Week 2 HW: DNA Sequencing and Synthesis
PART 2
 path to image: 2026a-zoe-isabel-senon/webpages/src/branch/main/assets/images/heartsim.jpg
path to image: 2026a-zoe-isabel-senon/webpages/src/branch/main/assets/images/heartsim.jpg
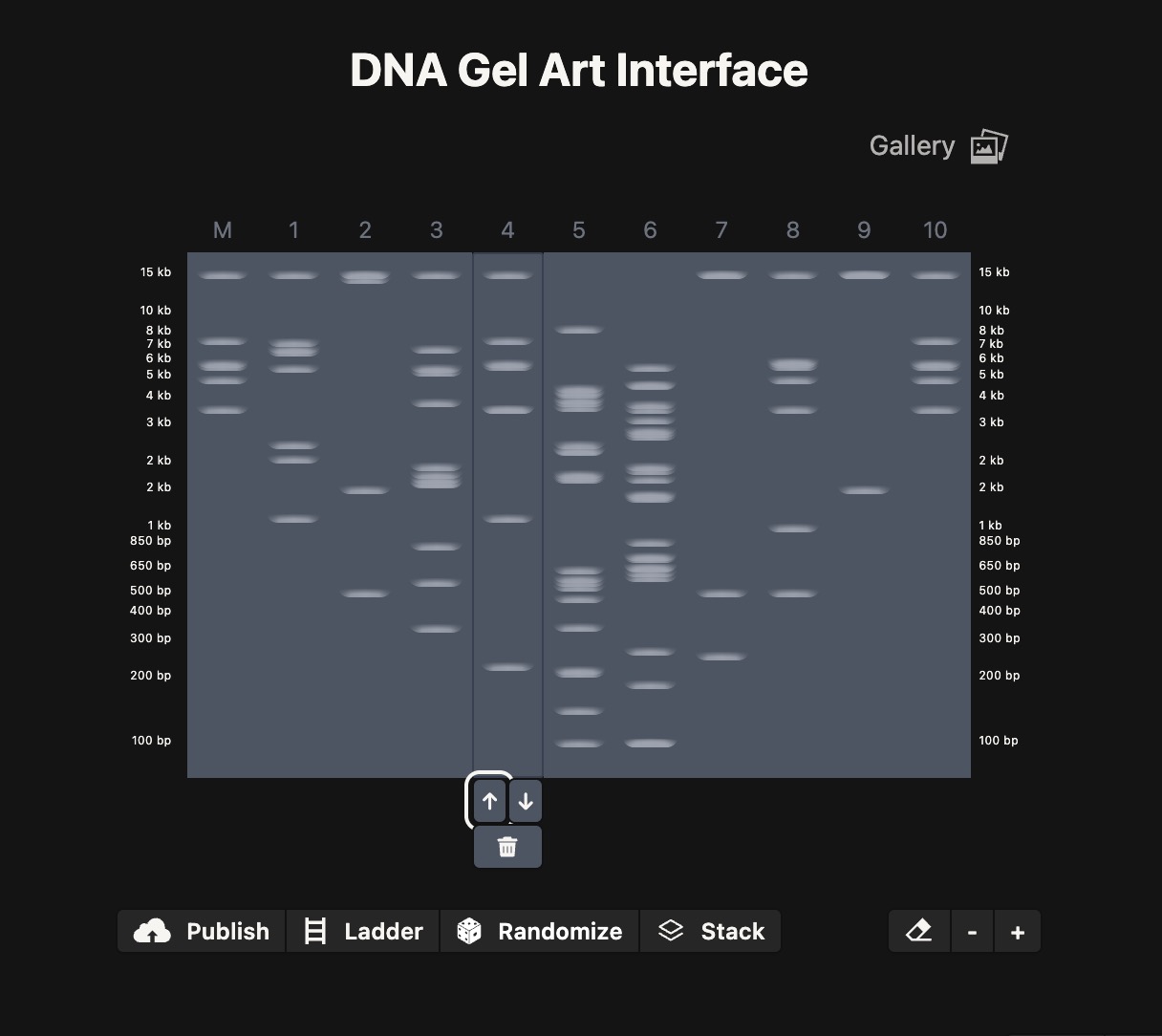
(I tried to recreate a heart shape…)
I did the digest simulation in Benchling and checked the virtual gel simulation result:
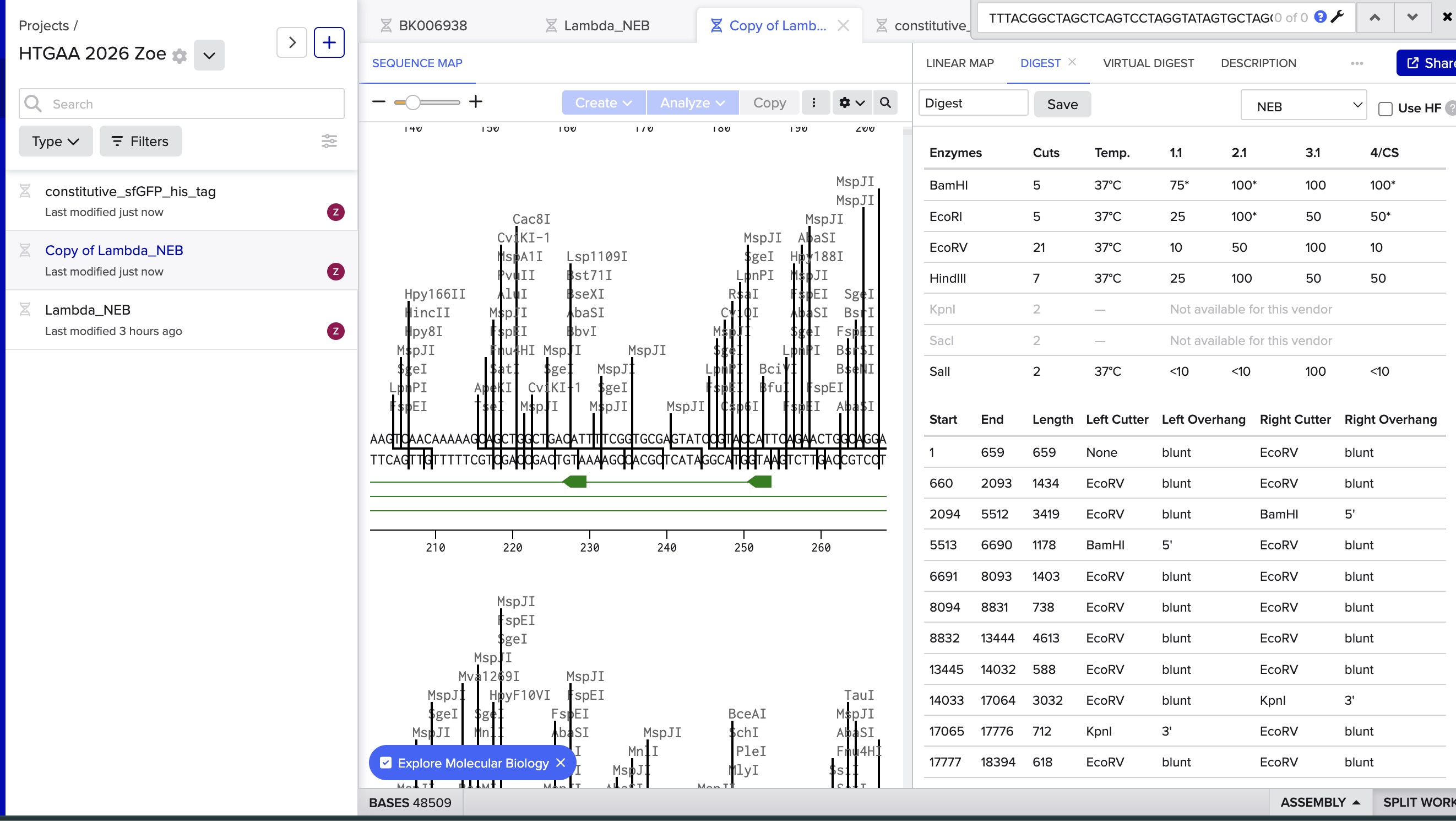
