Subsections of HTGAA 2026 - Georg Tremmel, Global TA & Node Lead: BioClub Tokyo
Homework
Weekly Homework:
Week 5 Homework - Protein Design ⓶
Notes Part A: SOD1 Binder Peptide Design A.1: Generate Binders with PepMLM A.1.1: Retrieve SOD1 and introduce the A4V mutation Note Begin by retrieving the human SOD1 sequence from UniProt (P00441) and introducing the A4V mutation.
Week 7 - Neuromorphic Networks
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs) Week 7 Homework Overview Questions What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions? Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal. Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation. Designing IANNs with the Neuromorphic Wizard
Subsections of Homework
Week 5 Homework - Protein Design ⓶
Notes
Part A: SOD1 Binder Peptide Design
A.1: Generate Binders with PepMLM
A.1.1: Retrieve SOD1 and introduce the A4V mutation
Note
Begin by retrieving the human SOD1 sequence from UniProt (P00441) and introducing the A4V mutation.
Find the entry for P00441 SODC_HUMAN at UniProt
!()[uniprot01.jpg?width=768px]
By clicking on the Amino Acids go to sequence link, we arrive at the sequence.
!()[uniprot02.jpg?width=768px]
The Download links downloads the FASTA file, but you can also just copy the sequence.
I manually introduce the A4V mutation, mutating Alanine to Valine at residue 4.
Note
Why exactly does A4V means?
If we start counting at 1, then the 4th residue is K.
If we start counting at 0, then the 4th residue is actually A.
Which one is correct?
Both. Neither?
Proteins usually start with the start codon, which translates to M (methionine), which is often removed after translation, therefore MATKAVC.. becomes ATKAVC... When we not start counting residues at 1, then 4 is A
A.1.2: Use the PepMLM Colab
Copy the [https://colab.research.google.com/drive/1u0i-LBog_lvQ5YRKs7QLKh_RtI-tV8qM?usp=sharing](PepMLM Colab), from ChatterjeeLab/PepMLM-650M
A.1.3: Generate four peptides of length 12
Binder,Pseudo Perplexity HRYYVAAVRHWK,23.837650573350423 WRSPVVVAEHKK,13.883698506608807 WLYYAAALRLKE,19.11173304337227 WRYYAAALAWGX,10.052019407114733
A.1.4: Add known SOD1-binding peptide FLYRWLPSRRGG
HRYYVAAVRHWK WRSPVVVAEHKK WLYYAAALRLKE WRYYAAALAWGX FLYRWLPSRRGG
A.2 Evaluate Binders with AlphaFold3
A.2.1 Evaluate Binders with AlphaFold3
alphafoldserver.com
Week 7 - Neuromorphic Networks
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
Questions
- What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
- Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
- Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.
Designing IANNs with the Neuromorphic Wizard
Assignment Part 2: Fungal Materials
- What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
- What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Subsections of Week 7 - Neuromorphic Networks
Week 7 - Neuromorphic Networks Lab
Pre-lab: Installing the Neuromorphic Wizard
Subsections of Week 7 - Neuromorphic Networks Lab
Week 7 - Neuromorphic Networks Lab
Info
Installation Guide and Walk-through of the Neuromorphic Wizard Software.
Pre-lab: Installing the Neuromorphic Wizard
The following instruction were made with macOS 14.
Downloading the Software
Evan is sharing the software as a Google Folder. Download the Folder by clicking Download in the Folder Menu.
