Week 9 HW: Cell-Free Systems
Week 9 Homework: Cell-Free Systems
[WIP!]
Homework Part A: General and Lecturer-Specific Questions
General homework questions
- Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
The benefit can be summarized as “it removes the constraints we usually face for protein synthesis when working with live cells”. For example, working with live cells requires culturing cells throughout the whole cell lifecycle, with all the possibilities for error that this implies (mistakes, unexpected cell behavior due to their non-deterministic nature, etc), as well as the required timelines (speed is limited by the fundamental speed constraints from the cell growth cycle), and costs. Cell-free systems also allow for much greater control, as the main system is boiled down to its most basic functional components, removing a lot of complexity (variables outside of our control), and therefore allowing the possibility of producing much more homogeneous products.
Two special cases that CFS allow are the use of molecules that would usually be toxic for cells, and working with a much wider variety of molecules that would otherwise be destroyed by the cell, as well as modified versions of the most basic components of this machinery (that could not work in a live cell), such as amino acids that don’t exist in nature.
- Describe the main components of a cell-free expression system and explain the role of each component.
The basic components are: 1) The lysate, which includes the “machinery”: basically ground up cells (prokaryotic or eukaryotic), providing the cellular components we need for the reactions, like ribosomes, enzymes, tRNAs and cofactors, which will translate mRNA into protein 2) The genetic template (DNA or RNA) that encodes the product we want 3) The “building blocks” (amino acids and nucleotides) 4) The energy system (needed to provide and replenish the ATP and GTP needed for the reactions) (ie Mg^2+ and K+) 1) Note: CFS use Phosphoenolpyruvate (PEP) for this 5) The “environmental tuning” factors (salts, buffers, temperature, etc)
Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Because protein synthesis is an energy-intensive process. In living cells, mitochondria handle the production of ATP, but in CFS we need to set up systems that can continually obtain ie the phosphates needed to convert ADP into ATP. Not being able to meet these energy needs will cause the reactions to stall or provide low yield.
Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
Only eukaryotic cells can perform PTM post-translational modifications (ie glycosylation), so any desired product requiring this process will require the use of eukaryotic cell lysate. This will also be required if the final desired product requires the usage of mammalian regulators. However, when these requirements are absent, prokaryotic systems are faster and cheaper.
How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
The main challenge when working with membrane proteins is that they tend to aggregate or misfold in aqueous environments (due to their hydrophobic regions trying to “escape” it) [ PENDING ]
- Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
- The required energy needs are not being met.
- The proteins might be misfolding. In this case, we would check and tune the environmental variables, like the temperature or the salsts, and possibly add chaperones or use different membrane-mimicking systems.
- The final or intermediary products might be suffering degradation. In this case, I would check for the presence of ie nucleases, proteases, etc.
Homework question from Kate Adamala
Design an example of a useful synthetic minimal cell as follows:
- Pick a function and describe it.
- What would your synthetic cell do? What is the input and what is the output?
- Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
- Could this function be realized by genetically modified natural cell?
- Describe the desired outcome of your synthetic cell operation.
- Design all components that would need to be part of your synthetic cell.
- What would be the membrane made of?
- What would you encapsulate inside? Enzymes, small molecules.
- Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
- How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?)
- Experimental details
- List all lipids and genes. (bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.)
- How will you measure the function of your system?
PENDING
Example solution
Based on: Lentini, R. et al., 2014. Nat comm, 5, p.4012.
- Pick a function and describe it.
- What would your synthetic cell do? What is the input and what is the output?
Expand the sensing capacity of bacteria. Input: theophylline (inert to bacteria). Output of the SMC: IPTG. Output of the whole system: GFP produced in bacteria. (Theophyline aptamer reference: Martini, L. & Mansy, S.S., 2011. Cell-like systems with riboswitch controlled gene expression. Chemical Communications, 47(38), p.10734.)- Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
No. If the IPTG were not encapsulated, it would go into the bacteria without the need of theophylline-induced membrane channel synthesis, thus the synthetic cell actuator would not exist.- Could this function be realized by genetically modified natural cell?
Yes, in this particular case: the theophylline aptamer could be incorporated into a transformed gene. This lacks generality though – it is easier to make SMC than modify bacteria, so in this system a single bacteria reporter can be used to detect various small molecules.- Describe the desired outcome of your synthetic cell operation.
In the presence of SMC, bacteria sense theophylline.- Design all components that would need to be part of your synthetic cell.
- What would be the membrane made of?
Phospholipids + cholesterol.- What would you encapsulate inside? Enzymes, small molecules.
cell-free Tx/Tl system, IPTG, gene for membrane transporter under the control of theophylline aptamer.- Which organism your Tx/Tl system will come from? Is bacterial OK, or do you need a mammalian system for some reason? (hint: for example, if you want to use small molecule modulated promotors, like Tet-ON, you need mammalian)
Bacterial, because of the theophylline riboswitch used as SMC input.- How will your synthetic cell communicate with the environment? (hint: are substrates permeable? or do you need to express the membrane channel?)
The membrane is permeable to the input molecule (theophylline), the output is IPTG that will cross the membrane via the membrane pore created after theophyline-initiated gene expression.- Experimental details
- List all lipids and genes. (bonus: find the specific genes; for example, instead of just saying “small molecule membrane channel” pick the actual gene.)