 image in: 2026a-zoe-isabel-senon/webpages/src/branch/main/assets/images/benchlingdigest1.jpg
image in: 2026a-zoe-isabel-senon/webpages/src/branch/main/assets/images/benchlingdigest1.jpg
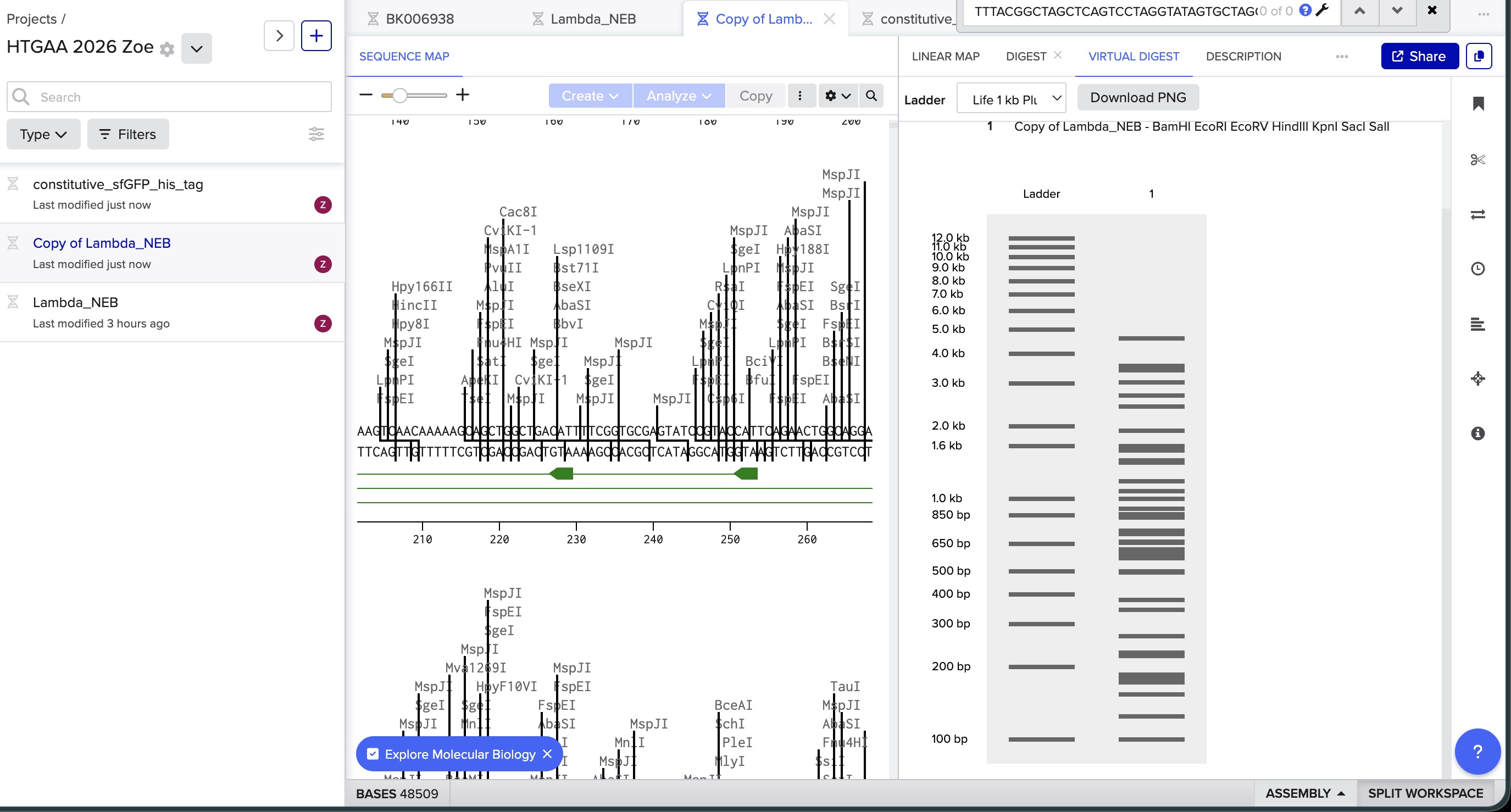
 image in: 2026a-zoe-isabel-senon/webpages/src/branch/main/assets/images/benchlingdigest2.jpg
image in: 2026a-zoe-isabel-senon/webpages/src/branch/main/assets/images/benchlingdigest2.jpg
3.1 PICK A PROTEIN & Find the protein sequence:
I picked SIRT1
And downloaded the sequence from Uniprot:
3.2 - Protein → DNA sequence
Obtained through the reverse translation tool after attaching the human codon table:
```jsx
Reverse Translate
Results for 747 residue sequence "sp|Q96EB6|SIR1_HUMAN NAD-dependent protein deacetylase sirtuin-1 OS=Homo sapiens OX=9606 GN=SIRT1 PE=1 SV=2" starting "MADEAALALQ"
>reverse translation of sp|Q96EB6|SIR1_HUMAN NAD-dependent protein deacetylase sirtuin-1 OS=Homo sapiens OX=9606 GN=SIRT1 PE=1 SV=2 to a 2241 base sequence of most likely codons.