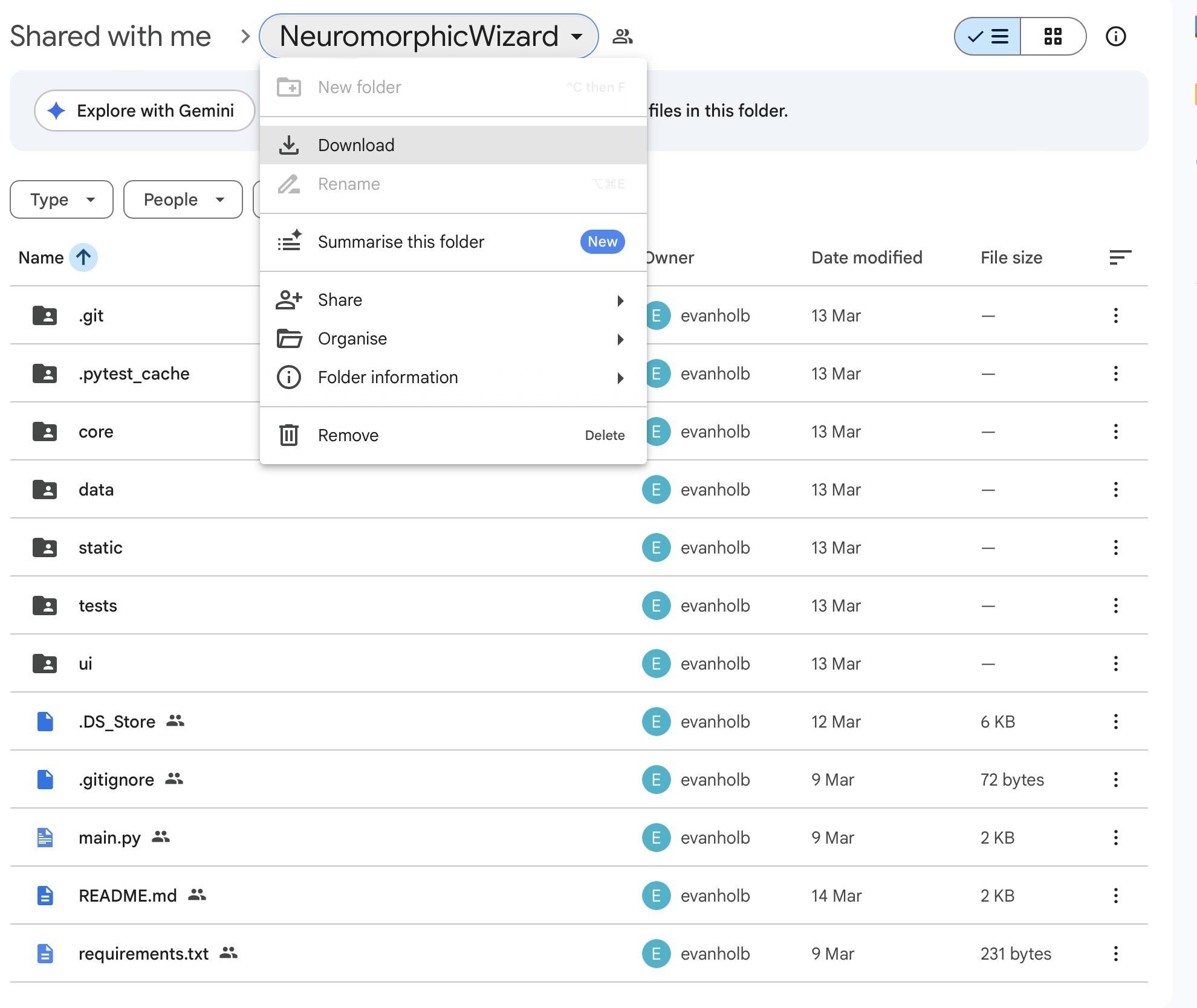

Installing the Anaconda Package Manager
The Neuromorphic Wizard needs the Python Distribution & Package ManagerAnaconda to run. If you don’t have Anaconda on your system, download and install it.
You can check if you successfully installed Anaconda, by opening a Terminal and saying the following:
Which should give your something like:
Make sure that Anaconda is running before you proceed.
Installing the Software
We are installing the software from the Terminal.
Change into the Download Directory
In my case, I the directory into my Development Folder.
