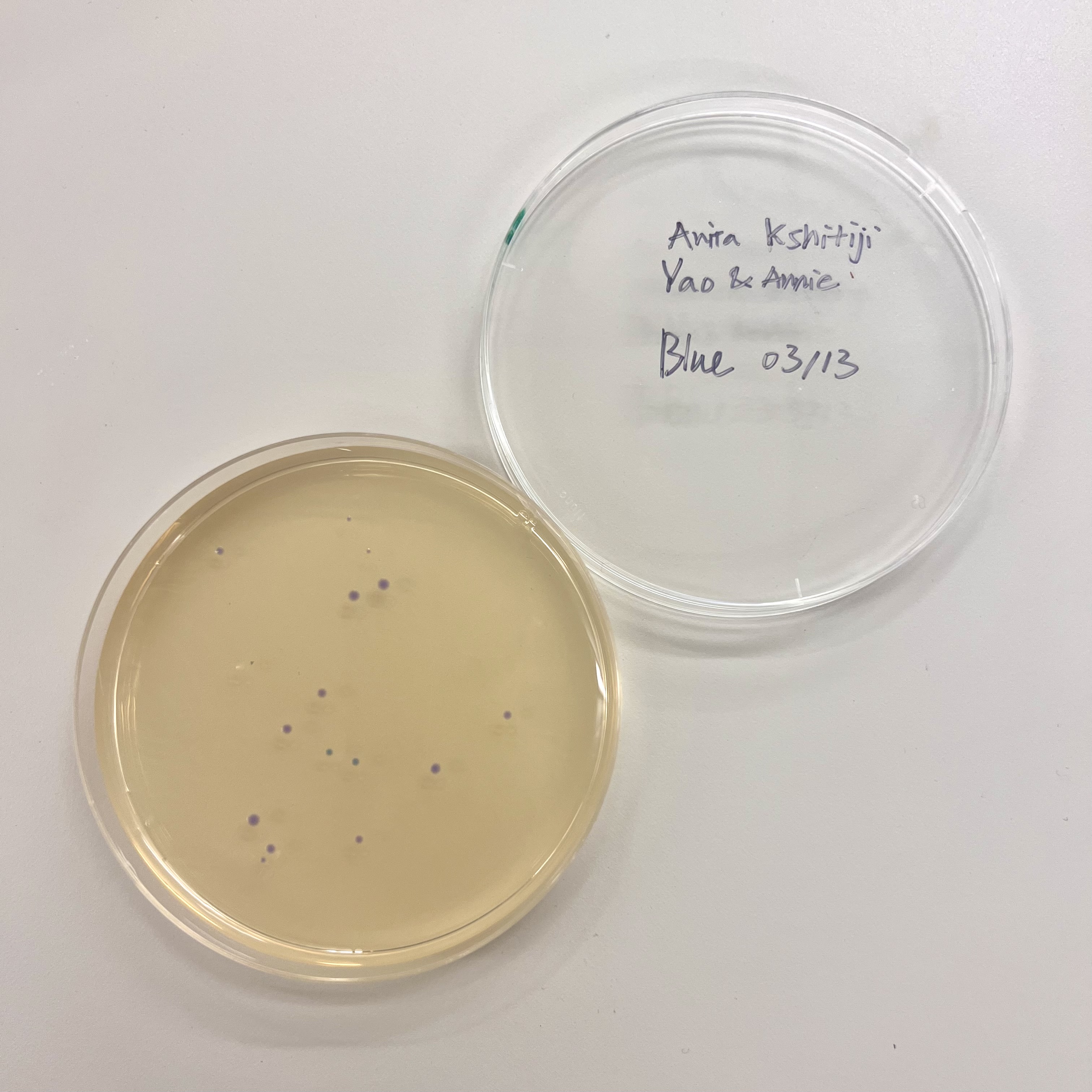
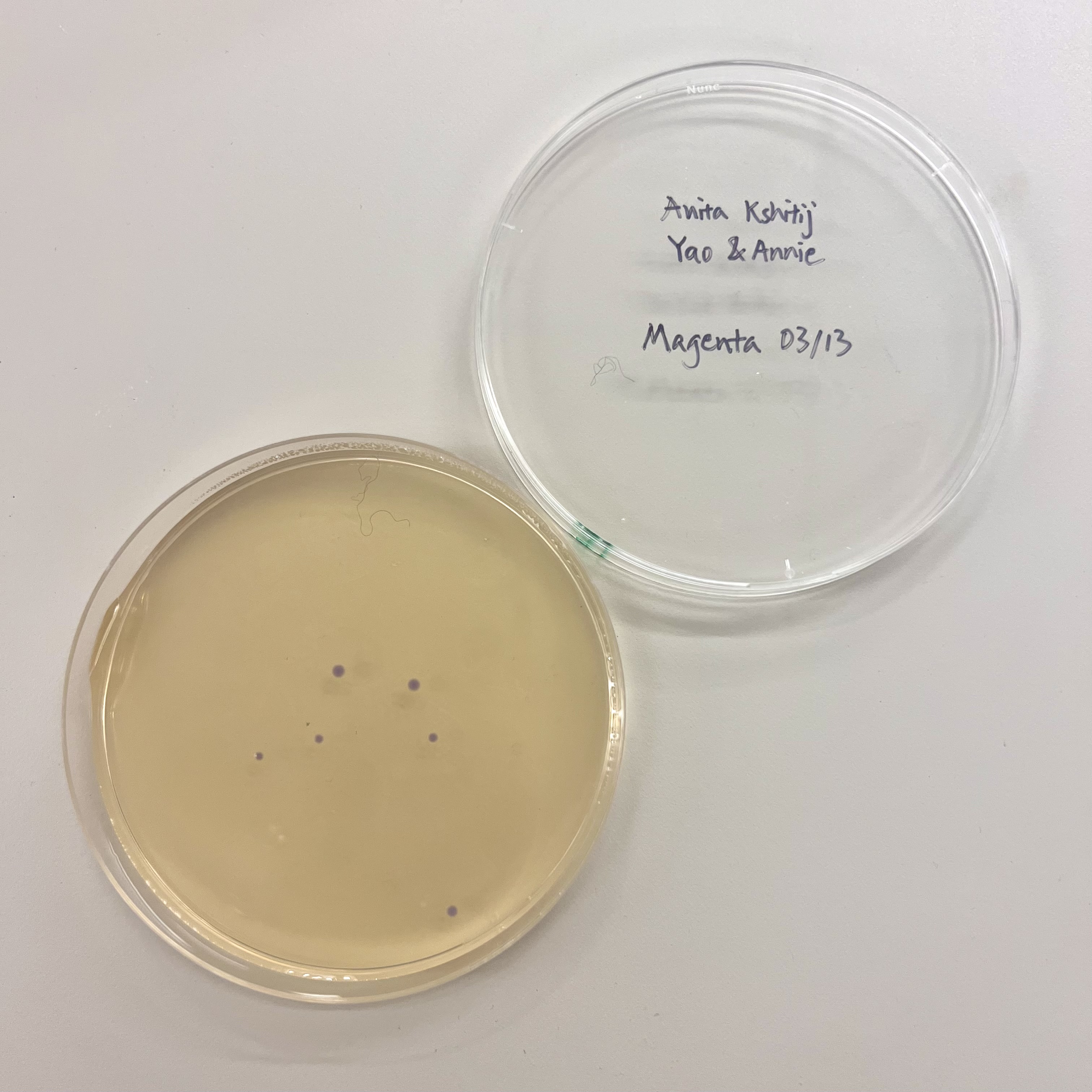

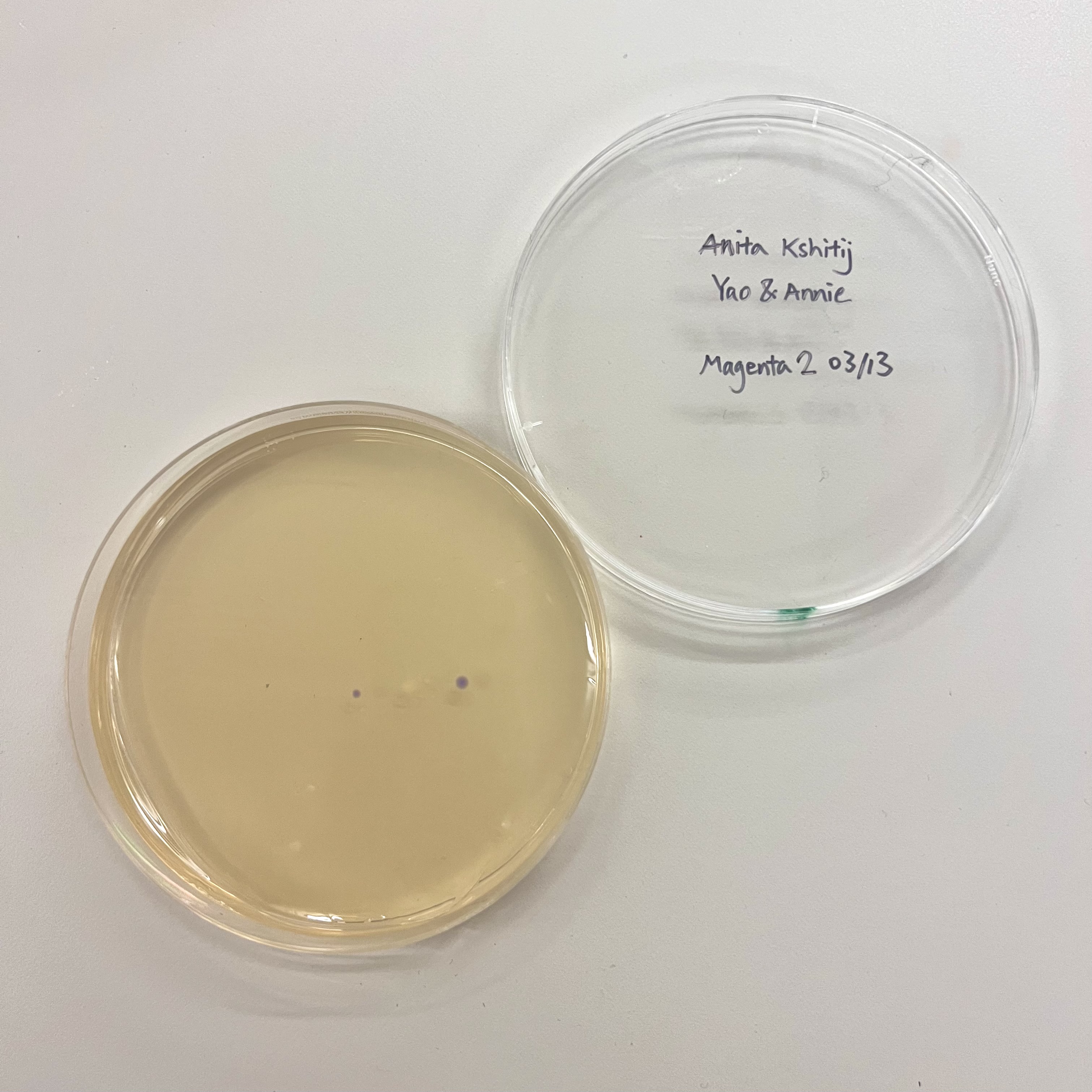


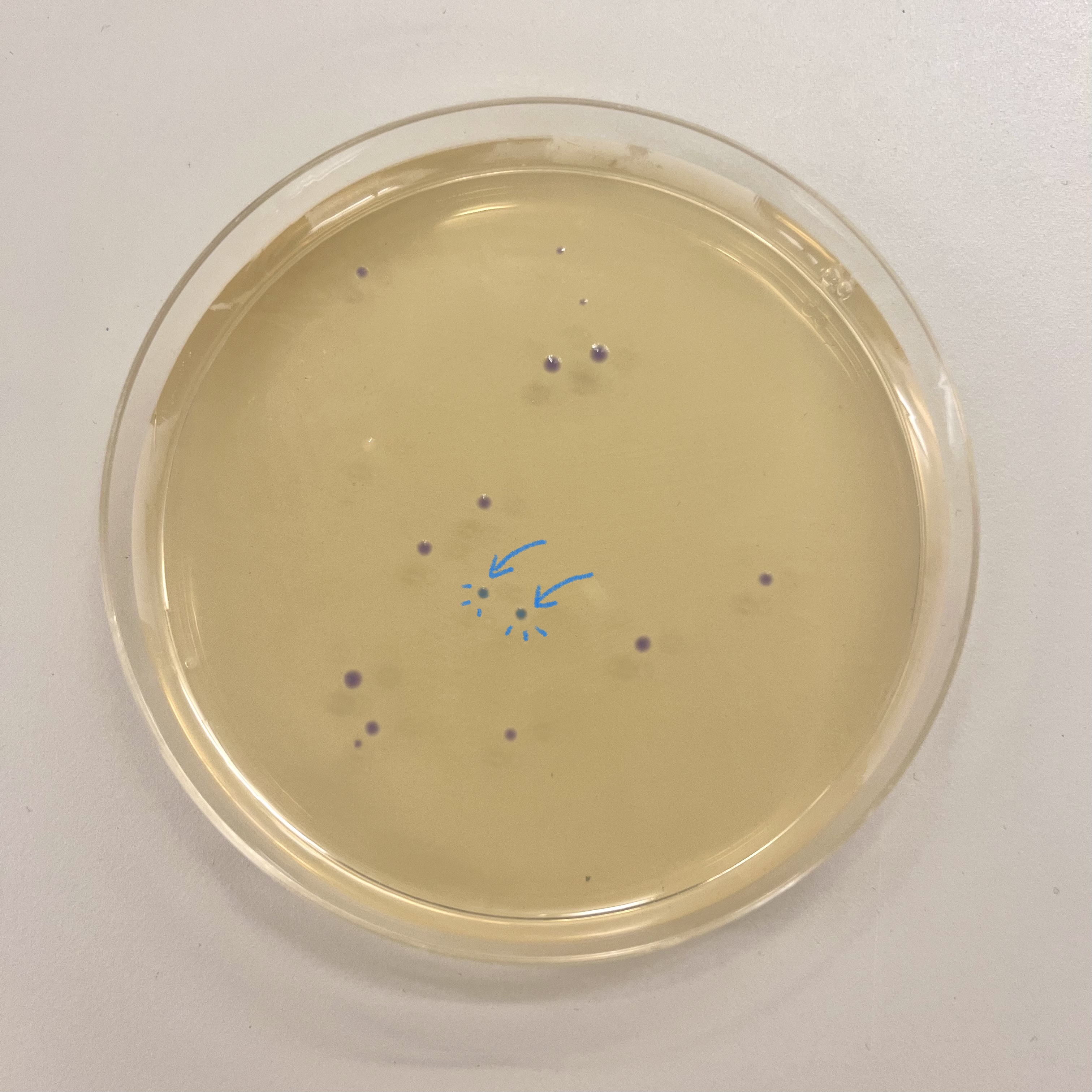
 All plates had very low colony count overall. The blue plate had 2 visibly blue-colored colonies, small partial success. Both magenta plates showed no magenta coloration, so the colonies that did grow were likely untransformed or reverted to original amilCP color of purple.
All plates had very low colony count overall. The blue plate had 2 visibly blue-colored colonies, small partial success. Both magenta plates showed no magenta coloration, so the colonies that did grow were likely untransformed or reverted to original amilCP color of purple.