atggccgacgaggccgccctggccctgcagcccggcggcagccccagcgccgccggcgcc
gacagagaggccgccagcagccccgccggcgagcccctgagaaagagacccagaagagac
ggccccggcctggagagaagccccggcgagcccggcggcgccgcccccgagagagaggtg
cccgccgccgccagaggctgccccggcgccgccgccgccgccctgtggagagaggccgag
gccgaggccgccgccgccggcggcgagcaggaggcccaggccaccgccgccgccggcgag
ggcgacaacggccccggcctgcagggccccagcagagagccccccctggccgacaacctg
tacgacgaggacgacgacgacgagggcgaggaggaggaggaggccgccgccgccgccatc
ggctacagagacaacctgctgttcggcgacgagatcatcaccaacggcttccacagctgc
gagagcgacgaggaggacagagccagccacgccagcagcagcgactggacccccagaccc
agaatcggcccctacaccttcgtgcagcagcacctgatgatcggcaccgaccccagaacc
atcctgaaggacctgctgcccgagaccatccccccccccgagctggacgacatgaccctg
tggcagatcgtgatcaacatcctgagcgagccccccaagagaaagaagagaaaggacatc
aacaccatcgaggacgccgtgaagctgctgcaggagtgcaagaagatcatcgtgctgacc
ggcgccggcgtgagcgtgagctgcggcatccccgacttcagaagcagagacggcatctac
gccagactggccgtggacttccccgacctgcccgacccccaggccatgttcgacatcgag
tacttcagaaaggaccccagacccttcttcaagttcgccaaggagatctaccccggccag
ttccagcccagcctgtgccacaagttcatcgccctgagcgacaaggagggcaagctgctg
agaaactacacccagaacatcgacaccctggagcaggtggccggcatccagagaatcatc
cagtgccacggcagcttcgccaccgccagctgcctgatctgcaagtacaaggtggactgc
gaggccgtgagaggcgacatcttcaaccaggtggtgcccagatgccccagatgccccgcc
gacgagcccctggccatcatgaagcccgagatcgtgttcttcggcgagaacctgcccgag
cagttccacagagccatgaagtacgacaaggacgaggtggacctgctgatcgtgatcggc
agcagcctgaaggtgagacccgtggccctgatccccagcagcatcccccacgaggtgccc
cagatcctgatcaacagagagcccctgccccacctgcacttcgacgtggagctgctgggc
gactgcgacgtgatcatcaacgagctgtgccacagactgggcggcgagtacgccaagctg
tgctgcaaccccgtgaagctgagcgagatcaccgagaagccccccagaacccagaaggag
ctggcctacctgagcgagctgccccccacccccctgcacgtgagcgaggacagcagcagc
cccgagagaaccagcccccccgacagcagcgtgatcgtgaccctgctggaccaggccgcc
aagagcaacgacgacctggacgtgagcgagagcaagggctgcatggaggagaagccccag
gaggtgcagaccagcagaaacgtggagagcatcgccgagcagatggagaaccccgacctg
aagaacgtgggcagcagcaccggcgagaagaacgagagaaccagcgtggccggcaccgtg
agaaagtgctggcccaacagagtggccaaggagcagatcagcagaagactggacggcaac
cagtacctgttcctgccccccaacagatacatcttccacggcgccgaggtgtacagcgac
agcgaggacgacgtgctgagcagcagcagctgcggcagcaacagcgacagcggcacctgc
cagagccccagcctggaggagcccatggaggacgagagcgagatcgaggagttctacaac
ggcctggaggacgagcccgacgtgcccgagagagccggcggcgccggcttcggcaccgac
ggcgacgaccaggaggccatcaacgaggccatcagcgtgaagcaggaggtgaccgacatg
aactaccccagcaacaagagc
>reverse translation of sp|Q96EB6|SIR1_HUMAN NAD-dependent protein deacetylase sirtuin-1 OS=Homo sapiens OX=9606 GN=SIRT1 PE=1 SV=2 to a 2241 base sequence of consensus codons.