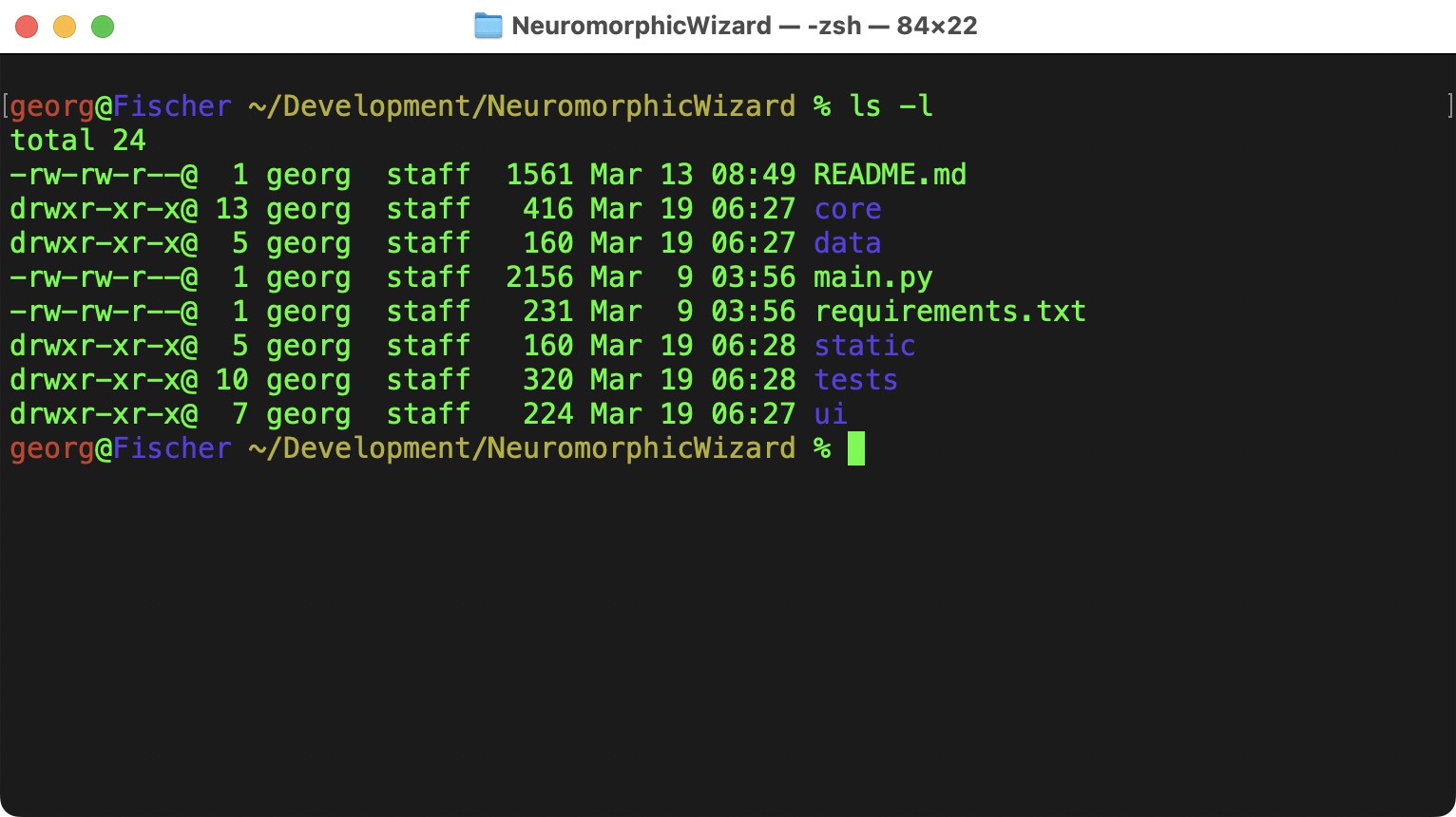

The README.md has more details about the installtion, but I am also summarizing them here.