- Lipids: POPC, cholesterol
- Enzymes: bacterial cell-free Tx/Tl
- Genes: a-hemolysin (aHL) to encapsulate in SMC
- Biological cells: E.coli transformed with GFP under T7 promoter and a lac operator
- How will you measure the function of your system?
Measure GFP output of the cells via flow cytometry. Alternatively, use enzymatic reporter, like luciferase, and measure bulk output of the enzyme.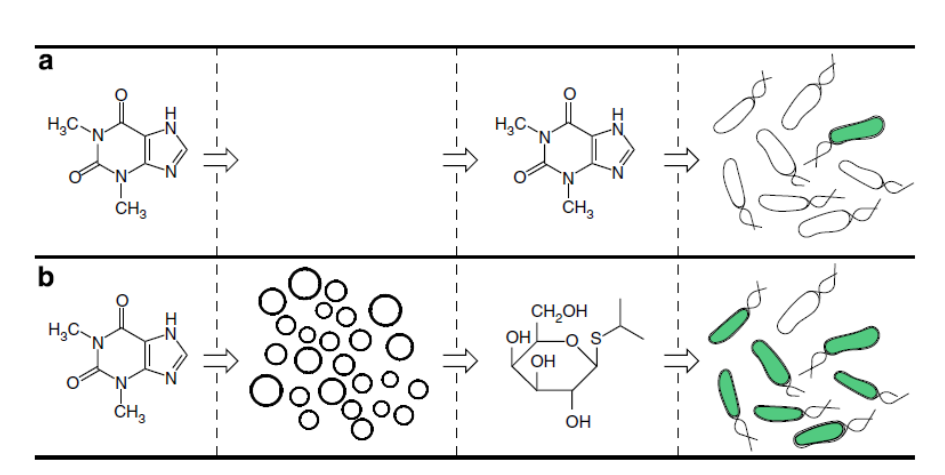
Artificial cells translate chemical signals for E. coli.
(a) In the absence of artificial cells (circles), E. coli (oblong) cannot sense theophylline.
(b) Artificial cells can be engineered to detect theophylline and in response release IPTG, a chemical signal that induces a response in E. coli.
Homework question from Peter Nguyen
Freeze-dried cell-free systems can be incorporated into all kinds of materials as biological sensors or as inducible enzymes to modify the material itself or the surrounding environment. Choose one application field — Architecture, Textiles/Fashion, or Robotics — and propose an application using cell-free systems that are functionally integrated into the material. Answer each of these key questions for your proposal pitch:
- Write a one-sentence summary pitch sentence describing your concept.
- How will the idea work, in more detail? Write 3-4 sentences or more.
- What societal challenge or market need will this address?
- How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
MY ANSWERS:
- Portable FD-CF pathogen sensor patches embedded in taxi and public transport vehicles to monitor airborne pathogen load
- Leveraging a freeze-dried system with an RNA switch (or CRISPR-based) detector, this device could be carried by transport service providers who could periodically measure the pathogen levels in the vehicle with a swab. It could be integrated into seatbelts or any other part of the car (it could be a patch or sticker).
- Use cases: 1) identify when the vehicle should be more thoroughly disinfected 2) driver safety 3) passenger safety (for example, riders who are especially immunocompromised could request vehicles with lower pathogen levels) 4) data collection for scientific purposes (ie epidemic detection and prevention)
- The device could be a patch with a cartridge containing the necessary water and buffers and a button or switch that the driver could click to easily mix the components, for example at the end of a day’s shift. The device would be single-use and replaced daily to ensure minimal contamination. The freeze-dried system ensures stability for at least 3-6 months at room temperature.
Homework question from Ally Huang
Freeze-dried cell-free reactions have great potential in space, where resources are constrained. As described in my talk, the Genes in Space competition challenges students to consider how biotechnology, including cell-free reactions, can be used to solve biological problems encountered in space. While the competition is limited to only high school students, your assignment will be to develop your own mock Genes in Space proposal to practice thinking about biotech applications in space!
For this particular assignment, your proposal is required to incorporate the BioBits® cell-free protein expression system, but you may also use the other tools in the Genes in Space toolkit (the miniPCR® thermal cycler and the P51 Molecular Fluorescence Viewer). For more inspiration, check out https://www.genesinspace.org/ .
- Provide background information that describes the space biology question or challenge you propose to address. Explain why this topic is significant for humanity, relevant for space exploration, and scientifically interesting. (Maximum 100 words)
- Name the molecular or genetic target that you propose to study. Examples of molecular targets include individual genes and proteins, DNA and RNA sequences, or broader -omics approaches. (Maximum 30 words)
- Describe how your molecular or genetic target relates to the space biology question or challenge your proposal addresses. (Maximum 100 words)
- Clearly state your hypothesis or research goal and explain the reasoning behind it. (Maximum 150 words)
- Outline your experimental plan - identify the sample(s) you will test in your experiment, including any necessary controls, the type of data or measurements that will be collected, etc. (Maximum 100 words)
MY ANSWERS (PENDING)