atggcngaygargcngcnytngcnytncarccnggnggnwsnccnwsngcngcnggngcn
gaymgngargcngcnwsnwsnccngcnggngarccnytnmgnaarmgnccnmgnmgngay
ggnccnggnytngarmgnwsnccnggngarccnggnggngcngcnccngarmgngargtn
ccngcngcngcnmgnggntgyccnggngcngcngcngcngcnytntggmgngargcngar
gcngargcngcngcngcnggnggngarcargargcncargcnacngcngcngcnggngar
ggngayaayggnccnggnytncarggnccnwsnmgngarccnccnytngcngayaayytn
taygaygargaygaygaygaygarggngargargargargargcngcngcngcngcnath
ggntaymgngayaayytnytnttyggngaygarathathacnaayggnttycaywsntgy
garwsngaygargargaymgngcnwsncaygcnwsnwsnwsngaytggacnccnmgnccn
mgnathggnccntayacnttygtncarcarcayytnatgathggnacngayccnmgnacn
athytnaargayytnytnccngaracnathccnccnccngarytngaygayatgacnytn
tggcarathgtnathaayathytnwsngarccnccnaarmgnaaraarmgnaargayath
aayacnathgargaygcngtnaarytnytncargartgyaaraarathathgtnytnacn
ggngcnggngtnwsngtnwsntgyggnathccngayttymgnwsnmgngayggnathtay
gcnmgnytngcngtngayttyccngayytnccngayccncargcnatgttygayathgar
tayttymgnaargayccnmgnccnttyttyaarttygcnaargarathtayccnggncar
ttycarccnwsnytntgycayaarttyathgcnytnwsngayaargarggnaarytnytn
mgnaaytayacncaraayathgayacnytngarcargtngcnggnathcarmgnathath
cartgycayggnwsnttygcnacngcnwsntgyytnathtgyaartayaargtngaytgy
gargcngtnmgnggngayathttyaaycargtngtnccnmgntgyccnmgntgyccngcn
gaygarccnytngcnathatgaarccngarathgtnttyttyggngaraayytnccngar
carttycaymgngcnatgaartaygayaargaygargtngayytnytnathgtnathggn
wsnwsnytnaargtnmgnccngtngcnytnathccnwsnwsnathccncaygargtnccn
carathytnathaaymgngarccnytnccncayytncayttygaygtngarytnytnggn
gaytgygaygtnathathaaygarytntgycaymgnytnggnggngartaygcnaarytn
tgytgyaayccngtnaarytnwsngarathacngaraarccnccnmgnacncaraargar
ytngcntayytnwsngarytnccnccnacnccnytncaygtnwsngargaywsnwsnwsn
ccngarmgnacnwsnccnccngaywsnwsngtnathgtnacnytnytngaycargcngcn
aarwsnaaygaygayytngaygtnwsngarwsnaarggntgyatggargaraarccncar
gargtncaracnwsnmgnaaygtngarwsnathgcngarcaratggaraayccngayytn
aaraaygtnggnwsnwsnacnggngaraaraaygarmgnacnwsngtngcnggnacngtn
mgnaartgytggccnaaymgngtngcnaargarcarathwsnmgnmgnytngayggnaay
cartayytnttyytnccnccnaaymgntayathttycayggngcngargtntaywsngay
wsngargaygaygtnytnwsnwsnwsnwsntgyggnwsnaaywsngaywsnggnacntgy
carwsnccnwsnytngargarccnatggargaygarwsngarathgargarttytayaay
ggnytngargaygarccngaygtnccngarmgngcnggnggngcnggnttyggnacngay
ggngaygaycargargcnathaaygargcnathwsngtnaarcargargtnacngayatg
aaytayccnwsnaayaarwsn
```
3.3 - In your own words, describe why you need to optimize codon usage. Which organism have you chosen to optimize the codon sequence for and why?