Creating a Virtual Environment with Conda
Although there is a version of Python on most computers, we can use Virtual Environment to create an Environment that does not interfere with any other Python installation on your computer.
To create an enviroment with Python 3.10:
To activate it:
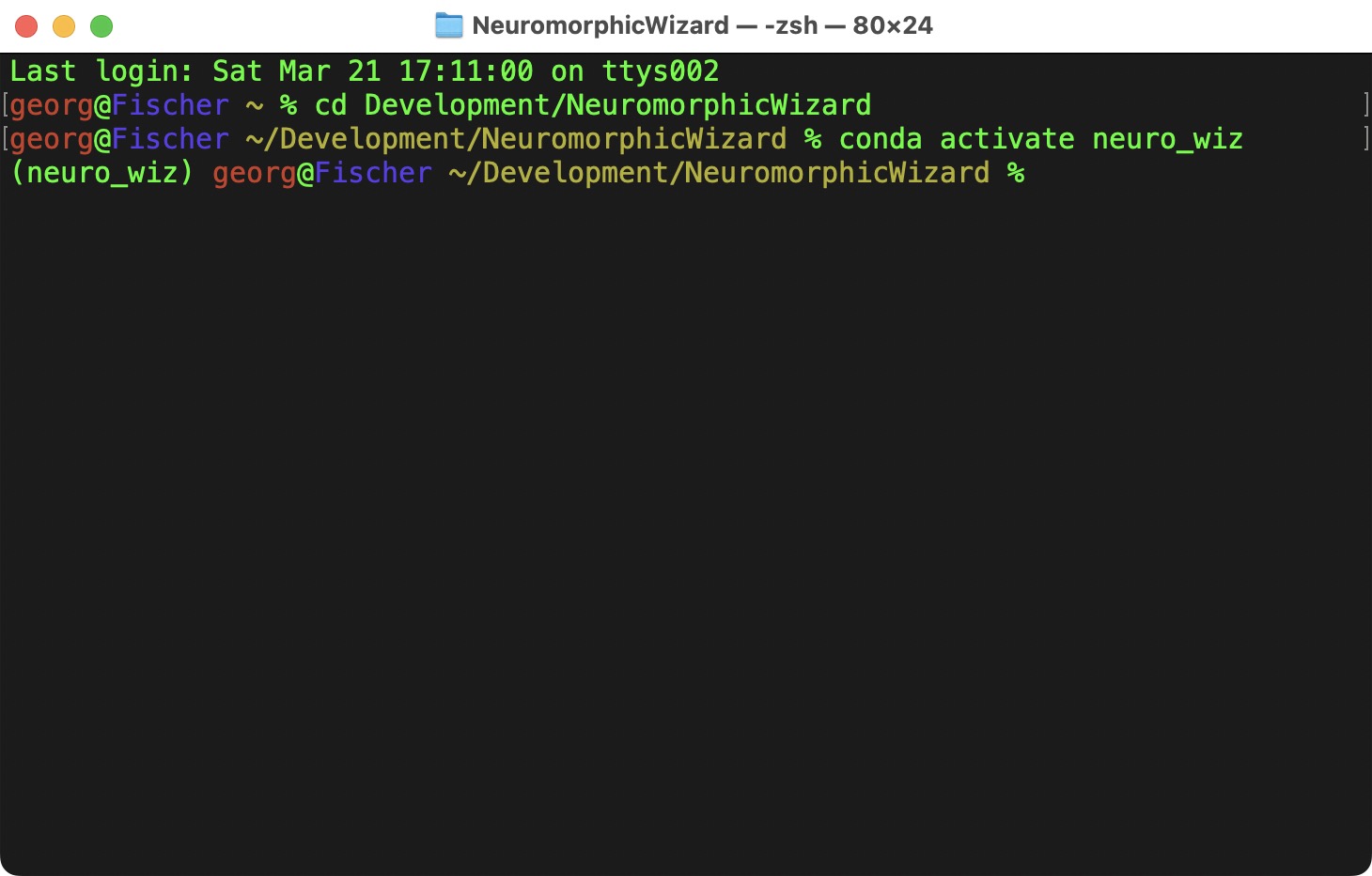

Now we have a fresh enviroment, where we can install the Neuromorphic Wizard and its dependencies into.