I have chosen to optimize it for homo sapiens, because I am interested in human medicine and I want to learn more about how certain proteins might affect human lifespans.
We need to optimize codon usage because different organisms have different availability of tRNA to match certain codons, and we want to optimize for tRNA abundance to maximize translation speed, and mRNA stability and structure. These optimizations can be made because the same amino acid can be produced with different codons.
3.4 - What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
I could transfect the necessary genes into mammalian cells (like neurons, or HEK cells, which divide rapidly!) to make them express this protein in large quantities. We do this by basically integrating these “instructions” (as, for example, plasmids) for producing different specific proteins into the cell, and letting their machinery decode them in the same way that it does with their own genome.
(I have done this in the past with HEK cells to make them express RFP, using lipofectamine as the transfection method/reagent.)
We optimize the plasmid content for the target organism by choosing a certain promoter, codon-optimized CDS, and regulatory elements.
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would want to sequence the genomes of healthy centenarians or supercentenarians (people who reach 110 years of age), so that we can learn if there is a genetic basis for longevity and find enhancing variants. I know that certain variants of the FOXO3 gene have been found to be more prevalently enriched among this population (paper), and I would like to learn more.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Also answer the following questions:
- Is your method first-, second- or third-generation or other? How so?**
- What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.**
- What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?**
- What is the output of your chosen sequencing technology?**
I would choose Illumina short-read WGS, which is a relatively quick, affordable and accurate current industry standard for consumer sequencing.
It is a second-generation method that allows to do massively-parallelized short-reads.
To do this, we would take a sample from the supercentenarians (saliva or blood), perform end-repair and A-tailing, ligate Illumina adapters, and PCR-amplify to make libraries.
The output is a FASTQ file which we align to the human reference genome to find (call) SNPs, indels, etc. Basically to identify how and where this given genome differs from the “reference” one (assembled as an average of a group of +100 individuals).
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond.
Klotho gene therapy for cognitive enhancement and longevity! I has been linked to better cognition in mice and humans (paper: https://pmc.ncbi.nlm.nih.gov/articles/PMC4176932/ )
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
Also answer the following questions:
- What are the essential steps of your chosen sequencing methods?**
- What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?**
I would use the industry standard (phosphoramidite chemical oligo synthesis + Gibson assembly) as it is currently the most battle-tested one.
The limitation is that it only allows us to assemble up to around 200 nucleotides at a time before the error rate becomes so high as to make the output unusable (too different from the referent we want to “print”). To print out longer sequences, we must separately assemble them using methods like Gibson assembly.
5.3 DNA Edit
(i) What DNA would you want to edit and why? What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
Genes associated with longevity or cognitive enhancement, like FOXO3 or Klotho. I would like to test these in both humans and animals, especially dogs, since it could be helpful to obtain faster data about the effects in mammals, and possibly buy a few people additional time with their beloved pets :)
(ii) What technology or technologies would you use to perform these DNA edits and why?
Also answer the following questions:
- How does your technology of choice edit DNA? What are the essential steps?**
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?**
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?**
I would use a CRISPR-based editing system and deliver it with AAV (the current most widely-used delivery mechanism for gene therapies). AAV is safer because it doesn’t integrate, which means that it doesn’t actually modify the existing genome (which can cause issues like cancer through insertional mutagenesis).
The main limitation of AAVs is their small cargo sizes, which severely limits the number of genes we could deliver through this method.
STEPS
- Found this paper using Perplexity: https://pmc.ncbi.nlm.nih.gov/articles/PMC5399099/
- Chose SIRT1 because of previous familiarity with it (it’s “famous” within the longevity field)
- Asked how to “reverse translate” this protein, got instructions to go to https://www.novoprolabs.com/tools/revtrans
- Found the Homo sapiens codon preference table: https://www.kazusa.or.jp/codon/cgi-bin/showcodon.cgi?species=9606&aa=SELECT+A+CODE&style=GCG and pasted into the tool
- (PENDING describe the rest and attach Perplexity conversation URL: https://www.perplexity.ai/search/spermidine-is-a-protein-right-lcAd5toGRwm1Cqu6_a_ang#0 )