pip is the Python Package Manager, it takes the requirements listed in requirements.txt and install all the necessary packages.
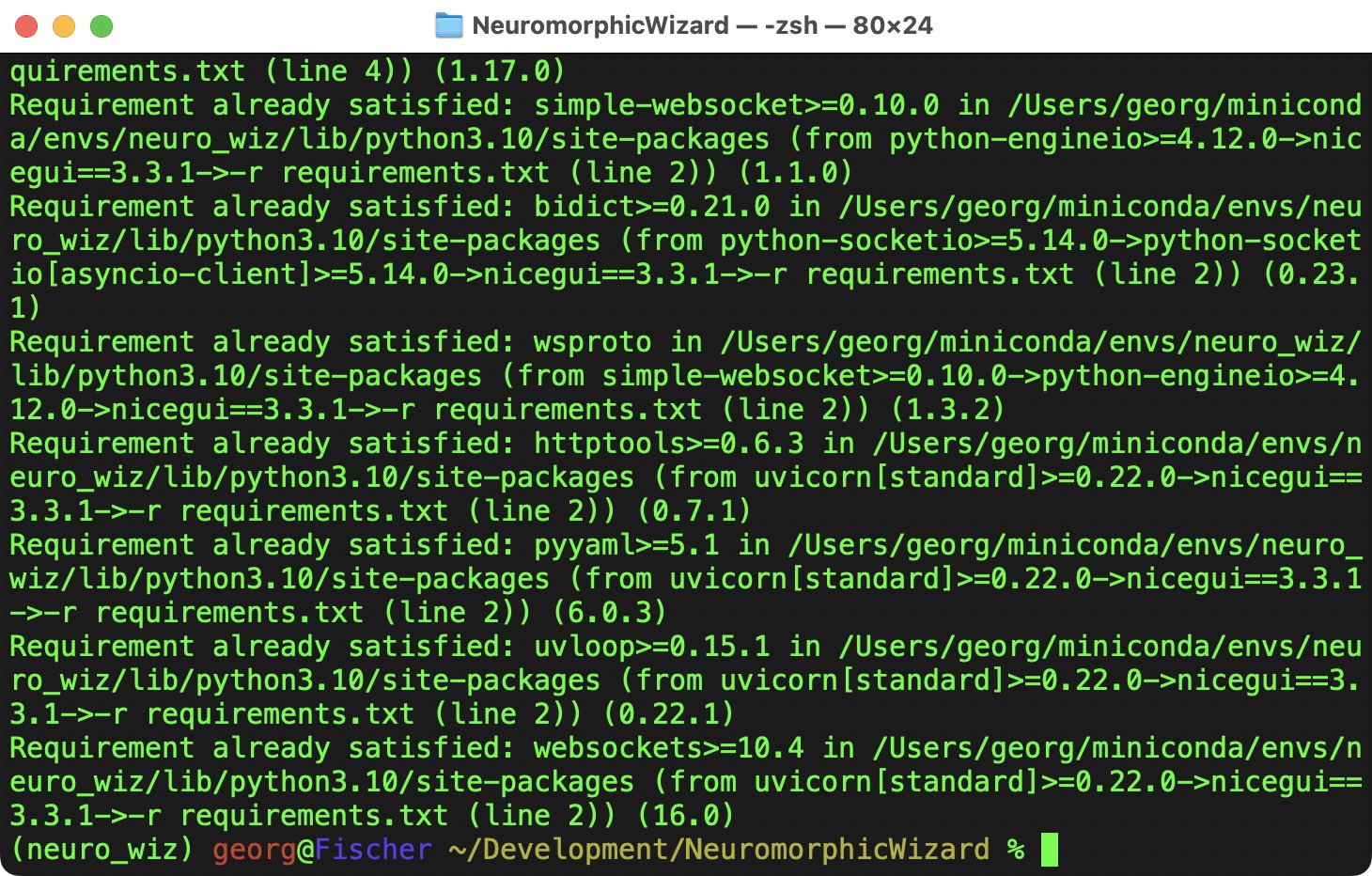

Starting the Application

I might take a couple of seconds to start the application. A new browser should open - if it does not open automatically, go to http://localhost:8080.
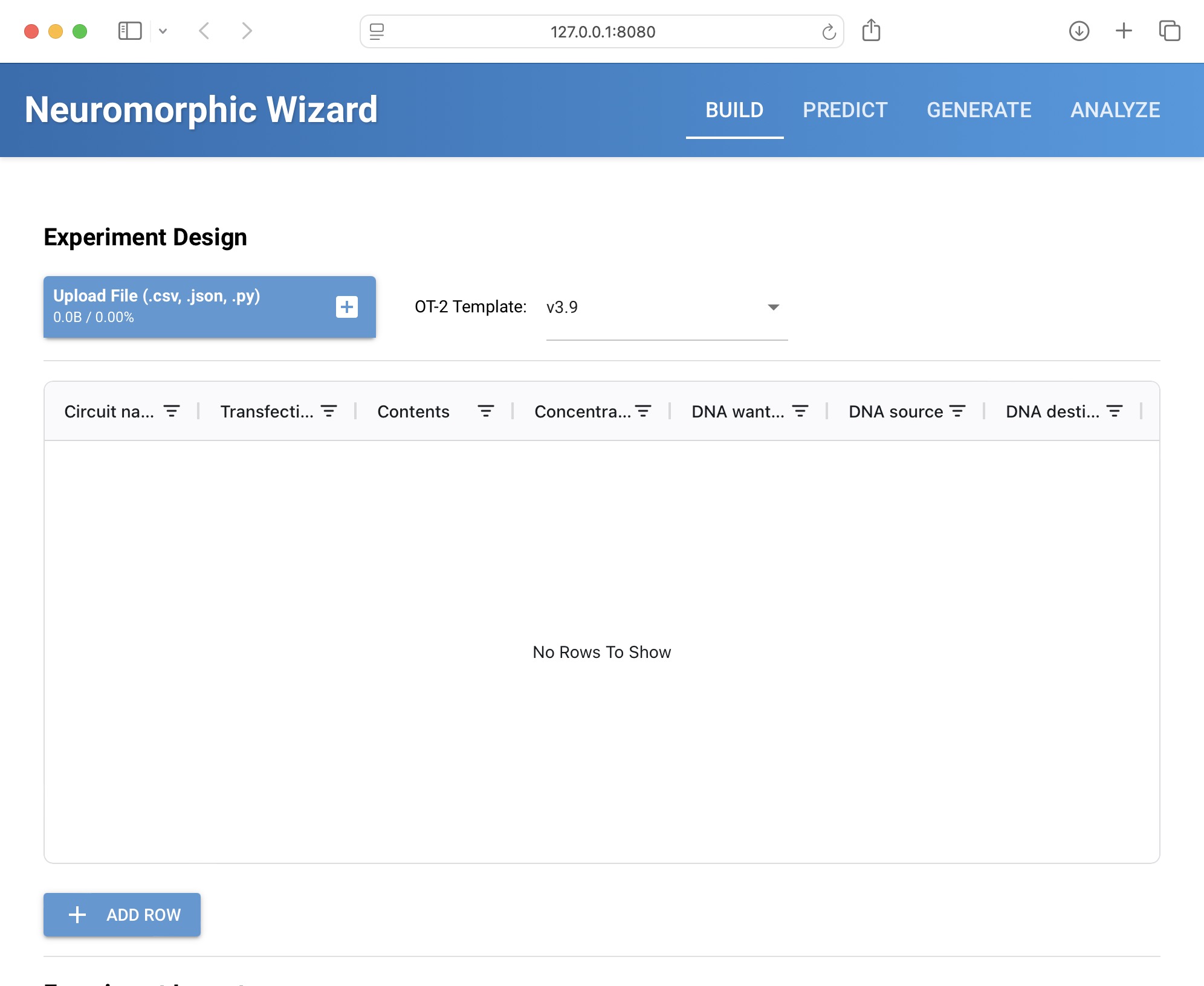

Now we are ready to design our IntraCellular Artificial Neural Networks!
Pages & Documentation
For 2026 HTGAA Committed Listeners are required to submit their homework and documentation with and through the git-based HTGAA 2026 Documentation Server. Each students will get their own git repository, from which your personal student website will be automatically build.
Access
Registered participants of HTGAA 2026 will be given access to the HTGAA Documentation Server after Class #1 on Feburary 3rd.
Example Repository:
https://edit.htgaa.org/georg-tremmel/webpages
Example Website: (will be build automatically from the repository)
https://pages.htgaa.org/georg-tremmel
All your homework and documentation will be publicly accessible.
- To Do: Add Screenshots
Authoring
Markdown Syntax
Your content will be written in Markdown, a simple syntax to add formatting to plain text. If you are unfamiliar with Markdown, please have a look at this Introduction to Markdown.
Editing Files with the Online Editor (easy)
The HTGAA Repositories have a web-based editor:
- Log into your account
- Navigate to the file you want to edit
- Click the
Edit fileicon - Edit
- After Editing, save the changes by clicking
Commit Changes
- To Do: Add Screenshots
Adding Images
- To Do: How to add images
- To Do: How to scale down images
Cloning your Repo and runnig a Local Instance of Hugo (advanced)
or
git clone https://edit.htgaa.org/georg-tremmel/webpages.git my-local-directory
Git
Files can now be edited on your computer. To push the changes to the server and update the website, the follow steps are necessary:
| Git Command | |
|---|---|
1. git add * | Add all new and changed files |
2. git commit | Locally commit your changed. You will be asked to provide add a short message. Can also be concatenated into `git commit -m ‘my message`` |
3. git push | Uploads and pushes the changes onto the HTGAA Server, the website is rebuilt after a few seconds. You will be asked for your credentials. |
Git is much more powerful - and complex - but these 3 commands will be used most often.
If you change file on the server, make sure to do a git pull to download the latest changes.
Running a Local Version of Hugo
- Install the lastest version of Hugo.
Technical Details
- Forgejo self-hosted lightweight git-based software forge’, functional similar to GitHub and GitLab
- Hugo SSG (Static Site Generator) with the Relearn Theme
You don’t need to edit/change your Hugo theme. But if you really (really) want, you can. But make sure we can navigate the site and find your homework.
To Do
- Add Screenshots showing the login process
MoUs
Sign the MoU by copying and committing it to your repository, and add your name and date.
- Memorandum of Understanding for BioClub Committed Listeners
- Memorandum of Understanding for BioClub GLobal Teaching Assistants
Here is my signed TA MoU.
Subsections of MoU
BioClub Committed Listener MoU
HTGAA Committed Listener (CL) Agreement
I am a HTGAA Committed Listener, my responsibilities are:
- Watching class lectures and recitations
- Participating in node reviews
- Developing and documenting my homework
- Actively communicating with other students and TAs on the forum
- Allowing HTGAA and BioClub to share my work (with attribution)
- Honestly reporting on my work, and appropriately attributing and citing the work of others (both human and non-human)
- Following locally applicable health and safety guidance
- Promoting a respectful environment free of harassment and discrimination
Signed by committing this file to my documentation page/repository,
{{ your name goes here }}
{{ date }}
BioClub Global TA MoU
HTGAA Teaching Assistant (TA) Memorandum of Understanding (MoU)
By signing this form, I agree to fulfill the following responsibilities as a teaching assistant for How To Grow Almost Anything (HTGAA) 2026.
As an HTGAA Teaching Assistant, I agree to:
- Collaboration: Work with the instructor team at my node to carry out agreed-upon tasks throughout the semester and ensure students at my node understand all requirements for success.
- Safety: Adhere to all applicable local, regional, and national health and safety guidelines.
- Communication: Communicate clearly with students and instructors, raising any issues or concerns promptly.
- Respect: Promote a respectful, inclusive, and welcoming learning environment, encouraging curiosity and peer support.
- Integrity: Demonstrate integrity as a scientist and educator by keeping data accurate, complete, and truthful.
- Attribution: Grant HTGAA permission to share my work produced for the course, with appropriate attribution.
- Mentorship: Mentor students with a continuous learning mindset, supporting their curiosity, growth, and resilience, and celebrating their achievements.
- Initiative: Proactively contribute ideas and solutions to enhance the course, lab experiences, and student learning.
- Adaptability: Approach challenges with flexibility and problem-solving, modeling resilience in a dynamic learning environment.
Node-Specific Agreements
I also agree to carry out responsibilities assigned to support my node’s success, including:
- Engagement: Participate in required node meetings and coordinate with your node team to support planning, collaboration, and student success.
- Preparation: Preparing instructional materials ahead of classes and labs and running validation experiments to strengthen the learning experience.
- Support: Maintaining availability to support students through established communication channels (e.g., email, Discourse).
- Review: Reviewing student homework and providing regular, constructive feedback.
- Attendance: Tracking and maintaining accurate attendance records for students at my node.
Signed by committing this file in my repository,
{{ your name }}
{{ date }}
My Signed TA MoU
HTGAA Teaching Assistant (TA) Memorandum of Understanding (MoU)
By signing this form, I agree to fulfill the following responsibilities as a teaching assistant for How To Grow Almost Anything (HTGAA) 2026.
As an HTGAA Teaching Assistant, I agree to:
- Collaboration: Work with the instructor team at my node to carry out agreed-upon tasks throughout the semester and ensure students at my node understand all requirements for success.
- Safety: Adhere to all applicable local, regional, and national health and safety guidelines.
- Communication: Communicate clearly with students and instructors, raising any issues or concerns promptly.
- Respect: Promote a respectful, inclusive, and welcoming learning environment, encouraging curiosity and peer support.
- Integrity: Demonstrate integrity as a scientist and educator by keeping data accurate, complete, and truthful.
- Attribution: Grant HTGAA permission to share my work produced for the course, with appropriate attribution.
- Mentorship: Mentor students with a continuous learning mindset, supporting their curiosity, growth, and resilience, and celebrating their achievements.
- Initiative: Proactively contribute ideas and solutions to enhance the course, lab experiences, and student learning.
- Adaptability: Approach challenges with flexibility and problem-solving, modeling resilience in a dynamic learning environment.
Node-Specific Agreements
I also agree to carry out responsibilities assigned to support my node’s success, including:
- Engagement: Participate in required node meetings and coordinate with your node team to support planning, collaboration, and student success.
- Preparation: Preparing instructional materials ahead of classes and labs and running validation experiments to strengthen the learning experience.
- Support: Maintaining availability to support students through established communication channels (e.g., email, Discourse).
- Review: Reviewing student homework and providing regular, constructive feedback.
- Attendance: Tracking and maintaining accurate attendance records for students at my node.
Signed by committing this file in my repository,
Georg Tremmel
28.2.2026